Team:HKU-HKBU/Polar Expression Results
From 2009.igem.org
(→Fluorescence Microscopy) |
|||
| (66 intermediate revisions not shown) | |||
| Line 3: | Line 3: | ||
{{Team:HKU-HKBU/header}} | {{Team:HKU-HKBU/header}} | ||
| - | = | + | =Strain Selection= |
| - | ===Swimming | + | ===Swimming Ability Assay=== |
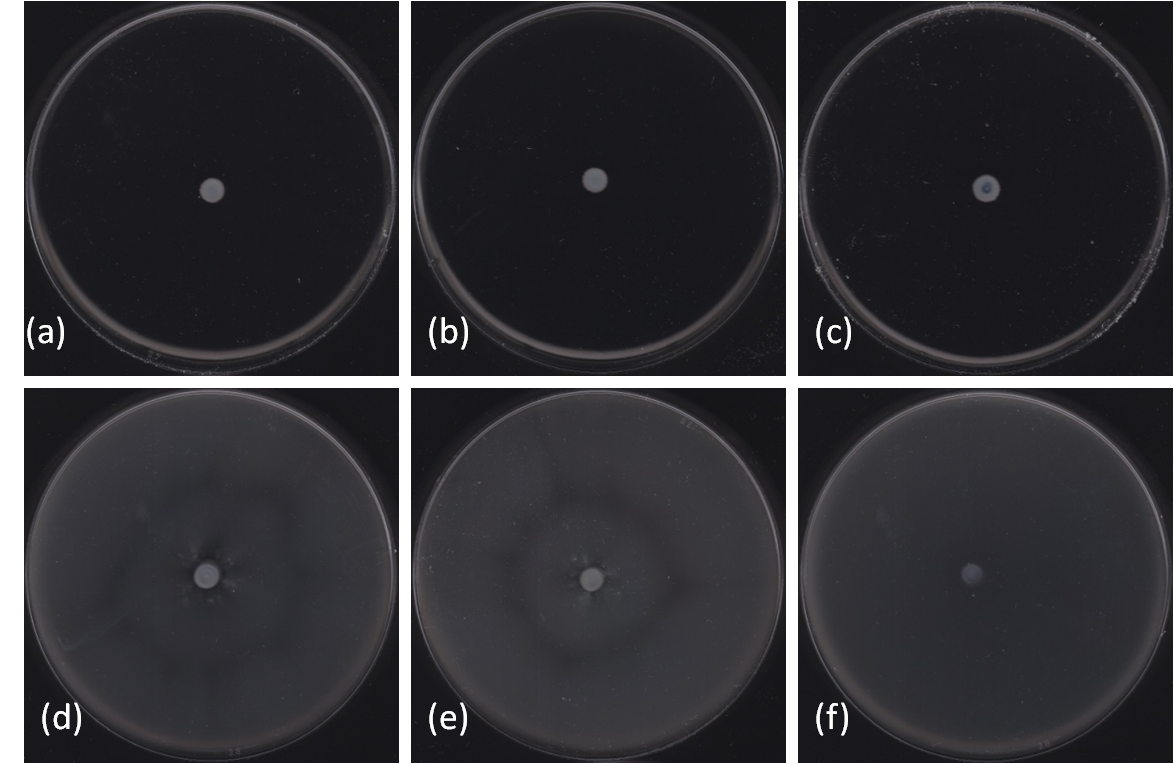
| - | The purpose of this part is to find one or | + | The purpose of this part is to find one or several strains of bacteria which could propel the motor efficiently. Swimming Ability Assay was used to identify the best bacterial strain. We tested several strains of ''Escherichia coli'' and ''Salmonella typhimurium''. We found that BL21, NCM3722 and YBE03 cannot swim whilst a ''Salmonella typhimurium'' strain - we named it YBS01 - is a bacteria with high swimming abilities (~4.5 mm/hr). ''Escherichia coli'', we named it YBE01, showed even more impressive performance, with a speed of ~5.5 mm/hr increase in radius at the end of eight-hour-experiment. |
| - | |||
| - | |||
| - | =''' | + | [[Image:HKU-BU-Swimming assay second3.png| center|thumb|300px|'''Figure 1.''' Swimming ability of different bacteria strains. '''a,''' BL21; '''b,''' NCM3722; '''c,''' YBE03; '''d,''' YBE01; '''e,''' YBS01; '''f,''' MG1655.]] |
| + | |||
| + | ===LPS Completeness Search=== | ||
| + | |||
| + | LPS takes vital part in AIDA polar expression system. Due to many mutations may exist in the strains of ''Escherichia coli'' or ''Salmonella typhimurium'', some of these mutations may cause the deficiency in the LPS layers. These mutants could survive in the culture. But for AIDA expression, when the microorganism's LPS is incomplete, the AIDA will express all over the bacteria surface; while the bacterial LPS is complete, the AIDA will express only on one side of the bacteria. After literature review, ''E. coli''-YBE01 [[#Reference | [1]]] and ''S.typhimurium''-YBS01 [[#Reference | [2]]] are identified to possess complete LPS layer. Their ability to express desirable proteins on the head is examined in later experiment. | ||
| + | |||
| + | =Polar Expression= | ||
| - | ==AIDA | + | ==AIDA Polar Expression System== |
| + | |||
| + | ===Fluorescence Microscopy=== | ||
| + | |||
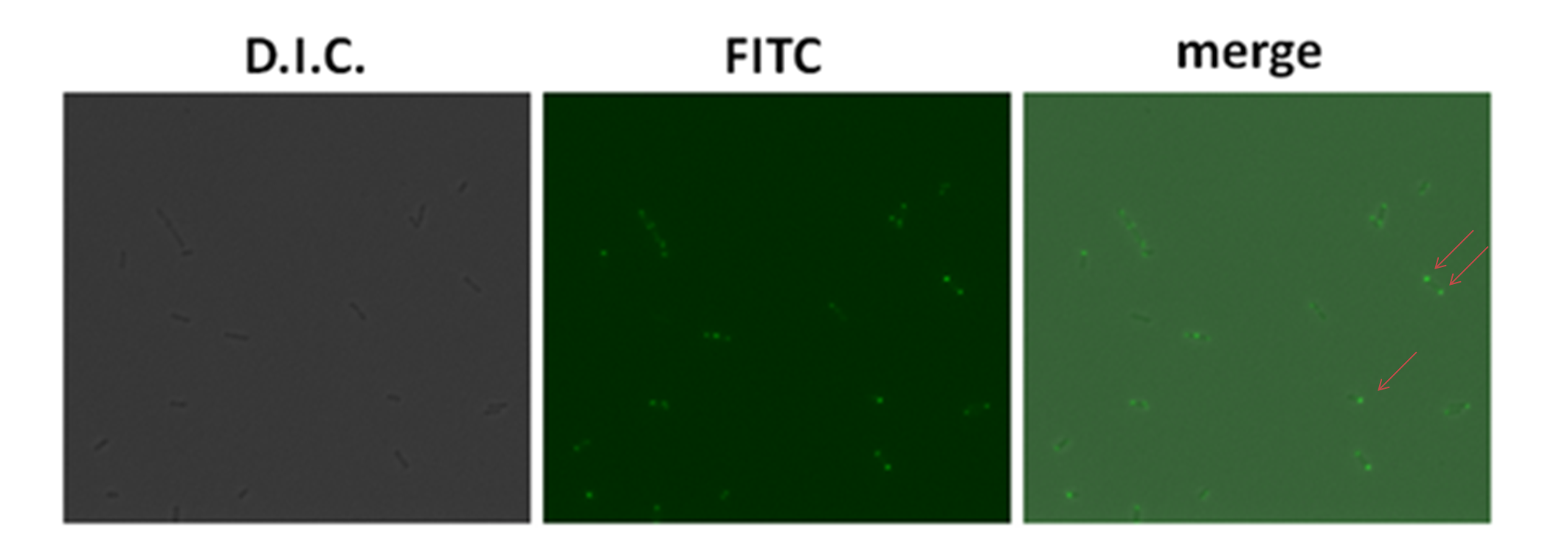
| + | After literature review, we found the ''E. coli'' BL21 was also an LPS complete strain. Therefore, we used the plasmid ([http://partsregistry.org/wiki/index.php?title=Part:BBa_K283001 '''BBa_K283001''']) containing T7 promoter which controlled the expression of '''GFP-Strp-AIDA''' to test the polar expression. BL21 strain also contains T7 polymerase which expression is induced by IPTG. When IPTG was added into the culture medium, '''GFP-Strp-AIDA''' would be strongly induced. In the figure below, it showes the expression of '''GFP'''. This expression was observed under the fluorescent microscope using oil lens with the magnification of 600 times. In the merged picture of dark field and FITC field, the fluorescent proteins were showed at one end of the bacteria. | ||
| + | |||
| - | |||
| - | |||
| - | [[Image:HKU- | + | [[Image:HKU-HKBU polar expression results E coli fluorescent3.png | center|thumb|300px|'''Figure 2.''' Polar Expression of GFP-Strp-AIDA in BL21. ''E.coli'' BL21 with plasmid pGFP-Strp-AIDA were induced by 0.5 mM IPTG for 2 h.]] |
| + | |||
| + | |||
| + | |||
| + | For this project, we required a strain that expresses specific proteins only in one side, and has swimming ability at the same time. However, the BL21 strain couldn't swim. Therefore we need to apply one side expression system into our new candidate YBE01. Unfortunately YBE01 does not contain T7 polymerase and is not able to activate the T7 promoter in the '''GFP-Strp-AIDA''' expressing plasmid ([http://partsregistry.org/wiki/index.php?title=Part:BBa_K283001 '''BBa_K283001''']). Therefore, a noval plasmid with strongly inducible or constitutive expression system was constructed. Attempts were made to use '''PLac''' without '''LacI''' as a strong constitutive promoter. However, only weak and unstable expression of '''GFP''' could be observed. One of the possible explanation could be that these systems were not strong or stable enough to ensure the consistent expression of our required proteins. | ||
| + | |||
| + | ===SDS-PAGE and Western Blot=== | ||
| + | |||
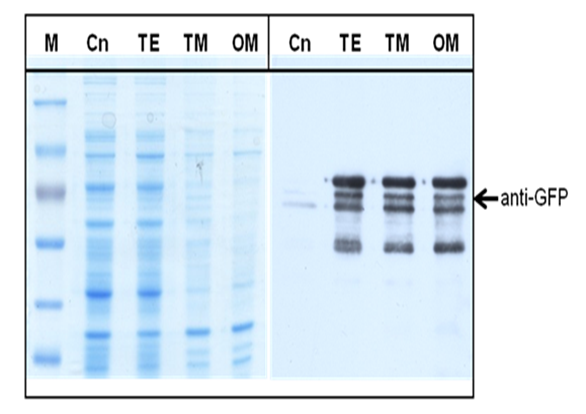
| + | The surface expression of streptavidin was essential for the specific binding of the bacteria to biotin-coated motor. To prove that the '''GFP-strp''' was expressed on the outer membrane of the bacteria through''' AIDA''' system, different contents of the bacteria were separated by ultracentrifuge. After separation, the outer membrane, inner membrane, total membrane and cytoplasm were collected as different samples. SDS-PAGE and Western Blot were conducted for verification. Each sample was quantified by BCA assay to make sure the equal loading of proteins. The western blotting results showed that most '''GFP''' proteins were localized on the outer membrane of the bacteria, which corresponded with the expected result that the specific expressions of proteins were on the outer membrane. | ||
| + | |||
| + | |||
| + | |||
| + | [[Image:HKU-BU-AIDA-west3.png| center|thumb|300px|'''Figure 3.''' SDS-PAGE and Western Blot of GFP-Strp-AIDA in BL21. Bacteria were induced by 0.5mM IPTG for 2 h; '''Cn,''' 0h control; '''TE,''' total cell extract; '''TM,''' total membrane; '''OM,''' outer membrane.]] | ||
| + | |||
| + | |||
| + | |||
| + | Observing the moving strains in the microscope, we found that all the '''GFP''' were expressed at the forehead end of the rod shaped bacteria. Therefore, this system was a perfect choice for binding with biotin-coated motor. It can express streptavidin at pole(s), which allows cells to adhere in the same orientation to a microrotatory motor through biotin-streptavidin interaction. | ||
| - | + | =='''Lpp-OmpA System'''== | |
| - | === | + | ===Fluorescence Microscopy=== |
| - | + | For AIDA system, the requirements for strains were critical, such as LPS completeness, strong expression, etc. Another polar expression system was constructed as our back-up plan. '''Lpp''' consists of 9 amino acids of signal peptide for targeting the inner membrane of the bacteria. '''OmpA (46-159)''' is a 5-cross trans-membrane protein which could bring cargos to the outer membrane of the bacteria. We constructed a plasmid ([http://partsregistry.org/wiki/index.php?title=Part:BBa_K283000 '''BBa_K283000''']) by fusing GFP-strp with '''Lpp-OmpA''' system. | |
| - | + | The surface expression of '''Lpp-ompA-GFP-Strp''' was consistent; however the polar expression of GFP-strp may not be achieved. Therefore, this plasmid was transformed into various strains of ''E.coli'' and ''S.typhimurium''. We finally found that in the stationary phase of YBS01, polar expression of the GFP-Strp was observed. | |
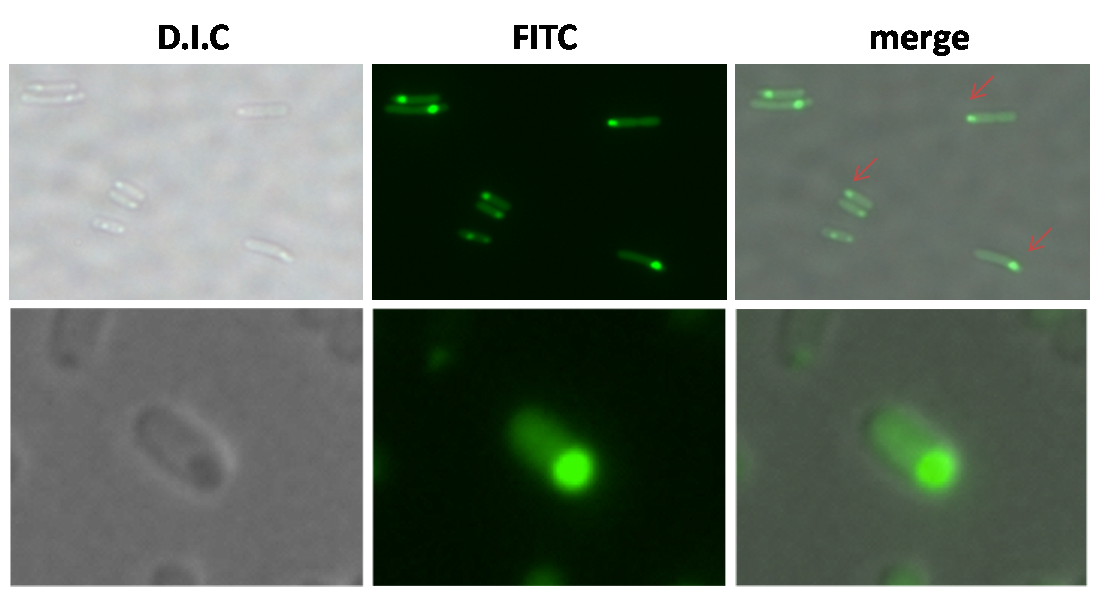
| - | + | In the figure below showed the polar expression of the plasmid ([http://partsregistry.org/wiki/index.php?title=Part:BBa_K283000 '''BBa_K283000''']) in the stationary phase of ''YBS01''. The right panel showed the merge picture of the bright field and FITC field under the fluorescence microscope with a magnification of 600 times. The result indicated that polar expression of proteins was achieved by this system in YBS01. | |
| - | |||
| - | + | [[Image:Polar.png| center|thumb|300px|'''Figure 4.''' Polar Expression of Lpp-ompA-GFP-Strp in YBS01. Bacteria were harvested through over-night culture.]] | |
| - | + | ===Western Blot=== | |
| + | To verify our experimental result in a more convincing way, Western Blot was conducted, which could reflect the surface expression of proteins. By studying the relative concentrations of expressed protein GFP in different samples, an obvious darker band was observed in the outer membrane samples. These results verified our hypothesis that the GFP proteins are particularly expressed on the outer membrane. | ||
| - | |||
| - | |||
| - | [[Image:HKU- | + | [[Image:HKU-HKBU polar expression results Salmonella western4.png| center|thumb|300px|'''Figure 5.''' Western Blot of Lpp-OmpA-GFP-Strp in YBS01. Bacteria were harvested through over-night culture ; '''OM,''' outer membrane; '''TM,''' total membrane; '''IM,''' inter membrane.]] |
| - | === | + | =='''Conclusion'''== |
| - | + | ||
| - | [ | + | After we tried different strains and polar expression systems, we decided to use bacteria YBS01 with Lpp-ompA system ([http://partsregistry.org/wiki/index.php?title=Part:BBa_K283000 '''BBa_K283000''']) to continue our project. |
| - | ==Reference== | + | =='''Reference'''== |
| - | # Sumita Jain, Peter van Ulsen, Inga Benz, M. Alexander Schmidt, Rachel Fernandez, Jan Tommassen, and Marcia B. Goldberg, Polar Localization of the Autotransporter Family of Large Bacterial Virulence Proteins, Journal of Bacteriology, | + | # Sumita Jain, Peter van Ulsen, Inga Benz, M. Alexander Schmidt, Rachel Fernandez, Jan Tommassen, and Marcia B. Goldberg, Polar Localization of the Autotransporter Family of Large Bacterial Virulence Proteins, ''Journal of Bacteriology'',2006, 188(13):4841-4850 |
| - | # Maurien M. A. Olsthoorn, Bent O. Petersen, Siegfried Schlecht, Johan Haverkamp, Klaus Bock, Jane E. Thomas-Oates and Otto Holst, Identification of a Novel Core Type in Salmonella Lipopolysaccharide, The Journal of Biological Chemistry, | + | # Maurien M. A. Olsthoorn, Bent O. Petersen, Siegfried Schlecht, Johan Haverkamp, Klaus Bock, Jane E. Thomas-Oates and Otto Holst, Identification of a Novel Core Type in Salmonella Lipopolysaccharide, ''The Journal of Biological Chemistry'', 1998, 273(7):3817-3829 |
{{Team:HKU-HKBU/footer}} | {{Team:HKU-HKBU/footer}} | ||
Latest revision as of 03:06, 22 October 2009
Contents |
Strain Selection
Swimming Ability Assay
The purpose of this part is to find one or several strains of bacteria which could propel the motor efficiently. Swimming Ability Assay was used to identify the best bacterial strain. We tested several strains of Escherichia coli and Salmonella typhimurium. We found that BL21, NCM3722 and YBE03 cannot swim whilst a Salmonella typhimurium strain - we named it YBS01 - is a bacteria with high swimming abilities (~4.5 mm/hr). Escherichia coli, we named it YBE01, showed even more impressive performance, with a speed of ~5.5 mm/hr increase in radius at the end of eight-hour-experiment.
LPS Completeness Search
LPS takes vital part in AIDA polar expression system. Due to many mutations may exist in the strains of Escherichia coli or Salmonella typhimurium, some of these mutations may cause the deficiency in the LPS layers. These mutants could survive in the culture. But for AIDA expression, when the microorganism's LPS is incomplete, the AIDA will express all over the bacteria surface; while the bacterial LPS is complete, the AIDA will express only on one side of the bacteria. After literature review, E. coli-YBE01 [1] and S.typhimurium-YBS01 [2] are identified to possess complete LPS layer. Their ability to express desirable proteins on the head is examined in later experiment.
Polar Expression
AIDA Polar Expression System
Fluorescence Microscopy
After literature review, we found the E. coli BL21 was also an LPS complete strain. Therefore, we used the plasmid ([http://partsregistry.org/wiki/index.php?title=Part:BBa_K283001 BBa_K283001]) containing T7 promoter which controlled the expression of GFP-Strp-AIDA to test the polar expression. BL21 strain also contains T7 polymerase which expression is induced by IPTG. When IPTG was added into the culture medium, GFP-Strp-AIDA would be strongly induced. In the figure below, it showes the expression of GFP. This expression was observed under the fluorescent microscope using oil lens with the magnification of 600 times. In the merged picture of dark field and FITC field, the fluorescent proteins were showed at one end of the bacteria.
For this project, we required a strain that expresses specific proteins only in one side, and has swimming ability at the same time. However, the BL21 strain couldn't swim. Therefore we need to apply one side expression system into our new candidate YBE01. Unfortunately YBE01 does not contain T7 polymerase and is not able to activate the T7 promoter in the GFP-Strp-AIDA expressing plasmid ([http://partsregistry.org/wiki/index.php?title=Part:BBa_K283001 BBa_K283001]). Therefore, a noval plasmid with strongly inducible or constitutive expression system was constructed. Attempts were made to use PLac without LacI as a strong constitutive promoter. However, only weak and unstable expression of GFP could be observed. One of the possible explanation could be that these systems were not strong or stable enough to ensure the consistent expression of our required proteins.
SDS-PAGE and Western Blot
The surface expression of streptavidin was essential for the specific binding of the bacteria to biotin-coated motor. To prove that the GFP-strp was expressed on the outer membrane of the bacteria through AIDA system, different contents of the bacteria were separated by ultracentrifuge. After separation, the outer membrane, inner membrane, total membrane and cytoplasm were collected as different samples. SDS-PAGE and Western Blot were conducted for verification. Each sample was quantified by BCA assay to make sure the equal loading of proteins. The western blotting results showed that most GFP proteins were localized on the outer membrane of the bacteria, which corresponded with the expected result that the specific expressions of proteins were on the outer membrane.
Observing the moving strains in the microscope, we found that all the GFP were expressed at the forehead end of the rod shaped bacteria. Therefore, this system was a perfect choice for binding with biotin-coated motor. It can express streptavidin at pole(s), which allows cells to adhere in the same orientation to a microrotatory motor through biotin-streptavidin interaction.
Lpp-OmpA System
Fluorescence Microscopy
For AIDA system, the requirements for strains were critical, such as LPS completeness, strong expression, etc. Another polar expression system was constructed as our back-up plan. Lpp consists of 9 amino acids of signal peptide for targeting the inner membrane of the bacteria. OmpA (46-159) is a 5-cross trans-membrane protein which could bring cargos to the outer membrane of the bacteria. We constructed a plasmid ([http://partsregistry.org/wiki/index.php?title=Part:BBa_K283000 BBa_K283000]) by fusing GFP-strp with Lpp-OmpA system.
The surface expression of Lpp-ompA-GFP-Strp was consistent; however the polar expression of GFP-strp may not be achieved. Therefore, this plasmid was transformed into various strains of E.coli and S.typhimurium. We finally found that in the stationary phase of YBS01, polar expression of the GFP-Strp was observed.
In the figure below showed the polar expression of the plasmid ([http://partsregistry.org/wiki/index.php?title=Part:BBa_K283000 BBa_K283000]) in the stationary phase of YBS01. The right panel showed the merge picture of the bright field and FITC field under the fluorescence microscope with a magnification of 600 times. The result indicated that polar expression of proteins was achieved by this system in YBS01.
Western Blot
To verify our experimental result in a more convincing way, Western Blot was conducted, which could reflect the surface expression of proteins. By studying the relative concentrations of expressed protein GFP in different samples, an obvious darker band was observed in the outer membrane samples. These results verified our hypothesis that the GFP proteins are particularly expressed on the outer membrane.
Conclusion
After we tried different strains and polar expression systems, we decided to use bacteria YBS01 with Lpp-ompA system ([http://partsregistry.org/wiki/index.php?title=Part:BBa_K283000 BBa_K283000]) to continue our project.
Reference
- Sumita Jain, Peter van Ulsen, Inga Benz, M. Alexander Schmidt, Rachel Fernandez, Jan Tommassen, and Marcia B. Goldberg, Polar Localization of the Autotransporter Family of Large Bacterial Virulence Proteins, Journal of Bacteriology,2006, 188(13):4841-4850
- Maurien M. A. Olsthoorn, Bent O. Petersen, Siegfried Schlecht, Johan Haverkamp, Klaus Bock, Jane E. Thomas-Oates and Otto Holst, Identification of a Novel Core Type in Salmonella Lipopolysaccharide, The Journal of Biological Chemistry, 1998, 273(7):3817-3829
 "
"