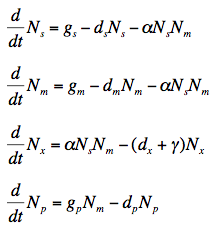Team:Illinois/Modeling
From 2009.igem.org
Modeling
sRNA regulation is a post-transcription regulation mechanism in which the mRNA transcript is prevented from being translated, either by RBS occlusion, degradation, or both. Traditionally, regulation via sRNA is faster acting than transcription factors because there is no delay due to translation and protein folding.[1] For this reason, sRNAs are often found to regulate stress and environmental responses. For example, the small RNA SgrS regulates PtsG, which codes for two protein subunits EIIB and EIIC which are involved in the glucose phosphate stress response. [2]
The rate for protein translation after sRNA regulation is governed by the following simple differential equations. N is the number of particles, g is the growth rate, d is the rate of decay or half life of a particle, and gamma is the rate of dissociation of the mRNA-sRNA complex. Subtext s, m, x, and p, denotes sRNA, mRNA, the bound sRNA-mRNA complex, and protein respectively.
The rate of change of the number of free sRNA molecules is found by taking the synthesis rate, subtracting the number of sRNA molecules multiplied by the decay rate, and subtracting the binding affinity to the mRNA multiplied by the number of mRNA molecules and the number of sRNA molecules. The process for finding the number of mRNA molecules is analogous.
Questions about our Wiki page? Please email Dave Korenchan at korench1@illinois.edu.
 "
"