Team:MIT/Projects/Project1
From 2009.igem.org
(→Engineering the PCB Synthesis Pathway into Yeast) |
(→Engineering the PCB Synthesis Pathway into Yeast) |
||
| (25 intermediate revisions not shown) | |||
| Line 1: | Line 1: | ||
| + | [[Image:Bilibuddies_project1.png]] | ||
__NOTOC__ | __NOTOC__ | ||
| - | <center>[[Team:MIT|Home]]|[[Team:MIT/Projects|Projects]]|[[Team:MIT/Protocols|Protocols]]|[[Team:MIT/ | + | <center>[[Team:MIT|Home]] | [[Team:MIT/Projects|Projects]] | [[Team:MIT/Protocols|Protocols]] | [http://partsregistry.org/cgi/partsdb/pgroup.cgi?pgroup=iGEM2009&group=MIT Parts for Registry] | [[Team:MIT/References|References]]</center> |
| + | <hr> | ||
| + | |||
| + | |||
| + | |||
| + | |||
=Metabolic Engineering of PCB Synthesis in Yeast= | =Metabolic Engineering of PCB Synthesis in Yeast= | ||
| + | [[Image:PCB_factory.jpg|center]] | ||
| + | == Background == | ||
| - | + | Phycocyanobilin (PCB) is a chromophore essential for the physical interaction between two proteins, phytochrome PhyB and integrating factor PIF3. PCB has two conformations: active and inactive. In the inactive Pr conformation, PCB has the ability to absorb red light at a wavelength of 680 nm. Once PCB absorbs red light, it switches to the active Pfr conformation and enables PhyB to bind to PIF3. In the Pfr conformation, PCB has the ability to absorb near infrared light, at a wavelength of 720 nm. Once PCB absorbs the near infrared light, it switches back to the inactive Pr conformation and causes dissociation of PhyB from PIF3. Shown below is the structure of PCB and how it switches between conformations. | |
| - | <center>[[Image: | + | <center> [[Image:PCB_Structure.gif|350px]] [[Image:Pr_Pfr_Cartoon.png|170px]]</center> |
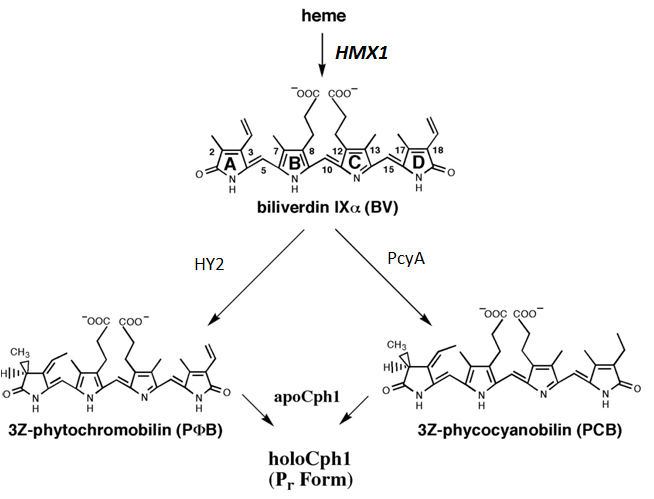
| - | PCB | + | The synthesis pathway of PCB begins with Heme to biliverdin (BV) then to PCB. Heme is produced in a wide range of organisms. Our main goal is to create a strain of yeast able to progress down the PCB synthesis pathway as shown below. |
| - | + | [[Image:PCB_Biosynthesis_Pathway.png|center|500px]] | |
| - | + | <center>''PCB biosynthesis pathway in Arabidopsis th. on the left and Synechocystis sp. on the right. Adapted from Gambetta, G. and Lagarias, JC., PNAS (2001)''</center> | |
| + | <br> | ||
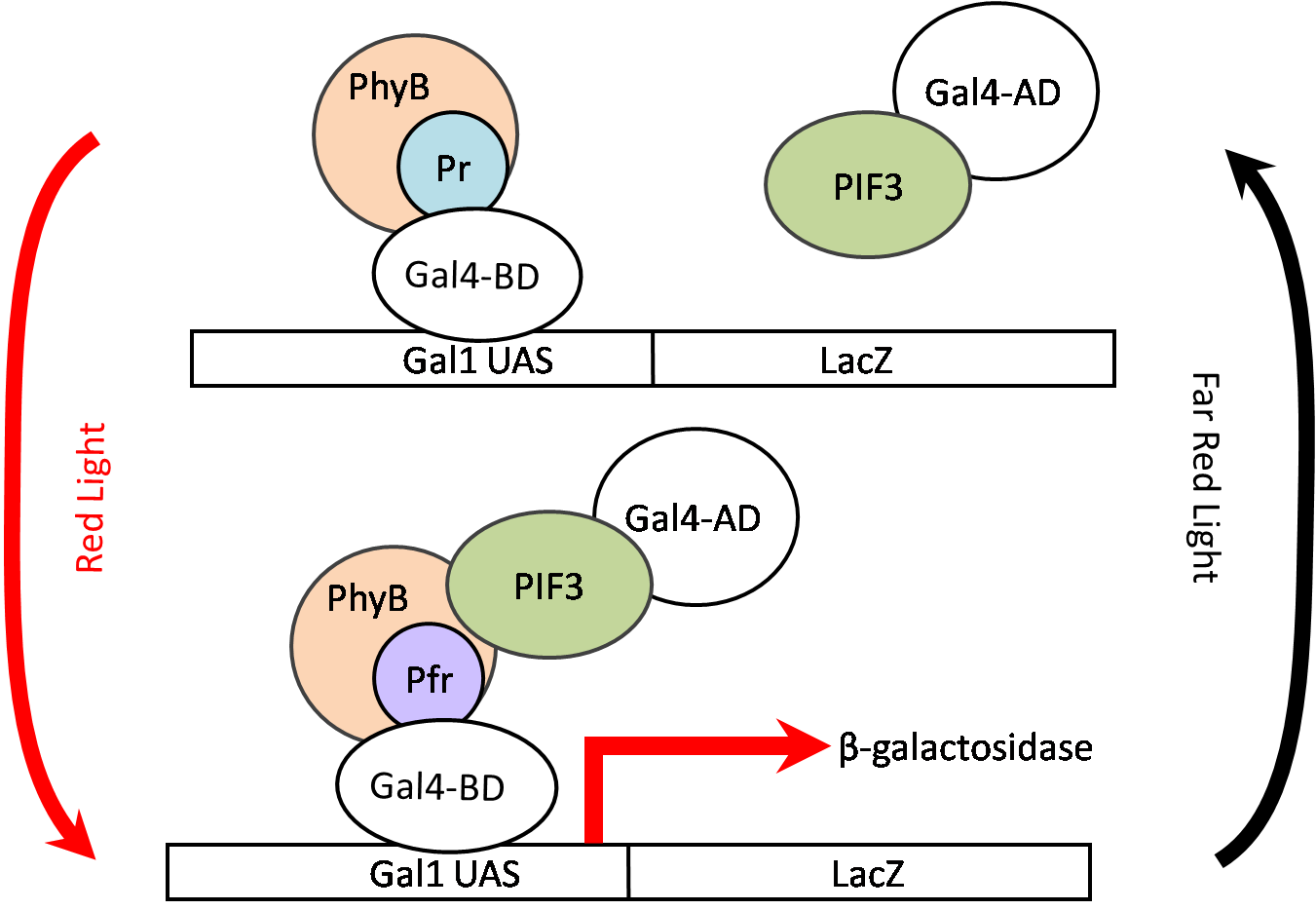
| + | In a 2002 ''Nature'' paper, the Quail Laboratory at University of California - Berekley used this PhyB-PIF3 system to induce ''LacZ'' expression. The GAL4 DNA-binding domain was fused to PhyB while the GAL4 activation domain was fused to PIF3. Once the system was pulsed with red light, ''LacZ'' expression was induced as shown below. | ||
| + | [[Image:PhyB_PIF3_LacZ_Expression.png|center|450px]] | ||
| - | + | <center>''Adapted from Shimizu-Sato, S., et al., Nature Biotechnology (2002)''</center> | |
| + | <br><br> | ||
| - | + | == Project Design == | |
| - | + | As mentioned above, PCB plays a crucial part in the PhyB-PIF3 system. Our goal is to make yeast synthesize PCB, so that it will not need to be supplemented in the medium. In order to confirm the production of PCB by our engineered yeast strain, we needed a sensitive, functional assay to detect the presence of PCB. We decided to use the two-hybrid system developed by the Quail lab, as illustrated below. The protocol for this functional assay can be found [http://openwetware.org/wiki/Functional_Assay_for_PCB here].<br> | |
| - | [[Image: | + | <center>[[Image:Donor-recipient.gif]]</center> |
| - | + | == Methods and Results == | |
| - | + | ===Developing the Standard: PCB from Spirulina=== | |
| - | + | We also purified PCB from a natural source so that we would have a standard for the PCB synthesized in yeast. | |
| + | Phycocyanobilin (PCB) extracted from ''Spirulina'' is a commonly used standard, as ''Spirulina'' produces a large amount of chromophores. We used ''Spirulina'' bought at Vitamin World, since ''Spirulina'' is commonly used as a dietary supplement. | ||
| - | [ | + | Typically, the chromophore has a very high absorbance around 680nm. A model of what the absorbance spectrum should look like is on the left, while the actual absorbance spectrum of PCB we extracted from ''Spirulina'' is on the right. The extraction protocol can be found at [http://openwetware.org/wiki/Extraction_of_PCB_from_Spirulina protocol that the Quail Lab]. |
| - | + | <center>[[Image:Theoretical_PCB_Absorbance.png|250px]] [[Image:Actual_PCB_Absorbance.png|250px]] </center> | |
| - | [[Image: | + | |
| - | |||
| - | + | We were able to get a concentration of 11.214 mM in a 4.5 mL solution. | |
| + | |||
| + | ===Engineering the PCB Synthesis Pathway into Yeast=== | ||
| - | |||
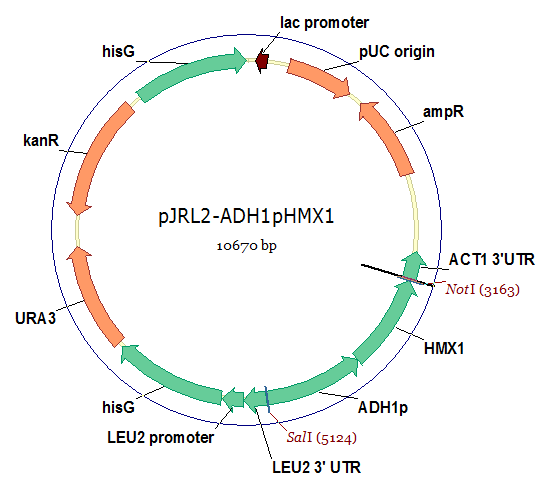
| - | ''HMX1'' is | + | The ''HMX1'' gene is already present in yeast. However, not enough of biliverdin is typically produced in yeast because ''HMX1'' is only transcribed under iron starvation conditions. Hence, we decided to clone ''HMX1'' from the genome and then overexpress it from a plasmid using the strong promoter from ''ADH1''. This plasmid is diagramed below. |
[[Image:HMX1_plasmid.png|center]] | [[Image:HMX1_plasmid.png|center]] | ||
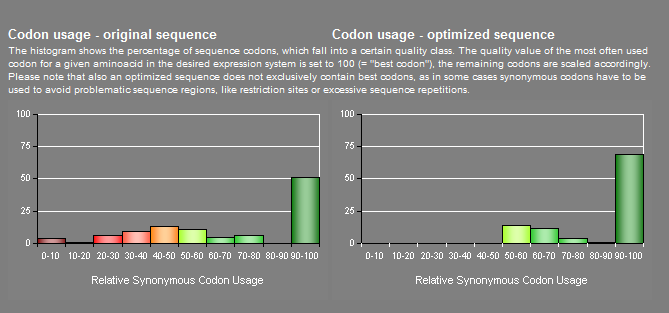
| - | + | For the synthesis of PCB from biliverdin, we decided to clone the ''PcyA'' gene from the ''Synechocystis sp.'' cDNA library. After looking at the codon usage charts for baker's yeast, we decided to codon optimize this gene to more effectively produce this enzyme in yeast. The codon usage comparisons of ''PcyA'' before and after codon optimization (done by GeneART), is seen below. | |
[[Image:PcyA_Usage.png|center]] | [[Image:PcyA_Usage.png|center]] | ||
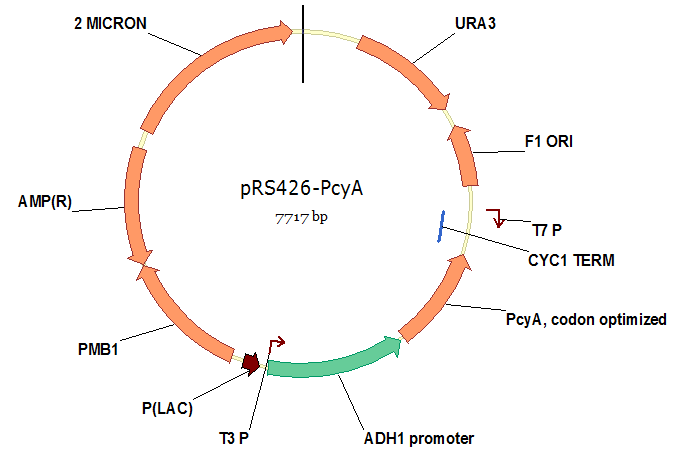
| - | We synthesized this optimized ''PcyA'' | + | We synthesized this optimized ''PcyA'' and cloned it into a vector, as shown below. |
[[Image:PcyA_plasmid.png|center]] | [[Image:PcyA_plasmid.png|center]] | ||
| - | + | Our next step is to transform both plasmids into a suitable yeast strain and employ the functional assay. By comparison with a known concentration of purified PCB and commercially available biliverdin, we will be able to assess the effectiveness of these enzymes in yeast. | |
Latest revision as of 03:51, 22 October 2009
Metabolic Engineering of PCB Synthesis in Yeast
Background
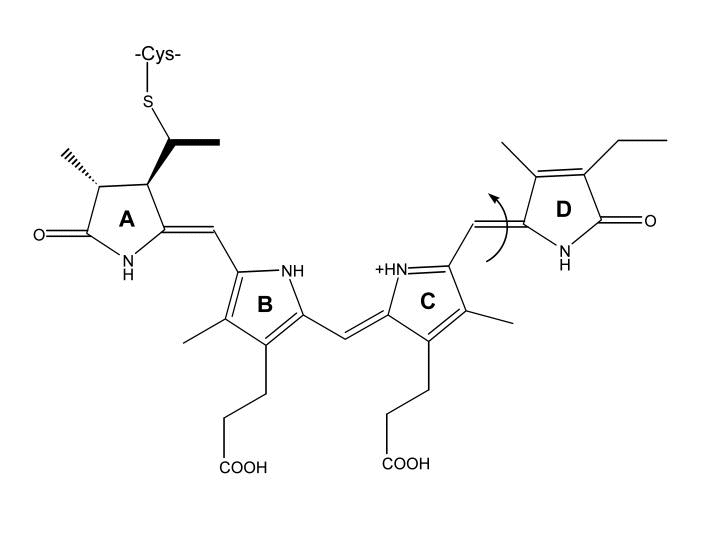
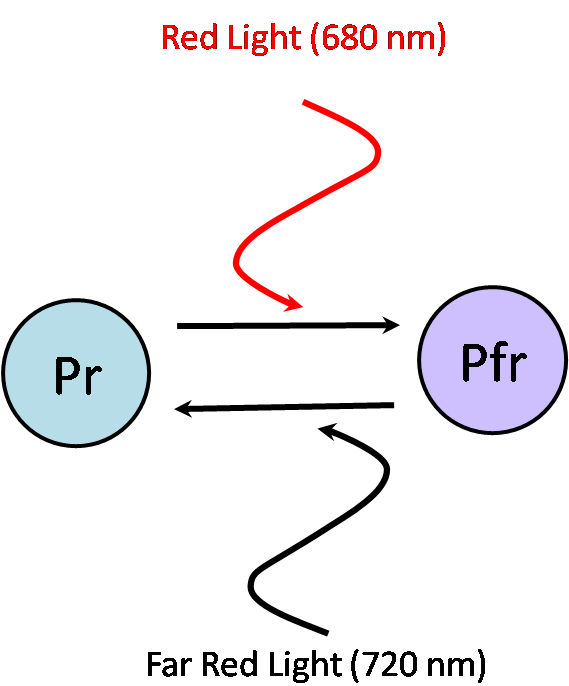
Phycocyanobilin (PCB) is a chromophore essential for the physical interaction between two proteins, phytochrome PhyB and integrating factor PIF3. PCB has two conformations: active and inactive. In the inactive Pr conformation, PCB has the ability to absorb red light at a wavelength of 680 nm. Once PCB absorbs red light, it switches to the active Pfr conformation and enables PhyB to bind to PIF3. In the Pfr conformation, PCB has the ability to absorb near infrared light, at a wavelength of 720 nm. Once PCB absorbs the near infrared light, it switches back to the inactive Pr conformation and causes dissociation of PhyB from PIF3. Shown below is the structure of PCB and how it switches between conformations.
The synthesis pathway of PCB begins with Heme to biliverdin (BV) then to PCB. Heme is produced in a wide range of organisms. Our main goal is to create a strain of yeast able to progress down the PCB synthesis pathway as shown below.
In a 2002 Nature paper, the Quail Laboratory at University of California - Berekley used this PhyB-PIF3 system to induce LacZ expression. The GAL4 DNA-binding domain was fused to PhyB while the GAL4 activation domain was fused to PIF3. Once the system was pulsed with red light, LacZ expression was induced as shown below.
Project Design
As mentioned above, PCB plays a crucial part in the PhyB-PIF3 system. Our goal is to make yeast synthesize PCB, so that it will not need to be supplemented in the medium. In order to confirm the production of PCB by our engineered yeast strain, we needed a sensitive, functional assay to detect the presence of PCB. We decided to use the two-hybrid system developed by the Quail lab, as illustrated below. The protocol for this functional assay can be found here.
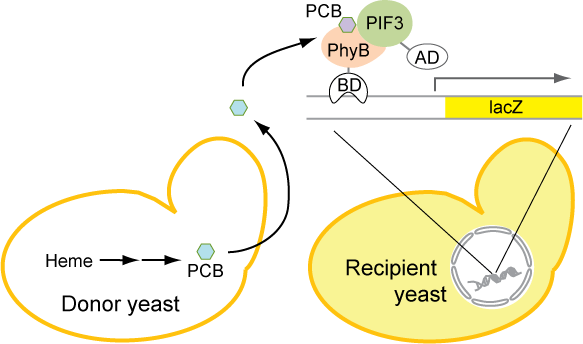
Methods and Results
Developing the Standard: PCB from Spirulina
We also purified PCB from a natural source so that we would have a standard for the PCB synthesized in yeast. Phycocyanobilin (PCB) extracted from Spirulina is a commonly used standard, as Spirulina produces a large amount of chromophores. We used Spirulina bought at Vitamin World, since Spirulina is commonly used as a dietary supplement.
Typically, the chromophore has a very high absorbance around 680nm. A model of what the absorbance spectrum should look like is on the left, while the actual absorbance spectrum of PCB we extracted from Spirulina is on the right. The extraction protocol can be found at protocol that the Quail Lab.
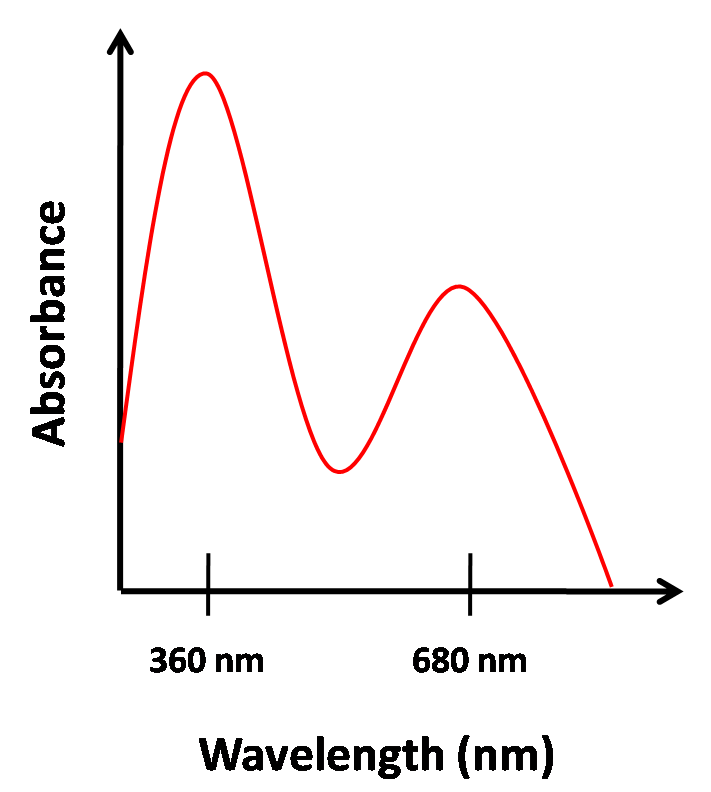
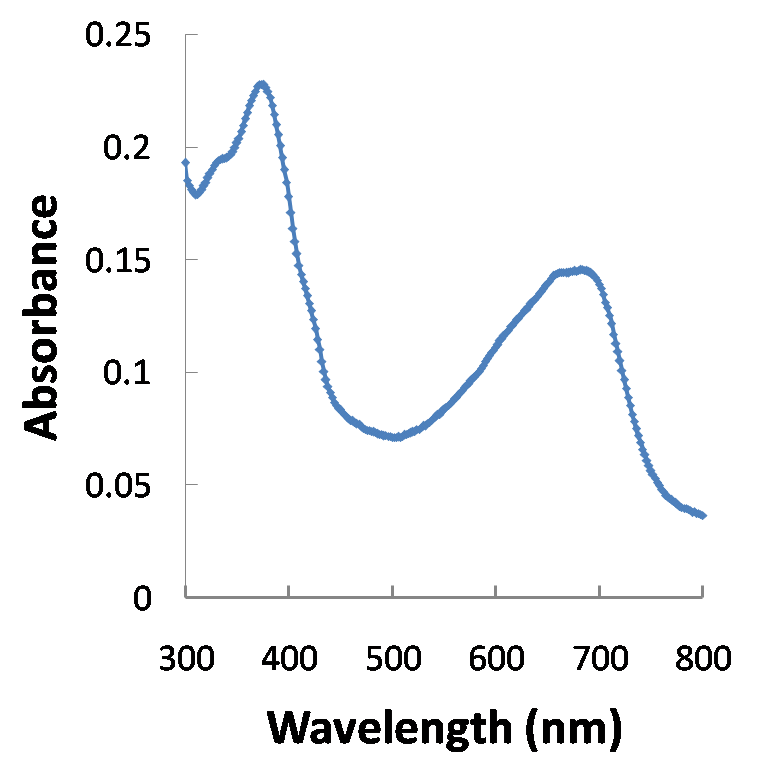
We were able to get a concentration of 11.214 mM in a 4.5 mL solution.
Engineering the PCB Synthesis Pathway into Yeast
The HMX1 gene is already present in yeast. However, not enough of biliverdin is typically produced in yeast because HMX1 is only transcribed under iron starvation conditions. Hence, we decided to clone HMX1 from the genome and then overexpress it from a plasmid using the strong promoter from ADH1. This plasmid is diagramed below.
For the synthesis of PCB from biliverdin, we decided to clone the PcyA gene from the Synechocystis sp. cDNA library. After looking at the codon usage charts for baker's yeast, we decided to codon optimize this gene to more effectively produce this enzyme in yeast. The codon usage comparisons of PcyA before and after codon optimization (done by GeneART), is seen below.
We synthesized this optimized PcyA and cloned it into a vector, as shown below.
Our next step is to transform both plasmids into a suitable yeast strain and employ the functional assay. By comparison with a known concentration of purified PCB and commercially available biliverdin, we will be able to assess the effectiveness of these enzymes in yeast.
 "
"