From 2009.igem.org
Wet
GVP Cluster
- → DONE isolate plasmids from overnight precultures
- → DONE perform a restriction on isolated plasmid (should result in 6000 and 3000bp fragments)
- → TODO plate on cultures for pure colony over the weekend
- → TODO cut pSB3K3 plasmid and GVP containing plasmid for ligation
- → TODO purify wanted fragments
Plasmid Purification
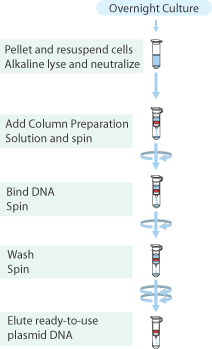
Plasmid isolation was performed on the cultures of E.coli TOP10 containing plasmids pSB1AC3 with high, medium and low constitutive promoters and GVP with the "Sygma-Aldrich™ GenElute™ Plasmid Miniprep Kit".
- From each tube 4mL of culture was collected in a 2.0mL cup, and the cells were pelleted by centrifugation for 1 min. at max. speed and the supernatant discarded.
- Plasmids were eluted with 20μL MQ and stored in the fridge
Concentration of Plasmids
| Plasmid
| Conc. ng/μL
| 260/280
| 260/230
| -20 box (michael
| Restriction Control
|
| pSB3K3-H
| 25.0
| ?
| ?
| ?
| ?
|
| pSB3K3-M
| 32.6
| ?
| ?
| ?
| ?
|
| pSB3K3-L
| 33.9
| ?
| ?
| ?
| ?
|
| GVP (biobrick)
| 484.0
| ?
| ?
| ?
| EcoRI/PstI
|
| pSB1AC3-H-GVP-amp
| 106.6
| 2.00
| 2.44
| ?
| EcoRI/PstI
|
| pSB1AC3-M-GVP-amp
| 165.2
| 1.92
| 2.40
| ?
| EcoRI/PstI
|
| pSB1AC3-L-GVP-amp
| 155.0
| 1.95
| 2.44
| ?
| EcoRI/PstI
|
| pSB1AC3-H-GVP-amp-conc.
| 116.3
| 1.96
| 2.43
| ?
| EcoRI/PstI
|
| pSB1AC3-M-GVP-amp-conc.
| 108.7
| 1.95
| 2.46
| ?
| ?
|
| pSB1AC3-L-GVP-amp-conc.
| 302.8
| 1.88
| 2.45
| ?
| ?
|
| pSB1AC3-H-GVP-amp/chl-conc.
| 257.1
| 1.92
| 2.41
| ?
| EcoRI/PstI
|
| pSB1AC3-M-GVP-amp/chl-conc.
| 224.7
| 1.94
| 2.37
| ?
| EcoRI/PstI
|
| pSB1AC3-L-GVP-amp/chl-conc.
| 166.1
| 1.95
| 2.44
| ?
| EcoRI/PstI
|
| pSB1AC3-L-GVP-amp/chl-oud
| 223.9
| 1.91
| 2.31
| ?
| EcoRI/PstI
|
Transporters
GlpF
The GlpF PCRs done yesterday (PCR 1 and2) were run on gel (lanes 2,3). Also the restriction done yesterday in order to check the ligations are shown on the gel below. The restriction was performed with HaeII for details see the notebook of 06-08-09.
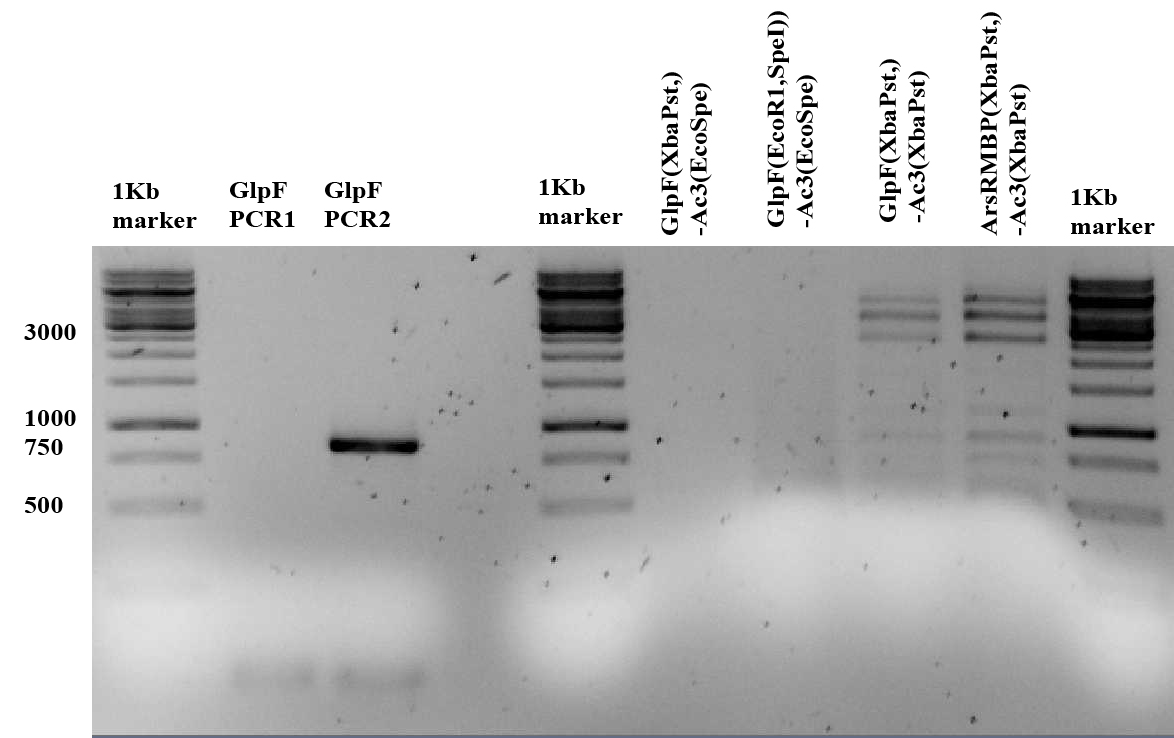
a band of 105 was expected for GlpF PCR1 which could not be seen on the above 1% agarose gel. Therefor 1uL of PCR1 (with 4 uL MQ and 1 uL 6x orange loading dye) was loaded to a 2% gel as shown below (arrow), since the amount of loaded DNA is low the band is not very dark.
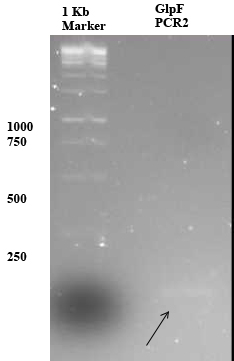
Metal Accumulation
- 2nd PCR to amplify SmtA and GST-SmtA
- Use pGEX-3X-GST-SmtA to amplify SmtA-GST and SmtA
- Use pET29a-SmtA to amplify SmtA
- Program: MT NIENKE (Left PCR machine)
- Globally: 10cycles touchdown from 65-50°C, 25 cycles with Tm gradient from 60-50°C (4steps), elongation time of 1:20.
- Mix
- MM NH4 21uL
- Primer fw [10mM] 1uL
- Primer rev [10mM] 1uL
- Vector 1uL
- HomeTaq pol 1uL
- Run program o/n
- Check on gel
- Transform E. coli with pGB68 (mymT)
- Use pGB68 from filter resuspended in 40uL MQ.
- As neg control use pSB1AC3 (incl ccdB), as pos control use pSB1AC3-low promoter.
- Mix 2-4uL of vector with 50uL chemically competent cells (prepared 31 aug)
- Transform using heat-shock @ 37°C
Vectors
Dry
 "
"