Protocols
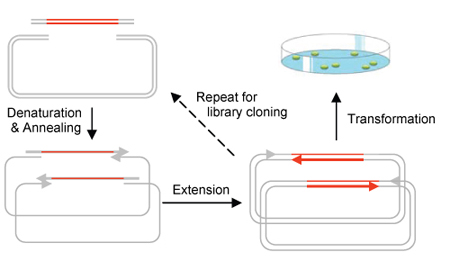
CPEC Cloning
Back to top
CPEC cloning
Materials:
• Phusion™ High-Fidelity PCR Kit (FINNZYMES, Cat. No. F-553)
• Thermocycler
Protocol:
5x Phusion HF Buffer 4 ul
10 mM dNTPs 0.4 ul
Vector 50 ng/1kb
Insert x ng*
Phusion DNA Polymerase 0.2 ul
H2O to 20 ul
- The amount of insert is determined so that the molar ratio for vector and insert is 1 to 2.
98°C 30sec
10X
98°C 10 sec
Annealing** 30 sec
72°C x sec***
72°C 5min
4°C hold
- Anneal at Tm + 3°C. The Tm should be calculated with the nearest-neighbor method.
- The extension time is usually calculated according to the shortest piece with 15 sec /kb if the cloning is not complicated. For example, if there is only one insert and is shorter than the vector, say, 600 bp, then I will use 15 sec for extension. Refer to the published paper for detailed information.
Back to top
DNA Purification
Back to top
DNA purification
Materials :
• E.Z.N.A Gel Purification Kit (Omega Bio-Tek, Cat No. D2500-02 )
• Water bath equilibrated to 55-65C
• Microcentrifuge capable of at least 10,000 x g
• Nuclease-free 1.5 ml centrifuge bottles
• Absolute (95%-100%) ethanol
• Protective eye-wear
• Isopropanol (for fragments < 500 bp only)
Protocol:
1. Perform agarose gel electrophoresis to fractionate DNA fragments. Any type or grade of agarose may be used. It is strongly recommended, however, that fresh TAE buffer or TBE buffer be used as running buffer. Do not re-use running buffer as its pH will increase and reduce yields.
2. When adequate separation of bands has occurred, carefully excise the DNA fragment of interest using a wide, clean scalpel.
3. Determine the approximate volume of the gel slice by weighing it in a clean 1.5 ml microfuge tube. Assuming a density of 1 g/ml of gel, the volume of gel is derived as follows: A gel slice of mass 0.3 g will have a volume of 0.3 ml. Add equal volume of Binding Buffer (XP2). Incubate the mixture at 55C-60C for 7 min or until the gel has completely melted. Mix by shaking or inverting the tube every 2-3 minutes. Centrifuge the tube briefly to collect all the liquid to the bottom of the tube.
Note: For DNA fragment less than 500bp, add 1 sample volume of isopropanol after the addition of Binding Buffer (XP2).
4. Apply up to 700 ul of the DNA/agarose solution to a HiBind® DNA spin column assembled in a clean 2 ml collection tube (provided) and centrifuge in a microcentrifuge at 8,000-10,000 x g for 1 min at room temperature. Discard the liquid. Re-use the collection tube in Steps 5-8. For volumes greater than 700 ul, load the column and centrifuge successively, 700 ul at a time. Each HiBind® spin-column has a total capacity of 25-30 ug DNA.
5. Discard liquid and add 300ul Binding Buffer. Centrifuge at 10,000 x g for 1 minutes.
6. Add 700 ul of SPW Buffer diluted with absolute ethanol into the column and wait 2-3 minutes. Centrifuge at 10,000 x g for 1 min at room temperature to wash the sample.
7. Discard liquid and repeat step 6 with another 700 ul SPW Buffer.
8. Discard liquid and, re-using the collection tube, centrifuge the empty column for 1 min at maxi speed (>13,000 x g) to dry the column matrix. This drying step is critical for good DNA yields.
9. Place column into a clean 1.5 ml microcentrifuge tube (not provided). Add 30-50 ul depending on desired concentration of final product) Elution Buffer (or sterile deionized water) directly to the center of the column matrix, then incubate for 1 minute. Centrifuge 1 min at maxi speed (>13,000 x g) to elute DNA. This represents approximately 70% of bound DNA. An optional second elution will yield any residual DNA, though at a lo
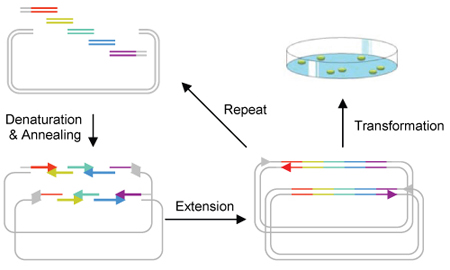
PCA (Polymerase Cycle Assembly
Back to top
Materials:
• Phusion™ High-Fidelity PCR Kit (FINNZYMES, Cat. No. F-553)
• Thermocycler
Protocols:
5x Phusion HF Buffer 5 ul
10 mM dNTPs 0.5 ul
Oligo mixture 125 ng /250 ng /500 ng /…
Phusion DNA Polymerase 0.25 ul
H2O to 25 ul
98°C 30sec
40X
98°C 7 sec
70-50°C slow ramp, 0.1°C/sec
50°C 30 sec
72°C 15 sec /kb
72°C 5 min
4°C hold
PCR Product Clean-up for DNA Sequencing
Back to top
Materials:
• ExoSAP-IT® (usb, Cat. No. 78200)
• Thermocycler
Protocol:
1. Remove ExoSAP-IT from -20°C freezer and keep on ice throughout this procedure.
2. Mix 5 μl of a post-PCR reaction product with 2 μl of ExoSAP-IT for a combined 7 μl reaction volume.
3. Incubate at 37°C for 15 min to degrade remaining primers and nucleotides.
4. Incubate at 80°C for 15 min to inactivate ExoSAP-IT.
5. The PCR product is now ready for use in DNA sequencing etc.
==PCR
Back to top
Materials:
• Phusion™ High-Fidelity PCR Kit (FINNZYMES, Cat. No. F-553)
• Thermocycler
Protocols:
5x Phusion HF Buffer 10 ul
10 mM dNTPs 1 ul
DNA template 1 pg – 10 ng
Forward primer (10 uM) 2.5 ul
Reverse primer (10 uM) 2.5 ul
Phusion DNA Polymerase 0.5 ul
H2O to 50 ul
98°C 30sec
30X
98°C 10 sec
Annealing* 30 sec
72°C 15 sec /1 kb
72°C 5min
4°C hold
- Anneal at Tm + 3°C. The Tm should be calculated with the nearest-neighbor method.
==Single Colony PCR
Back to top
Materials:
• Taq DNA Polymerase with Standard Taq Buffer (NEB, Cat. No. M0273)
• 10 mM dNTP Mix (NEB, Cat. No. N0447)
• Thermocycler
Protocols:
10x Standard Taq Buffer 2 ul
10 mM dNTPs 0.4 ul
Bacteria culture* 1 ul
Forward primer (10 uM) 1 ul
Reverse primer (10 uM) 1 ul
Taq DNA Polymerase 0.2 ul
H2O to 20 ul
- Bacteria culture refers to E. coli cultured in LB solution overnight.
94°C 5 min
25X
94°C 15 sec
Annealing ** 30 sec
72°C 1 min /1 kb
72°C 5min
4°C hold
- Anneal at Tm which is calculated with salt-adjusted method.
==Transformation
Back to top
Materials:
• GC5 Chemical Competent Cells (Genesee Scientific, Cat. No. 42-653)
• SOC Medium (Sigma, Cat. No. S1797)
• LB Agar (Sigma, Cat. No. L3027)
• Petri Dishes (VWR, Cat. No. SC25373-187)
• Cell Spreader (VWR, Cat. No. 89042-018)
• 37°C incubator
• 37°C shaker
• water bath
Protocol:
1. Thaw 1 tube of competent cells on ice;
2. Add 3 ul of cloning product or 1-50 ng of plasmid into competent cells while stirring gently;
3. Keep the tube covered by ice for 30min;
4. Heat-shock the competent cells in water bath for 45 sec at 42°C;
5. Put the tube on ice for 2 minutes;
6. Add 450 ul of SOC medium and then put it in a 37°C shaker for 1 hour;
7. Dilute and spread an appropriate amount on an LB agar plate with the appropriate antibiotics;
8. Place the plate up-side-down in 37°C incubator for 16-18 hours (overnight).
6/3/09
Experiment: PCR with contents of tube labeled 7 as DNA template, Gel with
100 bp ladder to check PCR
Results: Incorrect ladder and template
Conclusion:
Should use 1 kb ladder, Need to find correct template
6/4/09
E: Gel
to check Bioplastics #1-5
R: All correct sized bands
C: Use one as template
for PCR
E: PCR with Bioplastics #5 as template
5x HF buffer 5
ul
dNTPs 2 ul
forward primer 1.25 ul
reverse primer 1.25 ul
DNA template
(plasmid) 0.5 ul
Phusion DNA polymerase 0.3 ul
H2O 14.7 ul
25 ul
98°C 30s 1x
98°C 15s
55°C 30s
72°C 1m 10s 35x
72°C
5m
4°C infinity 1x
Fragment # Name Size/Length
1 pASKphaA 4.2
kb
2 phaB 900 bp
3 Termcat2 1.7 kb
4 phaC 1.9 kb
6/5/09
E:
Gel to check PCR
R: Correct band for fragments 2 and 4, No band for fragments
1 and 3
C: Cut correct bands out of gel, Try PCR with another template, different
conditions
E: PCR with Bioplastics #1 as template, increased extension
time to 1m 20s
6/6/09
R: Correct band for fragments 1 (weak) and 2,
No band for fragments 3 and 4
C: Cut out correct bands, Do PCR again with same
template for fragments 1 and 2, Try PCR with different template for fragments
3 and 4
E: PCR with Bioplastics #1 (fragments 1 and 2) and #3 (fragments
3 and 4) as templates
6/8/09
E: Gel to check PCR, Gel with leftover
PCR of right sized fragments
R: Correct bands for all fragments
C: Cut out
bands
6/9/09
Gel Extraction
1. Weigh cut gel piece(s). Add at least
1 ul binding buffer per mg of gel.
2. Place tube at 50-60°C for 10-15 min
or until gel is completely dissolved. Vortex every few minutes.
3. Add solution
to spin column. Centrifuge at max speed for 30s. Discard filtrate.
4. Add 300
ul binding buffer to column. Centrifuge for 30s. Discard filtrate.
5. Add 500
ul wash buffer. Centrifuge for 30s. Discard filtrate.
6. Repeat step 5.
7.
Centrifuge tubes for at least 2 min. Discard collection tube.
8. Obtain 1.7
ml centrifuge tube. Add 40-50 ul elution buffer to spin column. Centrifuge for
1 min. Discard spin column.
E: Measured concentration of DNA fragments
R:
Very low concentrations of fragments 1 and 3
C: Do PCR with fragments as template,
greater volume to increase amount
E: PCR with fragments 1 and 3 as templates 6/10/09
E: Gel to check PCR
R: Smears at bottom of gel
C: Should use Bioplastics
#1 as template to increase amount of fragment 1 and Bioplastics #3 as template
to increase amount of fragment
E: PCR with Bioplastics #1 and #3 as templates
and doubled volume, Gel to obtain more of fragments
R: Smears for fragment
1, Correct bands for fragment 2, Correct bands and smears for fragment 3
C:
PCR with different conditions
E: PCR with Bioplastics #3 as template,
extension time increased to 1m 30s, cycles increased to 40
6/11/09
E:
Gels to check PCR
R: Correct bands for fragment 1, 2 and 4 and possibly correct
band for fragment 3
C: Cut out bands
E: Measured concentration of
DNA fragments
R: Still low concentrations of fragments 1 and 3
C: Do more
PCRs to obtain more of fragments 1 and 3
6/15/09
E:
PCRs with Bioplastics #1 (fragments 1 and 2) and #3 (fragments 3 and 4) as templates,
doubled volume, eliminated annealing step, varied extension (plus annealing) time
based on fragment size, Gels to check PCRs
5x HF buffer 10 ul
dNTPs
4 ul
forward primer 2.5 ul
reverse primer 2.5 ul
DNA template (plasmid)
1 ul
Phusion DNA polymerase 0.8 ul
H2O 29.2 ul
50 ul
98°C
30s 1x
98°C 15s
72°C varies 35x
72°C 5m
4°C infinity
1x
Annealing Time: 30s
Extension Time: 15 s * size/length (kb)
Fragment # Name Size/Length (bp) Time (s)
1 pASKphaA 4215 100
2 phaB
896 50
3 Termcat2 1701 60
4 phaC 1872 65
R: Bands for fragments
1 and 2, No bands for fragments 3 and 4
C: Cut out bands of fragments 1 and
2, PCR again to obtain fragments 3 and 4
6/16/09
E: PCR with Bioplastics
#2 as template and Phusion mix (done by Maggie), Gels to check PCRs
2x
Phusion mix 25 ul
forward primer 2.5 ul
reverse primer 2.5 ul
DNA template
(plasmid) 1 ul
H2O 19 ul
50 ul
R: No bands for fragments 3, Bands
for fragment 4
C: Cut out bands of fragment 4, PCR again to obtain fragment
3
E: Measured concentration of DNA fragments 1 and 2
R: Concentrations
okay
C: Do assembly PCR once enough of fragments 3 and 4 obtained
6/17/09
E:
PCR with Bioplastics #4 as template, using Phusion enzyme protocol (shorter denaturation
time, lower annealing temperature), Gel to check PCRs
98°C 15s 1x
98°C
1s
50°C 5s
72°C 30s 35x
72°C 2m
4°C infinity 1x
R: Correct bands
C: Cut out bands
E: Measured concentration of
DNA fragments 3 and 4
R: Concentrations high
C: Do assembly PCR
6/18/09
E: Assembly PCR, Gels to check PCRs
2x Phusion mix 12.5 ul
insert
7.14 ul
H2O 5.36 ul
25 ul
98°C 30s 1x
98°C 10s
70-55°C
5s
72°C 30s 25x
72°C 2m
4°C infinity 1x
Fragment
# Name Size/Length (bp) Concentration (ng/ul) Mass (ng) Volume (ul)
1 pASKphaA
4215 41 194 4.73
2 phaB 896 68 41 0.60
3 Termcat2 1701 69 78 1.13
4 phaC
1872 126 86 0.68
~400 7.14
Length of plasmid: 8360 bp + 4 Histags
* 6 aa * 3 bp = 8432 bp R: Smear because of misformed gel,
no band
C: Change conditions
6/19/09
E: Assembly PCR (done by Maggie)
with different conditions (decreased extension time, increased number of cycles)
5x HF buffer 5 ul
dNTPs 2 ul
insert 7.18 ul
Phusion DNA polymerase
0.4 ul
H2O 10.42 ul
25 ul
98°C 30s 1x
98°C 10s
70-55°C
30s
72°C 2m 40x
72°C 5m
4°C infinity 1x
Fragment
# Name Mass (ng) Volume (ul)
1 pASKphaA 195.3 4.76
2 phaB 41 0.60
3 Termcat2
78.4 1.14
4 phaC 86.4 0.68
~400 7.14
6/20/09
E: Gel to check PCR
R:
Correct band (faint)
C: Low efficiency, Use different method (Infusion kit
or PCR with 2 fragments first)
6/22/09
LB Agar Plates with Chlorophenicol
(CAM), anhydrotetracycline, Nile Red (NR)
stock plate
CAM 50 mg/ul 20 ug/ml
anhyd
2 mg/ml 150 ng/ml
NR 0.25 mg/ml 0.5 ug/ml
E: PCR with fragments 1 and
2 and different conditions (decreased annealing and extension times and number
of cycles), Gel to check PCR
2x Phusion mix 12.5 ul
pASKphaA 4.76
ul
phaB 0.60 ul
H2O 7.14 ul
25 ul
98°C 30s 1x
98°C
10s
70-55°C 10s
72°C 1m 10s 20x
72°C 5m
4°C infinity
1x
R: No band/smear
C: Change conditions
6/23/09
E: Assembly
PCR with different conditions (no slow ramp)
6/24/09
E: Gel to check
PCR
R: Correct band (faint)
C: Do transformation with competent cells
6/24/09
Transformation with High Efficiency GC5 Competent Cells
1. Remove competent
cells from -70°C and place on ice. Thaw for 5-10 min.
2. Gently mix cells
by tapping tube.
3. Add 1-50 ng DNA (1 ul control) into 50 ul cells. Swirl
pipette tip while dispensing DNA. Gently tap tube to mix.
4. Place tubes on
ice for 30 min.
5. Heat-shock cells for 45 sec in 42°C (water) bath. Do
not shake!
6. Add 450 ul RT SOC Medium to each transformation reaction.
7.
Incubate at 37°C for 1 hr with shaking at 225-250 rpm.
8. Spread on LB
agar plates containing appropriate antibiotic.
9. Incubate plates at 37°C
overnight (12-16 hrs).
6/26/09
Pick a colony from a streaked selective
plate to inoculate 10 ml of LB medium supplemented with the appropriate selection
antibiotic. Incubate 12-18 hrs at 37°C while shaking at 200-250 rpm.
6/27/09
Harvest
bacterial culture by centrifuging at 8000 rpm (6800xg) in microcentrifuge for
2 min at RT. Decant supernatant and remove remaining medium.
Purification/Mini
prep
1. Resuspend pelleted cells in 250 ul Resuspension Solution. Vortex or
pipet up and down until no cell clumps remain.
2. Add 250 Lysis Solution and
mix thoroughly by inverting tube 4-6 times until solution becomes viscous and
slightly clear. Do not incubate for more than 5 min to avoid denaturation of supercoiled
plasmid DNA.
3. Add 350 ul Neutralization Solution and mix thoroughly by inverting
tube 4-6 times.
4. Centrifuge for 5 min to pellet cell debris and chromosomal
DNA.
5. Transfer supernatant to spin column by decanting or pipetting. Avoid
disturbing or transferring white precipitate.
6. Centrifuge for 1 min. Discard
flow-through and replace column in collection tube.
7. Add 500 ul Wash Solution
to spin column. Centrifuge for 30-60s and discard flow-through.
8. Repeat step
7.
9. Centrifuge for 1 min to remove residual Wash Solution.
10. Transfer
spin column into fresh 1.7 ml microcentrifuge tube. Add 50 ul Elution Buffer to
center of spin column membrane. Do not touch pipette tip to membrane. Incubate
2 min at RT and centrifuge for 2 min.
Note: An additional elution step with
Elution buffer or water will recover residual DNA from the membrane and increase
overall yield by 10-20%.
11. Discard column and store purified plasmid DNA
at -20°C.
6/29/09
E: Restriction digests, Gel to check digests
Digest 1
NdeI (FastDigest) 1 ul
SpeI (NEB) 1 ul
10x FastDigest buffer
2 ul
plasmid 1 ul (~100 ng)
H2O 15 ul
20 ul
Band Sizes/Lengths: 5
kb, 3 kb, 460 bp, (80bp)
Digest 2 (Did not do)
NdeI (FastDigest) 2
ul
10x FastDigest buffer 2 ul
plasmid 1 ul (~100 ng)
H2O 15 ul
20
ul
Band Sizes/Lengths: 5.4 kb, 3 kb
Digest 3
BamHI (FastDigest)
1 ul
XhoI (FastDigest) 1 ul
10x FastDigest buffer 2 ul
plasmid 1 ul (~100
ng)
H2O 15 ul
20 ul
Band Sizes/Lengths: 6.6 kb, 1.8 kb
Digest
4
BamHI (FastDigest) 2 ul
10x FastDigest buffer 2 ul
plasmid 1 ul (~100
ng)
H2O 15 ul
20 ul
Band Size/Length: 8.4 kb
R: Correct bands
and sizes for plasmid 2 and digests 1, 3, and 4, correct band and size for plasmid
4
C: Do more tests on plasmid 2 such as sequencing to determine if plasmid
is carryover, Try different primers
|
Advisors

| Dr. Jingdong Tian
jtian(at)duke.edu
Duke BME Department & Duke IGSP
| 
| Dr. Lingchong You
you(at)duke.edu
Duke BME Department & Duke IGSP
| 
| Dr. Fan Yuan
fyuan(at)duke.edu Duke BME Department
|
Graduate Students

| Maggie Jiayuan Quan
jq7(at)duke.edu
Graduate Student
| 
| Faisal Reza
faisal.reza(at)duke.edu
Graduate Student
|
Students

| Andrew Ang
andrew.ang(at)duke.edu
Andrew Ang is a freshman at Duke, majoring in Biomedical Engineering. Apart from class and iGEM, he is involved in the Jazz Ensembles program and Asian Students Association at Duke. His hobbies include piano, saxophone, tennis, and squash. He is interested in molecular and synthetic biology, biomolecular engineering and medical research. He has previously worked as part of the MIT 2008 team, and he is excited to participate in iGEM again this year, and many more years to come.
|

| Kevin Chien
kevin.chien(at)duke.edu
|

| Yaoyao Fu
yf21(at)duke.edu
|

| Faith Kung
fk8(at)duke.edu
Faith Kung is a senior at Duke majoring in Biomedical Engineering with minors in Music and Biology. She enjoys working in a lab. Besides academics, her hobbies
include arts and crafts, dance, and figure skating. Also, she is actively involved in the IV Christian Fellowship. Faith is applying to PhD programs
in Biomedical Science and hopes to pursue a career in scientific research and
education. She is excited about attending the iGEM competition this year.
|

| Sahil Prasada
sahil.prasada(at)duke.edu
Sahil Prasada is a freshman at Duke. He plans to pursue medicine as a career. His interests lie in Detroit sports, tennis, and dancing. He is a member of the DBS Raas team on campus. He is currently in the Trinity School of the Arts and Sciences but is considering transferring to the Pratt School of Engineering. He hopes that the Detroit Lions may one day win the Superbowl. While waiting for this to occur, he will attend the iGEM competition and is looking forward to winning an award.
|

| Nicholas Tang
nicholas.tang(at)duke.edu
|

| Peter Zhu
peter.zhu(at)duke.edu
Peter Zhu Is a freshman at Duke University and a North Carolina local. Though he's not sure yet what to do with his life, he thinks Biomedical Engineering and pre-Law is looking pretty good. When he's not busy with the routines of life, he is listening to the Billboard Top 100, playing Chopin Preludes, searching for new places to eat, playing tennis, studying poker, and gaming Starcraft/DoTA. Peter is a regular at Bail Hai Mongolian Grill, Lime and Basil Vietnamese Pho, and Five Guy's Burgers---bacon cheeseburger with all toppings of course.
|
|












 "
"