Team:Illinois-Tools/Project
From 2009.igem.org

Overall project
Synthetic biology is the creation of new functions using existing existing biological systems. In order to assist synthetic biologists, the 2009 Illinois tools team is trying to create a web-based open source program that outputs a theoretical pathway to synthesize a desired chemical product given an input and an output compound. This network will meet several constraints such as maximal reaction rates and mass balance. The ideal network will consist of known reactions taken from the Kyoto Encyclopedia of Genes and Genomes (KEGG)database, which consists of well-studied organisms. The capabilites of our program will be exemplified by establishing a network for the synthesis of biofuels from various input compounds.
Our program is modular and has 4 major components. The first aspect is obtaining reactions from the KEGG database via KEGG API and determining the shortest paths from the starting compound to the ending compound. The second is putting in an input and output for which you want a theoretical pathway. From then on you use our program to add weights in the pathway and customize it as you would like. We then give you the top three results as well as a stoichmetric matrix and a graphical representation. We also show how it would look like in Biobrick format and check it for compatablity with the registry. Now that you have your theoretical pathways you can proceed to the lab to test it.
Project Details
Database
As mentioned above we used information from the KEGG database to make our metabolic pathway tool. In order to organize the information and be able to retrieve it as desired, we created our own My SQL database with the necessary components for our network. The Kyoto Encyclopedia for Genes and Genomes has his own application programming interface (API) that can be used to allow other software, such as our own, to interact with it.
Initially we thought that we would create our network using the API functions each time a request was submitted so that we would have the most up-to-date information. Upon testing that out we determined that it would take way too long for each query. This is due to the time it takes to access the database through the API as well as the limited function calls that are available via the API. To fix this problem, we created out own database that has all the information we need it. By doing this, we decreased runtime exponentially. We explored several options and began with sqlite3. This was sufficient for our purposes but in order to ensure robustness for future versions, we switched over to MySql just before launching our program.
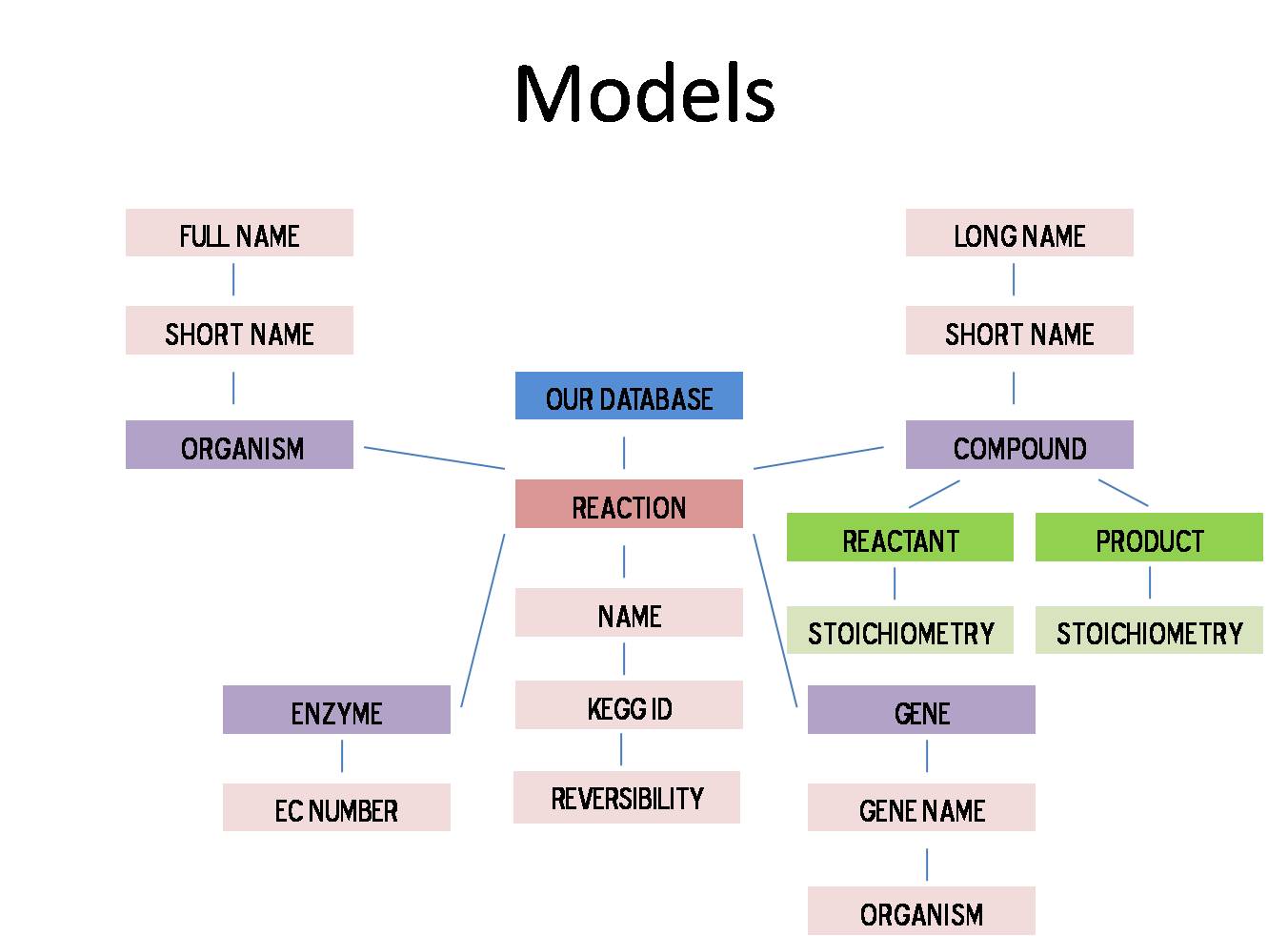
Using the Django web framework, creating a database was fairly straight forward but precise organization of the database was crucial to keeping runtime and update time to a minimum. After trying several different methods, we settled on creating a database centered on the reaction. Essentially all components can be reached through the reaction model. The picture below depicts the major parts of our database and how they are linked.
Although we decreased the time it takes for the program to run by creating our own database, there are disadvantages to this method. The biggest disadvantage is that our information will not be as up-to-date as KEGG. KEGG updates parts of their database monthly, other parts weekly, and still others monthly. We decided that in order to keep our program useful we will update our database to ensure that it matches KEGG every two weeks.
The Experiments
Part 3
Results
Future Plans
Specifications for the next version of IMP tools
The Illinois software tools team has explored into several new ideas that could potentially become projects for next summer’s team and other future teams. These ideas have focused on adding several new features that improve the present program and expand on what the current project accomplishes. There are several options that the team is currently considering. The user input and feedback that we receive will be taken into much consideration, and based on that, we will decide which changes should be implemented first. In order of importance, the main focus of expansion will be on incorporating the biobrick database, improving graphical representation of pathways, adding more constraints and improving the weighting system, and adding additional helpful features like displaying pricing information on compounds and biobricks.
One of the main areas of focus for the future teams will be the inclusion of biobricks and how to make use of the biobrick registry in a better way. This year’s team was able to convert the nucleotide sequence for the genes responsible for a particular pathway into standardized biobricks by cleaving restriction sites, and assigning appropriate prefixes and suffixes. Next year’s team could focus on creating better biobricks that could actually be tested out in an experimental lab. They could research into dividing the biobricks into genes that code for single proteins and enzymes, and ones that code for a series of enzymes that are used in the same pathway. Then, these biobricks can be put together on a single plasmid that contains the code for an entire pathway.
Another part of the project that can be improved is the graphical representation of the pathways generated. The pathway maps can be provided in a variety of different formats and programs. Some features that could be added are zoom features that could allow users to zoom into pathways and be able to see several alternate paths to a desired compound. This would involve being able recreate and put entire existing databases into one huge map. These interactive maps could be designed by members who are proficient at using software programs like Adobe Flash and Cytoscape.
Furthermore, next year’s team can improve the pathway generating algorithm by adding more constraints and developing a more efficient weighting system. Global computational studies can be used to evaluate immense numbers of possible input and output combinations. One new method of weighting can be added by assigning price values. This could be used to determine what the most economically valuable metabolic transformation is. Another improvement is picking the right host organism, which contains the highest fraction of the enzymes needed to conduct a transformation. Enzymes from other organisms can also be added to the initial host organism in order to add the desired functionality (considering from which sources the enzyme engineering challenge in the host is likely to be minimized).
Moreover, some other helpful features that can be added are assigning price values to all compounds and genes used, and then provide a feature that enables users to calculate the total expenses for ordering the compounds needed for a pathway, and also to order biobricks to carry out the pathway reactions in lab.
These are some of the additional features that could be added to enhance the project. Other ideas that will also be considered are; adding a tutorial that shows users how to create a biobrick if it does not already exist, and a way to favor reactions and pathways that have been genetically altered over time to create a specific product. Another tool that could be added is a flux balance analysis, to balance all the inputs and the outputs of the reaction. Thus, there are several options that are being looked into, for next summer’s team to work on, in order to enhance the current project.
== The End == "
"