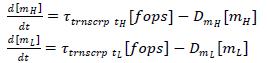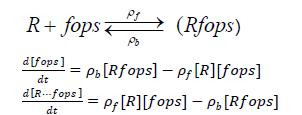Team:Michigan/Modeling
From 2009.igem.org
Charlieyan (Talk | contribs) |
Charlieyan (Talk | contribs) |
||
| Line 7: | Line 7: | ||
Cell Lysis happens when holin creates pores in membrane and lysozyme enters | Cell Lysis happens when holin creates pores in membrane and lysozyme enters | ||
Assume rate of cell death ∝[holin](# cells), rate of cell growth in exponential phase | Assume rate of cell death ∝[holin](# cells), rate of cell growth in exponential phase | ||
| - | + | [[Image:Equation1.jpg]] | |
== Transcription of Repressor: == | == Transcription of Repressor: == | ||
Revision as of 22:49, 20 October 2009
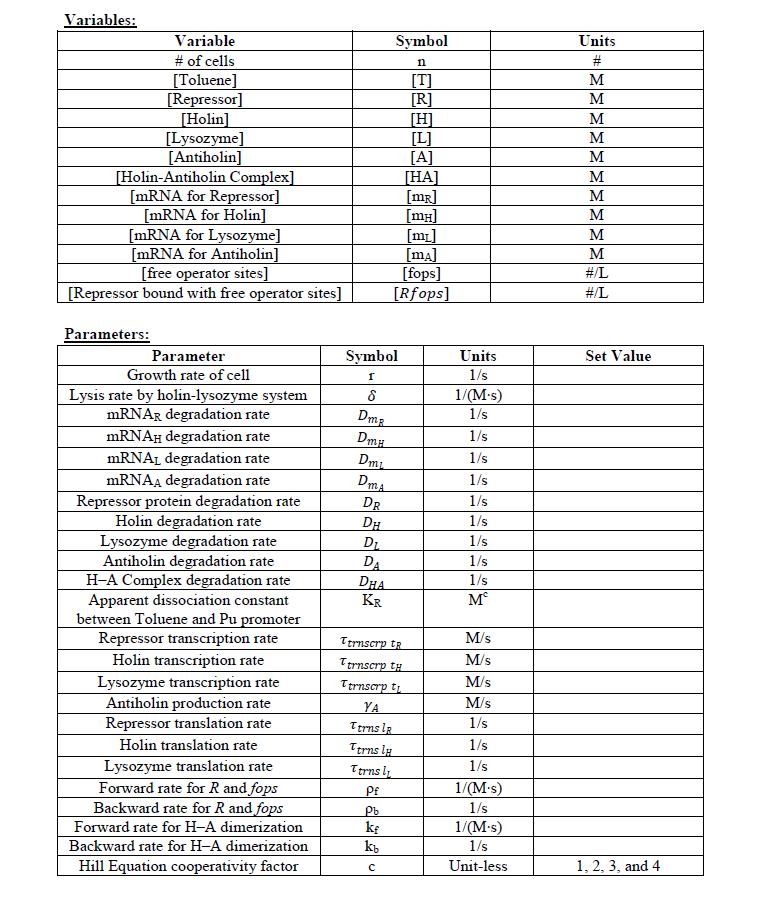
| ||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
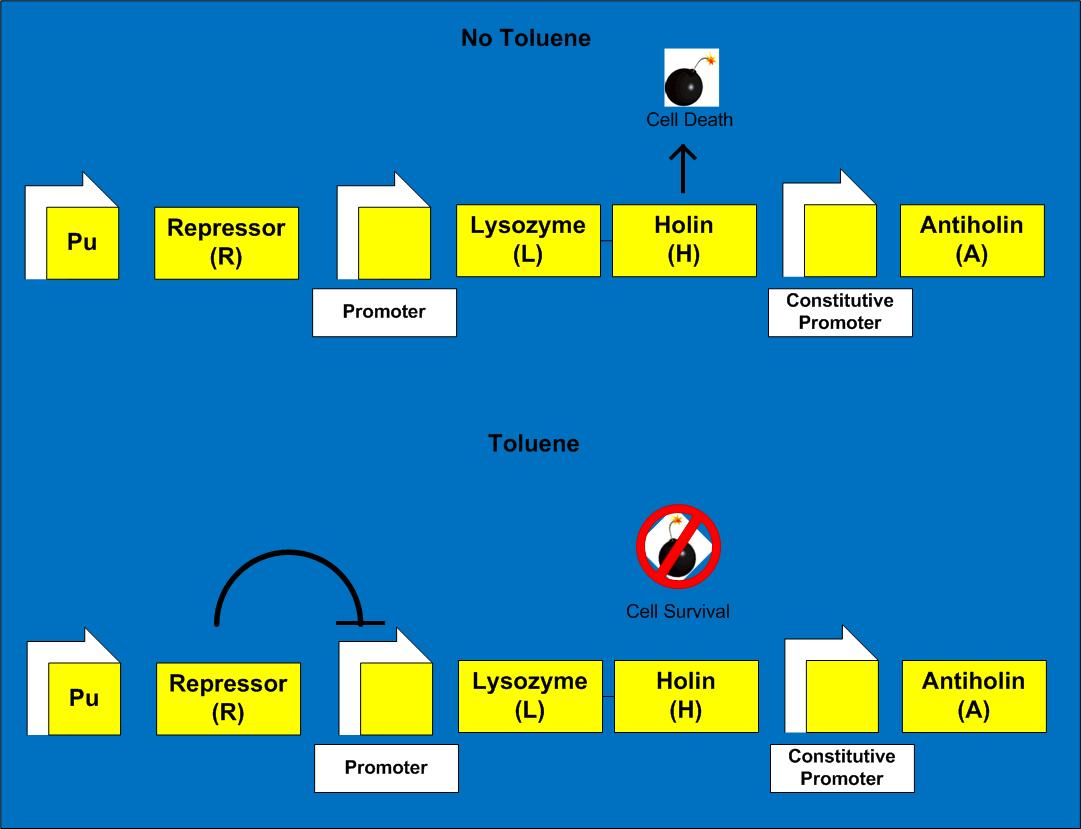
Cell Growth/Death
Cell Lysis happens when holin creates pores in membrane and lysozyme enters
Assume rate of cell death ∝[holin](# cells), rate of cell growth in exponential phase
![]()
Transcription of Repressor:
![]() 𝑓([𝑇]) is a function of toluene concentration
𝑓([𝑇]) is a function of toluene concentration
Use Hill Equation to describe binding affinity of toluene to Pu promoter.
![]()
Transcription of Lysozyme and Holin:
Assume transcription rate is proportional to # of free operator sites (which should be the same for both lysozyme and holin, since they are downstream of the same promoter)
Assume transcription rates for H and L are the same
Repression of Lysozyme and Holin Transcription
Repressor binds with free operator site, preventing transcription of lysozyme and holin. Use a model invoking law of mass action.
Assuming that fops and Rfops do not undergo spontaneous degradation.
Production of Repressor
Using the translation rate for R and taking into consideration the binding of R with fops,
Production of Proteins
Production of antiholin, under constitutive promoter, is at a constant rate γA, which depends on the promoter that is used
Dimerization:
Since holin and antiholin form a complex
![]()
Using the translation rates and incorporating dimerization using the law of mass action,
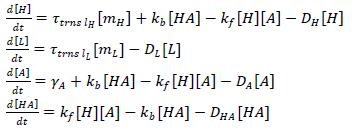
Assumptions
Can set γA equal to rate of production of holin in the case that all operating sites are free
- This is in order to balance antiholin and holin levels without repression
- This can be tuned so that timing of cell death works out
- Set antiholin production rate to that of holin in the absence of repression
- Put this through the transcription and translation equations to obtain production rate
Assume degradation rates of all mRNAs are the same
For protein degradation rate:
Case 1: Proteins are stable:
- Degradation rates equal reproduction rate of cell
- This is due to dilution of proteins across daughter cells
Case 2: Proteins are unstable
- If not other degradation rate information is provided, can possibly assume that the degradation rates are a few orders of magnitude above the reproduction rate of cell
- Will have to look into this further
 "
"