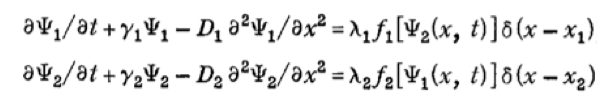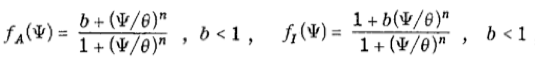Team:McGill/Modeling
From 2009.igem.org
(→Introduction) |
|||
| Line 63: | Line 63: | ||
This is modeled using the following system of PDEs: | This is modeled using the following system of PDEs: | ||
[[Image: Mcgill09PDEs.png|frame|center]] | [[Image: Mcgill09PDEs.png|frame|center]] | ||
| + | where Ψ1 and Ψ2 represent the concentrations of the activating and inhibiting molecules, respectively, γi the degradation constant, Di the diffusion constant, λi the maximal synthesis rate of molecule i, and δ the Dirac function. fi represents the Hill function describing the dependence on the opposing molecule: | ||
| + | [[Image: Mcgill09HillFunctions.png|frame|center]] | ||
Revision as of 05:41, 21 October 2009

Introduction
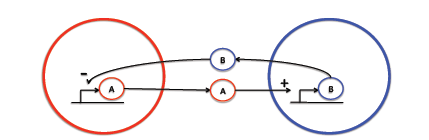
Many models examining intercellular signaling do not take into account the separation distances of the signaling bodies. We use a partial differential equation (PDE) based model to gain insight into spatially heterogeneous activation-inhibition intercellular signaling.
Two types of signaling molecules exist: activating and inhibiting. Each molecule is synthesized by a unique strain of cells and affects the synthesis rate of the other strain.
This is modeled using the following system of PDEs:
where Ψ1 and Ψ2 represent the concentrations of the activating and inhibiting molecules, respectively, γi the degradation constant, Di the diffusion constant, λi the maximal synthesis rate of molecule i, and δ the Dirac function. fi represents the Hill function describing the dependence on the opposing molecule:
 "
"