Team:TUDelft/Modeling
From 2009.igem.org
(→Conjugation) |
(→Conjugation) |
||
| Line 15: | Line 15: | ||
We would like to give credit to the [https://2008.igem.org/Team:Bologna/Modeling Bologna '08 team] and [https://2008.igem.org/Team:UNIPV-Pavia/Modeling UNIPV-Pavia '08 team], who's project was useful in determining our own equations. | We would like to give credit to the [https://2008.igem.org/Team:Bologna/Modeling Bologna '08 team] and [https://2008.igem.org/Team:UNIPV-Pavia/Modeling UNIPV-Pavia '08 team], who's project was useful in determining our own equations. | ||
='''Conjugation'''= | ='''Conjugation'''= | ||
| - | Using the paper "[http://www3.interscience.wiley.com/journal/118837812/abstract?CRETRY=1&SRETRY=0 A model for bacterial conjugal gene transfer on solid surfaces]", we established the assumptions and | + | Using the paper "[http://www3.interscience.wiley.com/journal/118837812/abstract?CRETRY=1&SRETRY=0 A model for bacterial conjugal gene transfer on solid surfaces]", we established the assumptions and parameters that would be needed for the . |
| + | =='''Assumptions'''== | ||
| + | |||
| + | =='''Parameters'''== | ||
| + | *Surface area of media (A) | ||
| + | *Initial colony radius (r_0) | ||
| + | *Specific growth rate (g_n) | ||
| + | *Colony radial growth rate (g_r) | ||
| + | *Maximum numbers of cells sustained by system (N_max) | ||
| + | *Initial number of donors (N_d) and recipients (N_r) | ||
| + | All these parameters can be determined experimentally, or by literary research into previous conjugation experiments. | ||
| + | =='''Spatial distribution of bacteria'''== | ||
| + | |||
| + | =='''Growth of colonies'''== | ||
| + | |||
| + | =='''Contact between colonies'''== | ||
| + | |||
| + | =='''Conjugation modeling'''== | ||
| + | |||
| + | =='''Final cell number'''== | ||
{{Template:TUDelftiGEM2009_end}} | {{Template:TUDelftiGEM2009_end}} | ||
Revision as of 10:02, 22 July 2009
Delay Device
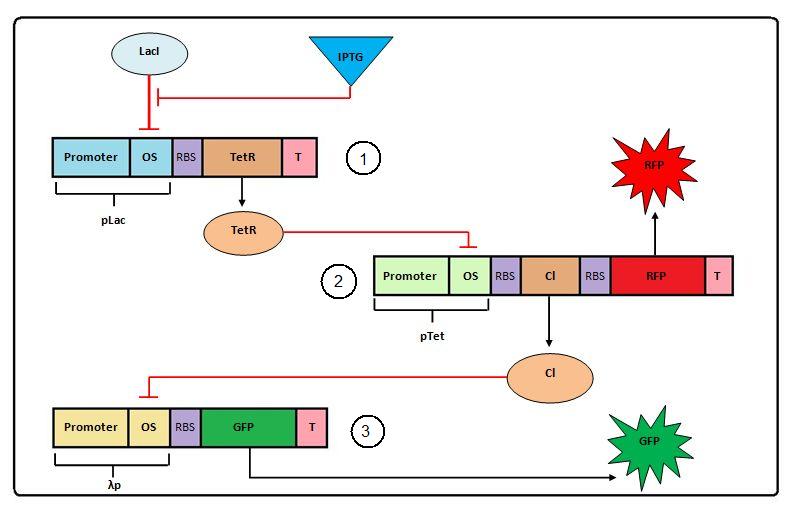
Figure 1: Negative Feedforward model
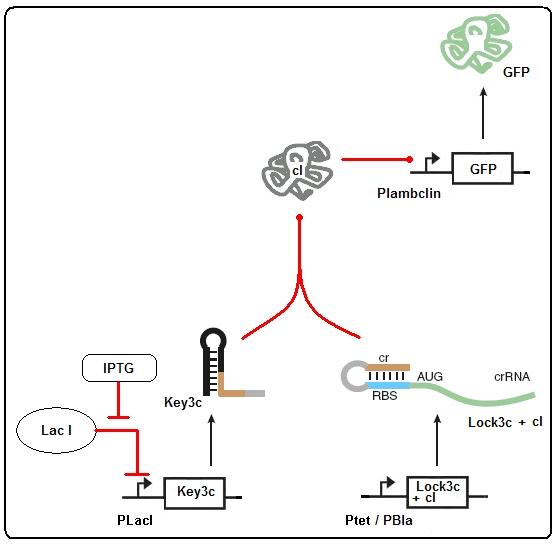
Figure 2: Riboregulator model
Mathematical Modeling
The Negative Feedforward model shown above can be expressed with the following set of ODEs (ordinary differential equations):
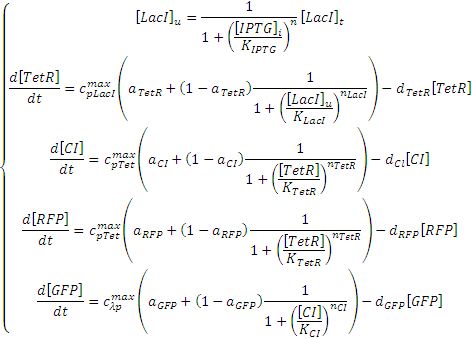
The Riboregulator model shown above can be expressed with the following set of ODEs:
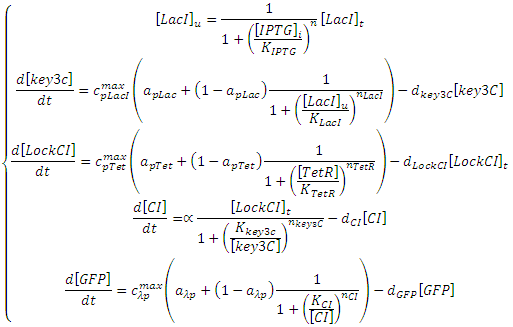
The symbols in this system of equations are found in the table below:
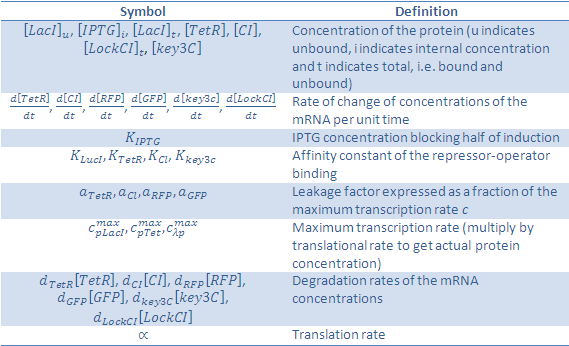
We would like to give credit to the Bologna '08 team and UNIPV-Pavia '08 team, who's project was useful in determining our own equations.
Conjugation
Using the paper "A model for bacterial conjugal gene transfer on solid surfaces", we established the assumptions and parameters that would be needed for the .
Assumptions
Parameters
- Surface area of media (A)
- Initial colony radius (r_0)
- Specific growth rate (g_n)
- Colony radial growth rate (g_r)
- Maximum numbers of cells sustained by system (N_max)
- Initial number of donors (N_d) and recipients (N_r)
All these parameters can be determined experimentally, or by literary research into previous conjugation experiments.
Spatial distribution of bacteria
Growth of colonies
Contact between colonies
Conjugation modeling
Final cell number
 "
"







