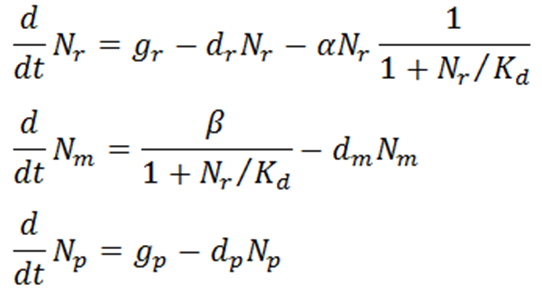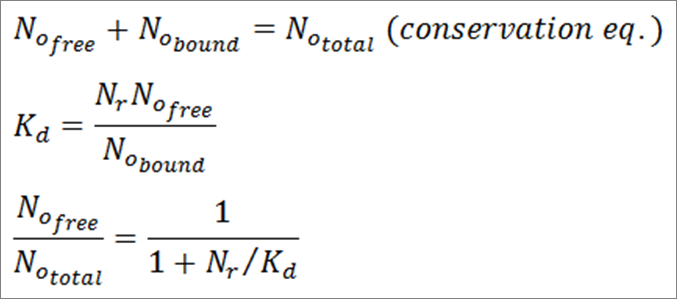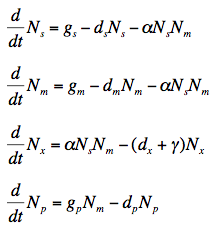Team:Illinois/Modeling
From 2009.igem.org
Modeling
This page includes mathematical descriptions and models of the biological processes involved in the function of our bacterial decoder.
Simple Transcriptional Regulation
Transcriptional regulation involves the binding of a regulator protein to a promoter and either activating or repressing transcription of the genes in the operon. Oftentimes, the regulator protein must be converted to an active state by binding to a signal molecule that changes the conformation of the protein and allows it to bind to the promoter. [3] In cases such as the Tet operon, the repressor protein TetR will bind to the operator site and repress transcription without binding to a signal molecule.
Transcriptional regulation by a repressor protein (such as TetR) can be described by the following ordinary differential equations
where subtexts r, m, and p represent repressor protein, mRNA, and output protein, respectively, Ni denote numbers of molecules for each species, gi denote synthesis rates, di denote degradation rates, α is the binding affinity of repressor protein to DNA, β is the maximal transcription rate of the promoter for the output protein, and Kd is the dissociation constant of repressor protein from DNA (in units of molecules). In this scenario, it is assumed that synthesis rates of repressor and output protein were constant and that there is only one DNA operator site for the repressor protein to bind to. Therefore, the third term of the first equation is a mass-action kinetics expression where the fraction part represents the probability that the operator site is free. This is also seen in the first term of the second equation, which is the maximal transcription rate β multiplied by the probability that the operator site is free. The probability is calculated by combining the first two equations below to get the third:
Here, No represents the number of operator sites in the cell (whether bound to repressor, free, or total).
Simple sRNA Regulation
sRNA regulation is a post-transcription regulation mechanism in which the mRNA transcript is prevented from being translated, either by RBS occlusion, degradation, or both. Traditionally, regulation via sRNA is faster acting than transcription factors because there is no delay due to translation and protein folding.[1] For this reason, sRNAs are often found to regulate stress and environmental responses. For example, the small RNA SgrS regulates PtsG, which codes for two protein subunits EIIB and EIIC which are involved in the glucose phosphate stress response. [2]
The rate for protein translation after sRNA regulation is governed by the following simple differential equations. N is the number of particles, g is the growth rate, d is the rate of decay or half life of a particle, and gamma is the rate of dissociation of the mRNA-sRNA complex. Subtext s, m, x, and p, denotes sRNA, mRNA, the bound sRNA-mRNA complex, and protein, respectively.
The rate of change of the number of free sRNA molecules is found by taking the synthesis rate, subtracting the number of sRNA molecules multiplied by the decay rate, and subtracting the binding affinity to the mRNA multiplied by the number of mRNA molecules and the number of sRNA molecules. The process for finding the number of mRNA molecules is analogous.
Analysis
Detailed information about how we created the models can be found on our Modeling Parameters page.
 "
"