Team:Illinois/Project
From 2009.igem.org
Abstract
Recently, synthetic biologists have been focusing on creating a logic circuit in bacteria by controlling gene expression at various levels: transcription, translation, and post-translational processes. Successfully installing a logic circuit in bacteria has large implications for engineering, and such a logic circuit could be used in applications involving drug development or biofuel fermentation. Various challenges have presented themselves in regard to this objective, such as the tendencies of outputs to be expressed at intermediate levels rather than being "on" or "off". While simple logic gates, such as AND, OR, NAND, and NOR gates, have been successfully created in bacteria, more complex logic circuits are still waiting to be designed.
Our team has chosen to create a two-to-four logic circuit in Escherichia coli. Our novel bacteria will be able to sense concentrations of two inputs (lactose and arabinose), identify these concentrations as "on" or "off", and produce one of four unique outputs (fluorescent proteins) depending on the combination of inputs. Thus, our four outputs can be represented in terms of our binary inputs as 00, 01, 10, and 11, where 0 corresponds to the "off" state (low concentration), 1 corresponds to the "on" state (high concentration), the first digit corresponds to the state of lactose, and the second digit corresponds to the state of arabinose.
One feature of our logic gate will be that it will be modular. In other words, it will be very simple to change the two inputs that our logic gate will respond to. Our design uses only two promoters that are each sensitive to a specific input. Swapping inputs simply involves using two different promoters sensitive to two different inputs in the proper locations. This ensures that our logic circuit does not just work with one specific scenario but that it will be flexible and have a wide variety of applications.
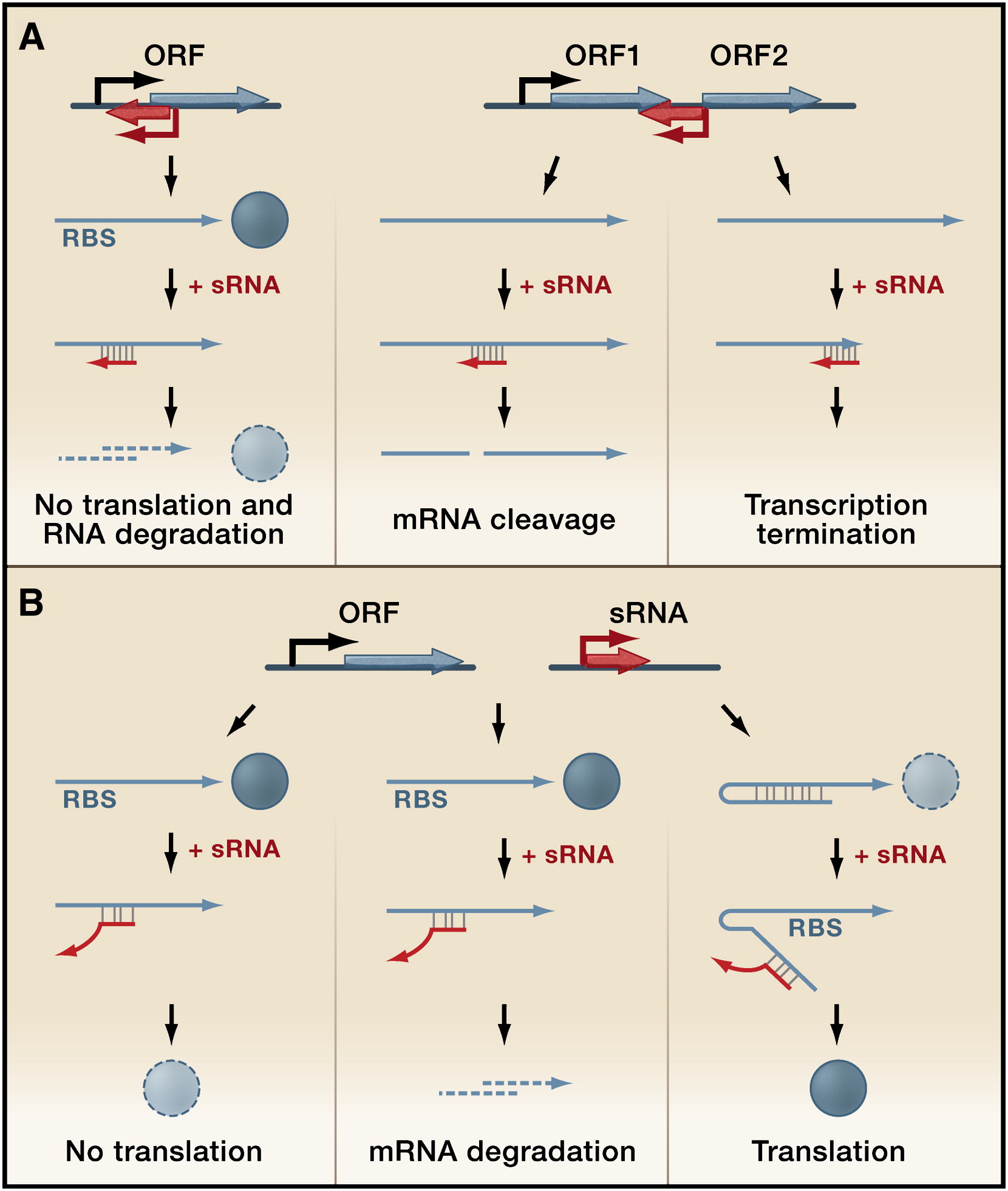
Our logic gate will also involve the usage of small RNAs, or sRNAs. sRNAs are very short transcripts of RNA (usually ~100 base pairs in E. coli) that are used in bacteria to positively or negatively regulate genes at the translational level by binding to the mRNA of the gene and either occluding the ribosomal binding site (RBS) and preventing ribosome binding or recruiting nucleases to degrade the mRNA. The roles and mechanisms of sRNAs in bacteria currently comprise a very prominent topic in the field of biology, and many sRNAs have been extensively studied and characterized. Using sRNAs in our logic circuit will help us achieve on/off behavior rather than intermediate levels of outputs, since sRNAs do little at low concentrations but demonstrate strong regulation at high concentrations. They also offer other advantages over transcriptional regulation; they use fewer cell resources, and they turn genes on or off more quickly. Currently, there are very few working sRNAs in the Parts Registry. During the course of our project, we hope to test several sRNAs and add them to the Parts Registry as working components for the use of other iGEM teams.
Sources
Barry, Patrick. "Bacteria That Do Logic". Science News, Vol. 174 #10, pg. 20. 8 November 2008. http://www.sciencenews.org/view/generic/id/37724/title/Bacteria_that_do_logic.
Garren, Emma. "Logic Gates". GcatWiki. http://gcat.davidson.edu/GcatWiki/index.php/Logic_Gates_-_Emma_Garren.
Storz, Gisela and Waters, Lauren. "Regulatory RNAs in Bacteria". Cell Vol. 136, Issue 4, pg. 615-28. 20 Feb 2009.
Questions about our Wiki page? Please email Dave Korenchan at korench1@illinois.edu.
 "
"
