Team:MoWestern Davidson/project physicalmodel
From 2009.igem.org
Eric.sawyer (Talk | contribs) |
Eric.sawyer (Talk | contribs) |
||
| Line 2: | Line 2: | ||
= Frameshift Suppressor tRNA Physical Modeling = | = Frameshift Suppressor tRNA Physical Modeling = | ||
| - | == | + | == tRNA Structure == |
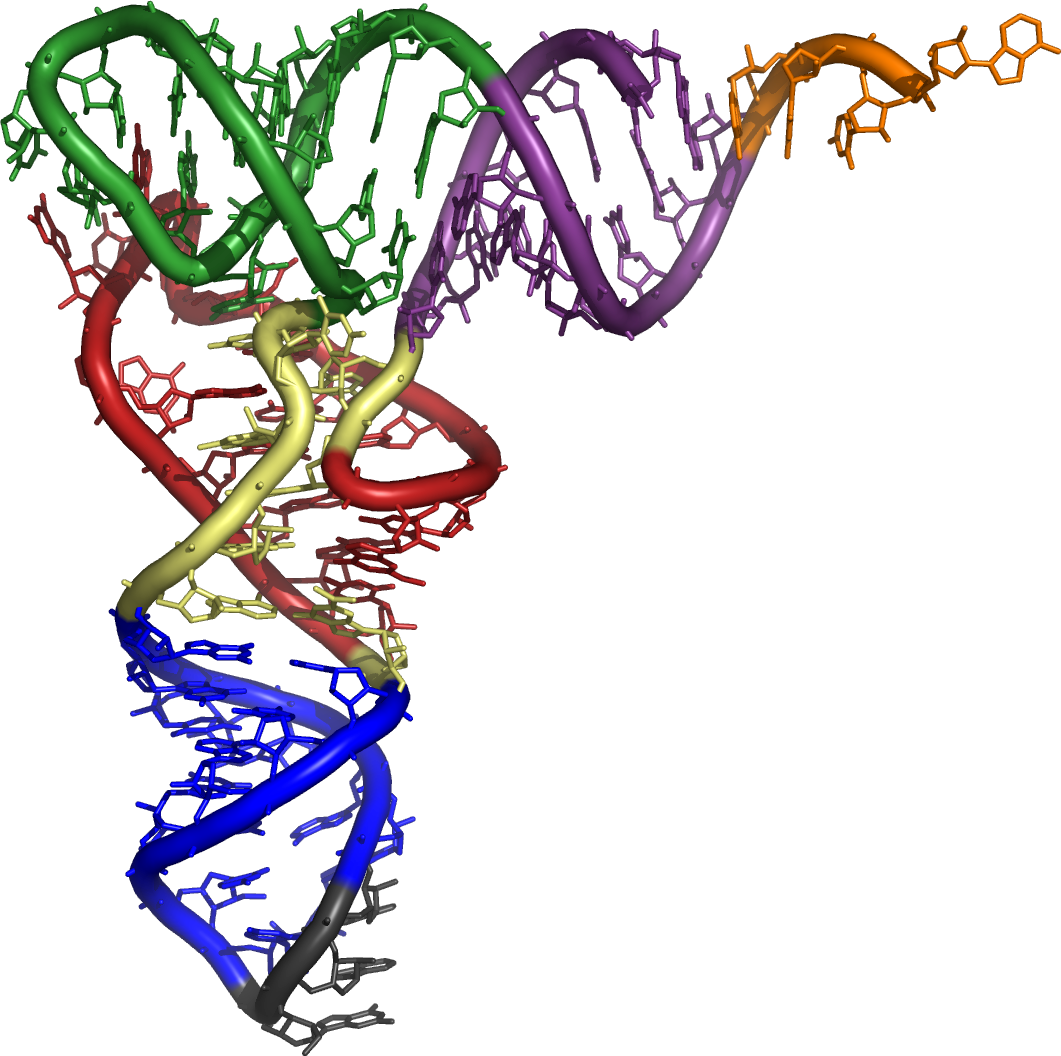
All tRNAs, whether they contain a 3- or 5-base anticodon, conform to a general structure. They feature a series of loops and stems folded into a tertiary L-shape, with the amino acid acceptor arm opposite the anticodon. The structure of tRNA molecules is important because their unique structural motifs are recognized by the ribosome and enzymatic pathways, and allow tRNAs to carry out their life-sustaining function. | All tRNAs, whether they contain a 3- or 5-base anticodon, conform to a general structure. They feature a series of loops and stems folded into a tertiary L-shape, with the amino acid acceptor arm opposite the anticodon. The structure of tRNA molecules is important because their unique structural motifs are recognized by the ribosome and enzymatic pathways, and allow tRNAs to carry out their life-sustaining function. | ||
[[Image:TRNAcloverleaf2.gif|right|thumb|150px|The cloverleaf model shows how the RNA strand base pairs with itself to form loops and stems, but does not accurately depict its structural motif.]] | [[Image:TRNAcloverleaf2.gif|right|thumb|150px|The cloverleaf model shows how the RNA strand base pairs with itself to form loops and stems, but does not accurately depict its structural motif.]] | ||
| Line 10: | Line 10: | ||
=== The Spacial tRNA Structural Motif === | === The Spacial tRNA Structural Motif === | ||
Base-pairing between arms stabilizes the structure of the tRNA molecule into a L-shape. The acceptor arm is positioned at one and of the L, and the anticodon at the other. The size and sequence of each arm influences the spacial structure of the tRNA, and allows enzymes responsible for maintaining function to recognize each tRNA. | Base-pairing between arms stabilizes the structure of the tRNA molecule into a L-shape. The acceptor arm is positioned at one and of the L, and the anticodon at the other. The size and sequence of each arm influences the spacial structure of the tRNA, and allows enzymes responsible for maintaining function to recognize each tRNA. | ||
| - | == | + | == Using the PDB Format to Design a Frameshift Suppressor tRNA == |
The PDB format is a standard file type used in molecular modeling and known structures to code for the Cartesian space coordinate triple for each atom within the molecule. It also includes portions describing which atoms combine to form nucleotides or amino acids, the locations of hydrogen bonds, and other information pertaining to the structure of molecules. A PDB file coding for a theorized model of a 5-base anticodon was made by modifying the anticodon loop of the existing PDB file 1EHZ—yeast phenylalanine tRNA. This particular tRNA was one of the first to undergo extensive study, and remains a model molecule among the countless tRNA molecules within the diverse variety of life. | The PDB format is a standard file type used in molecular modeling and known structures to code for the Cartesian space coordinate triple for each atom within the molecule. It also includes portions describing which atoms combine to form nucleotides or amino acids, the locations of hydrogen bonds, and other information pertaining to the structure of molecules. A PDB file coding for a theorized model of a 5-base anticodon was made by modifying the anticodon loop of the existing PDB file 1EHZ—yeast phenylalanine tRNA. This particular tRNA was one of the first to undergo extensive study, and remains a model molecule among the countless tRNA molecules within the diverse variety of life. | ||
| + | === The PDB Format Defined === | ||
| + | === Modifying a PDB File === | ||
== [[Team:MoWestern_Davidson/build model | Building a Physical Model]] == | == [[Team:MoWestern_Davidson/build model | Building a Physical Model]] == | ||
{{Template:MoWestern_Davidson2009_end}} | {{Template:MoWestern_Davidson2009_end}} | ||
Revision as of 21:05, 27 July 2009

Contents |
Frameshift Suppressor tRNA Physical Modeling
tRNA Structure
All tRNAs, whether they contain a 3- or 5-base anticodon, conform to a general structure. They feature a series of loops and stems folded into a tertiary L-shape, with the amino acid acceptor arm opposite the anticodon. The structure of tRNA molecules is important because their unique structural motifs are recognized by the ribosome and enzymatic pathways, and allow tRNAs to carry out their life-sustaining function.
The Cloverleaf Model
The structure of tRNA molecules is often represented by showing how stems and loops are formed by base pairing within a two-dimensional context—useful for depicting secondary structure, but unable to show three dimensional folding. Each tRNA molecule consists of five arms, and each, with the exception of the acceptor arm, contains both stems and loops. A stem is formed by self base-pairing along the RNA strand, and loops are formed from non-paired nucleotides. The 3′ terminus of the TRNA strand protrudes beyond the 5′ terminus by several nucleotides, but the 5′ terminus is base-paired. The appropriate amino acid is connected to the 3′ terminus by an amino acyl tRNA synthetase, an enzyme that uses the unique structural motif of each tRNA to consistently pair it with its designated amino acid. Moving from the 3′ terminus in the 5′ direction, the TΨC arm is encountered. It is so named because of the presence of the sequence 5′-TΨC-3′, where Ψ stands for the base pseudouracil, an isomer of uracil formed from post-transcriptional modification. Continuing in the 5′ direction, the variable loop is encountered. This portion of the tRNA family of molecules shows high size variation, from only a few nucleotides to well over a dozen. Next is the anticodon arm, which contains the anticodon itself. The anticodon recognizes its mRNA complement, and may contain a wobble base-pairing at the 5′ end residue of the anticodon. This wobble base-pairing often utilizes the abundant assortment of non-standard nucleotides. The final arm before the 5′ terminus is the D arm. It contains the D base, or dihydrouracil.
The Spacial tRNA Structural Motif
Base-pairing between arms stabilizes the structure of the tRNA molecule into a L-shape. The acceptor arm is positioned at one and of the L, and the anticodon at the other. The size and sequence of each arm influences the spacial structure of the tRNA, and allows enzymes responsible for maintaining function to recognize each tRNA.
Using the PDB Format to Design a Frameshift Suppressor tRNA
The PDB format is a standard file type used in molecular modeling and known structures to code for the Cartesian space coordinate triple for each atom within the molecule. It also includes portions describing which atoms combine to form nucleotides or amino acids, the locations of hydrogen bonds, and other information pertaining to the structure of molecules. A PDB file coding for a theorized model of a 5-base anticodon was made by modifying the anticodon loop of the existing PDB file 1EHZ—yeast phenylalanine tRNA. This particular tRNA was one of the first to undergo extensive study, and remains a model molecule among the countless tRNA molecules within the diverse variety of life.
The PDB Format Defined
Modifying a PDB File
Building a Physical Model
 "
"