Team:Groningen/Notebook/18 August 2009
From 2009.igem.org
(→Metal Accumulation) |
(→GVP Cluster) |
||
| Line 4: | Line 4: | ||
===GVP Cluster=== | ===GVP Cluster=== | ||
| + | |||
| + | '''Plates''' | ||
| + | |||
| + | Showed single colony growth on plates with J61002-Metal Promoter-GVP plasmids, and were stored in the fridge for future preculture growth. | ||
| + | |||
| + | :→ The plates with high and low concentration of transformed cells showed colonies in the expected ratio. On all plates 1 in 20 colonies was dark red, indication of ligation of the original high constitutive promoter back into the J61002 vector, or uncut plasmid which was transformed. | ||
| + | |||
| + | '''Over Night Cultures''' | ||
| + | |||
| + | :→ All cultures showed growth, and can be used for plasmid isolation. | ||
| + | |||
| + | :→ By growing colonies on medium with either amp. or kan. as a selection tool to find cultures with correct plasmid it was expected that only the tubes with kan. would show growth. Instead the transformed E.coli TOP10 cells seemed resistent to both antibiotics? | ||
| + | |||
| + | [[Image:Plasmid Miniprep Kit Flow Protocol.gif|thumb|150px| www.sigmaaldrich.com]] | ||
| + | |||
| + | '''Plasmid Purification''' | ||
| + | |||
| + | Plasmid isolation was performed on the cultures of E.coli TOP10 containing plasmids [http://partsregistry.org/wiki/index.php?title=Part:pSB3K3 pSB3K3] with [http://partsregistry.org/wiki/index.php?title=Part:BBa_J23100 high], [http://partsregistry.org/wiki/index.php?title=Part:BBa_J23106 medium] and [http://partsregistry.org/wiki/index.php?title=Part:BBa_J23109 low] constitutive promoters and GVP with the "Sygma-Aldrich™ [http://www.sigmaaldrich.com/life-science/molecular-biology/dna-and-rna-purification/plasmid-miniprep-kit.html GenElute™] Plasmid Miniprep Kit". | ||
| + | |||
| + | * From each tube 4mL of culture was collected in a 2.0mL cup, and the cells were pelleted by centrifugation for 1 min. at max. speed and the supernatant discarded. | ||
| + | * Plasmids were eluted with 30μL MQ and stored in the fridge | ||
| + | |||
| + | |||
| + | '''Concentration''' | ||
| + | |||
| + | {|cellpadding="2" cellspacing="1" border="4" | ||
| + | |'''Plasmid''' | ||
| + | |'''Conc. ng/μL''' | ||
| + | |'''260/280 | ||
| + | |'''260/230 ''' | ||
| + | |'''-20 box (michael | ||
| + | |'''Restriction Control''' | ||
| + | |- | ||
| + | |pSB3K3-BBa_J23100-GVP (amp.1) | ||
| + | |39.0 | ||
| + | |1.82 | ||
| + | |0.95 | ||
| + | |No | ||
| + | |Negative | ||
| + | |- | ||
| + | |pSB3K3-BBa_J23100-GVP (amp.2) | ||
| + | |23.5 | ||
| + | |2.11 | ||
| + | |1.51 | ||
| + | |No | ||
| + | |Negative | ||
| + | |- | ||
| + | |pSB3K3-BBa_J23100-GVP (kan.1) | ||
| + | |20.3 | ||
| + | |2.52 | ||
| + | |2.72 | ||
| + | |No | ||
| + | |Negative | ||
| + | |- | ||
| + | |pSB3K3-BBa_J23100-GVP (kan.2) | ||
| + | |23.0 | ||
| + | |2.17 | ||
| + | |2.56 | ||
| + | |No | ||
| + | |Negative | ||
| + | |- | ||
| + | |pSB3K3-BBa_J23109-GVP (amp.1) | ||
| + | |29.1 | ||
| + | |2.05 | ||
| + | |1.69 | ||
| + | |No | ||
| + | |Negative | ||
| + | |- | ||
| + | |pSB3K3-BBa_J23109-GVP (amp.2) | ||
| + | |40.6 | ||
| + | |1.92 | ||
| + | |1.16 | ||
| + | |No | ||
| + | |Negative | ||
| + | |- | ||
| + | |pSB3K3-BBa_J23109-GVP (kan.1) | ||
| + | |44.1 | ||
| + | |1.88 | ||
| + | |1.08 | ||
| + | |No | ||
| + | |Negative | ||
| + | |- | ||
| + | |pSB3K3-BBa_J23109-GVP (kan.2) | ||
| + | |24.6 | ||
| + | |2.26 | ||
| + | |2.22 | ||
| + | |No | ||
| + | |Negative | ||
| + | |} | ||
| + | |||
| + | |||
| + | '''Restriction Control''' | ||
| + | |||
| + | Isolated plasmids were cut with fast-digest enzymes EcoRI and PstI to cut out GVP, and create ~6000bp and ~2700bp fragments. | ||
| + | |||
| + | [[Image:Gel 18-8 no.1.jpg|400px]] [[Image:Generulers_1kb_marker_Fermentas.jpg]] | ||
| + | |||
| + | :→ From left to right: (2x) 1kB ladder, (2x) pSB3K3-J23100-GVP (amp.), (2x) pSB3K3-J23100-GVP (kan.), (2x) pSB3K3-J23109-GVP (amp.), (2x) pSB3K3-J23109-GVP (kan.) | ||
| + | |||
| + | :→ The restriction pattern was not expected (~6000bp and ~2700bp), and can be explained by ligation of vector pSB3K3 to J61035, which gives two EcoRI and one PstI site with ~2700bp, ~2000bp and ~1400bp fragments. The gel purification of GVP cluster must have gone wrong during the cutting of wanted fragment, and the vector instead of GVP has been isolated. | ||
| + | |||
| + | :→ During growth of cells the negative control plate with ampicillin also contained colonies, and can be explained by both amp. and kan. resistence on the double vector ligation product. | ||
| + | |||
| + | :→ New o.n. cultures for isolation of pSB3K3 plasmid with J23100, J23106, and J23109 promoters were made in 6mL LB-amp<sub>100</sub> medium. | ||
===Transporters=== | ===Transporters=== | ||
Revision as of 13:55, 18 August 2009
Wet
GVP Cluster
Plates
Showed single colony growth on plates with J61002-Metal Promoter-GVP plasmids, and were stored in the fridge for future preculture growth.
- → The plates with high and low concentration of transformed cells showed colonies in the expected ratio. On all plates 1 in 20 colonies was dark red, indication of ligation of the original high constitutive promoter back into the J61002 vector, or uncut plasmid which was transformed.
Over Night Cultures
- → All cultures showed growth, and can be used for plasmid isolation.
- → By growing colonies on medium with either amp. or kan. as a selection tool to find cultures with correct plasmid it was expected that only the tubes with kan. would show growth. Instead the transformed E.coli TOP10 cells seemed resistent to both antibiotics?
Plasmid Purification
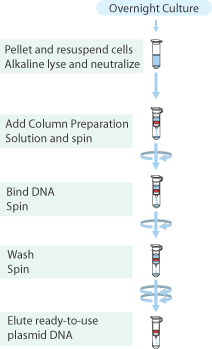
Plasmid isolation was performed on the cultures of E.coli TOP10 containing plasmids pSB3K3 with high, medium and low constitutive promoters and GVP with the "Sygma-Aldrich™ GenElute™ Plasmid Miniprep Kit".
- From each tube 4mL of culture was collected in a 2.0mL cup, and the cells were pelleted by centrifugation for 1 min. at max. speed and the supernatant discarded.
- Plasmids were eluted with 30μL MQ and stored in the fridge
Concentration
| Plasmid | Conc. ng/μL | 260/280 | 260/230 | -20 box (michael | Restriction Control |
| pSB3K3-BBa_J23100-GVP (amp.1) | 39.0 | 1.82 | 0.95 | No | Negative |
| pSB3K3-BBa_J23100-GVP (amp.2) | 23.5 | 2.11 | 1.51 | No | Negative |
| pSB3K3-BBa_J23100-GVP (kan.1) | 20.3 | 2.52 | 2.72 | No | Negative |
| pSB3K3-BBa_J23100-GVP (kan.2) | 23.0 | 2.17 | 2.56 | No | Negative |
| pSB3K3-BBa_J23109-GVP (amp.1) | 29.1 | 2.05 | 1.69 | No | Negative |
| pSB3K3-BBa_J23109-GVP (amp.2) | 40.6 | 1.92 | 1.16 | No | Negative |
| pSB3K3-BBa_J23109-GVP (kan.1) | 44.1 | 1.88 | 1.08 | No | Negative |
| pSB3K3-BBa_J23109-GVP (kan.2) | 24.6 | 2.26 | 2.22 | No | Negative |
Restriction Control
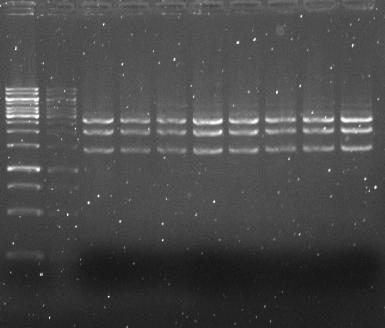
Isolated plasmids were cut with fast-digest enzymes EcoRI and PstI to cut out GVP, and create ~6000bp and ~2700bp fragments.
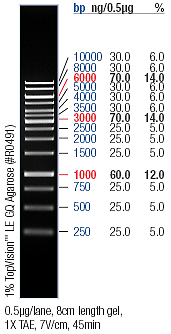
- → From left to right: (2x) 1kB ladder, (2x) pSB3K3-J23100-GVP (amp.), (2x) pSB3K3-J23100-GVP (kan.), (2x) pSB3K3-J23109-GVP (amp.), (2x) pSB3K3-J23109-GVP (kan.)
- → The restriction pattern was not expected (~6000bp and ~2700bp), and can be explained by ligation of vector pSB3K3 to J61035, which gives two EcoRI and one PstI site with ~2700bp, ~2000bp and ~1400bp fragments. The gel purification of GVP cluster must have gone wrong during the cutting of wanted fragment, and the vector instead of GVP has been isolated.
- → During growth of cells the negative control plate with ampicillin also contained colonies, and can be explained by both amp. and kan. resistence on the double vector ligation product.
- → New o.n. cultures for isolation of pSB3K3 plasmid with J23100, J23106, and J23109 promoters were made in 6mL LB-amp100 medium.
Transporters
HmtA curios about the dirty experiment. a 1% gel control will show if expected product is there. Unfortunately there was no product, the primer dimer changed and pcr 1 and 2 remain available. PCR again. Different program. Plus starting an other PCR1 and PCR2 to get new megaprimers.
Metal Accumulation
Restriction analysis MBP-ArsR
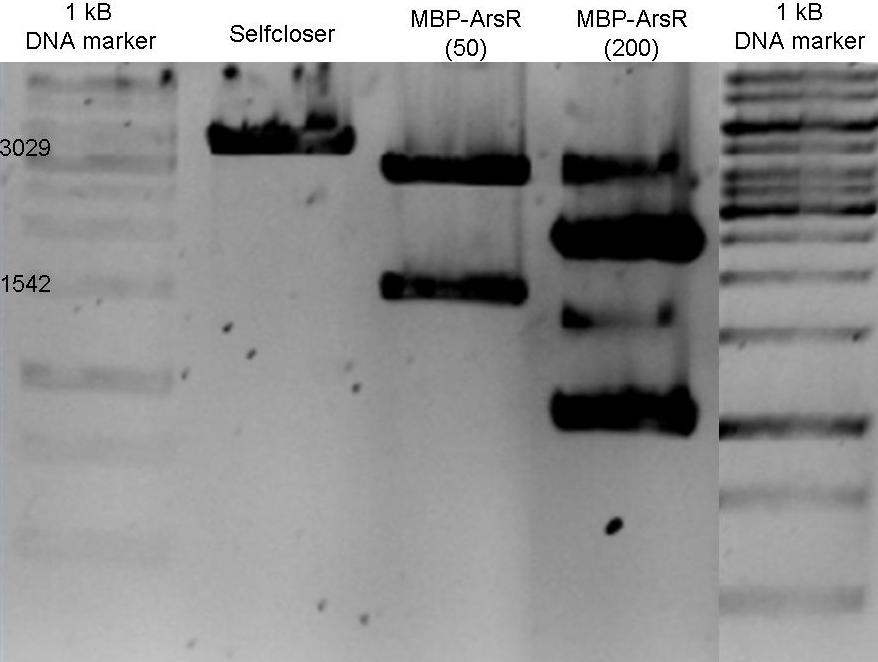
1% Agarose (1xTBE) gel with restriction analysis of MBP-ArsR fusion protein plasmids, with selfcloser (1), sample (white colony) (2), sample (red colony??) (3). Expected sizes of positive construct are shown. Samples were digested using PstI & XbaI, ~45 min 37 °C. Sample MBP-ArsR (200) was isolated from red organims, indicating the presence of RFP or similar, this was however not engineered in this plasmid. Therefore a deviant restriction pattern was to be expected. It appears as if two different plasmids were present in this sample.
Vectors
Restriction analysis metal promotors
Dry
|
|
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
|
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
 "
"