Team:Groningen/Notebook/28 September 2009
From 2009.igem.org
(Difference between revisions)
m (→GVP Cluster) |
(→GVP Cluster) |
||
| Line 21: | Line 21: | ||
:→ For GVP two fragments of ~6000bp and ~3000bp are expected, and for BBa_K190017 fragments of ~900bp and ~2000bp are expected. | :→ For GVP two fragments of ~6000bp and ~3000bp are expected, and for BBa_K190017 fragments of ~900bp and ~2000bp are expected. | ||
| + | ::* The sample of Steven-Jelle did not contain any DNA visible on gel! | ||
| + | ::* The GVP lanes showed the expected fragment sizes, with the addition of single cut vector at ~9000bp. The fragments were cut from the gel for purification. | ||
| + | ::* The restriction test to check SpeI/PstI confirmed the normal funtioning of the enzymes. This makes the RBS plasmid from -80 stock less attractive, because the unwanted RFP is not located behind the RBS to be cut out! | ||
| + | ::* To check another posibility, the stored plasmids from June were cut with SpeI/PstI and the stock plasmid was cut with EcoRI/XbaI to check if the RBS is placed infront of RBS. | ||
| + | |||
| + | |||
| + | [[Image:28-9 no.2.jpg|340px]] [[Image:Generulers_1kb_marker_Fermentas.jpg]] | ||
| + | |||
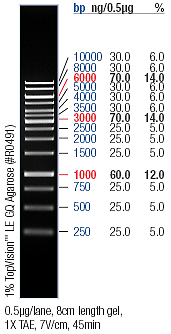
| + | :→ From left to right: 1kB ladder, RBS (EcoRI/XbaI), Empty Slot, RBS no.1, RBS no.2, RBS no.3 | ||
===Transporters=== | ===Transporters=== | ||
Revision as of 09:44, 28 September 2009
Wet
GVP Cluster
TODO Restrict GVP biobrick with XbaI/PstI and purify both vector and GVP fragment
TODO Restict GVP fragment and pMA-gvpL with MvaI/XhoI and purify 310bp. 2200bp, and 3300bp fragments
TODO Ligate the three fragments into the vector and transform E.coli DB3 cells
TODO Find out wath the problem is with RBS??
- Is it one of the restriction enzymes (SpeI/PstI) failing work
- Something wrong with biobrick in the sense that there is a red colour where none was expected (where did the RFP come from, and why is it activated so strongly)
- → From left to right: 1kB ladder, Sample Steven, SpeI/PstI control with BBa_K190017, Empty Slot, GVP (6x)
- → For GVP two fragments of ~6000bp and ~3000bp are expected, and for BBa_K190017 fragments of ~900bp and ~2000bp are expected.
- The sample of Steven-Jelle did not contain any DNA visible on gel!
- The GVP lanes showed the expected fragment sizes, with the addition of single cut vector at ~9000bp. The fragments were cut from the gel for purification.
- The restriction test to check SpeI/PstI confirmed the normal funtioning of the enzymes. This makes the RBS plasmid from -80 stock less attractive, because the unwanted RFP is not located behind the RBS to be cut out!
- To check another posibility, the stored plasmids from June were cut with SpeI/PstI and the stock plasmid was cut with EcoRI/XbaI to check if the RBS is placed infront of RBS.
- → From left to right: 1kB ladder, RBS (EcoRI/XbaI), Empty Slot, RBS no.1, RBS no.2, RBS no.3
Transporters
Metal Accumulation
Vectors
Dry
|
|
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
|
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
 "
"