Team:Uppsala-Sweden/Ethanol
From 2009.igem.org

The Ethanol Project
Our goal is to introduce ethanol producing capability into a cyanobacteria using the biobrick system.
How to
Cyanobacteria have the capability to harvest the energy from the sun and convert it into other forms of energy. The natural way for these organisms is to store it as sugars or other carbohydrates in a way similar to plants. By introducing a casette of genes for ethanol production, we could get a cyanobacteria to start producing ethanol. However the yield from this might be very low since the bacteria have already evolved good pathways in the metabolism to take care of the energy. The source of carbon for the ethanol is pyruvate which is also used in the krebs cycle. To get a higher yield the pyruvate pool must be large enough for the ethanol anabolism. By blocking the pyruvate dehydrogenase the usage of pyruvate in the Krebs cycle can be reduced and the pyruvate used for ethanol production instead.
The Constructs
We designed multiple constructs as we ran into some unexpected restriciton site problem during the assembling process. The Z. mobilis strain which was source of the pdc and adh, actually had most probably a mutation inside the adh2 gene, which lead to a novel EcoRI restriction site. Thus we decided to build a back up construct using the adh2 from S. cerevisiae.
We build as well versions for testing purposes in E. Coli.
: The Construct for Testing Purposes in E. Coli with adh2 from Z. mobilis
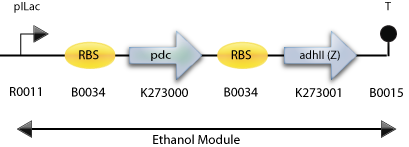
: The Final Construct for Synechocystis sp PCC6803 with adh2 from Z. mobilis
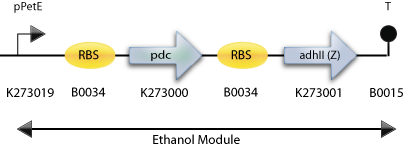
: The Construct for Testing Purposes in E. Coli with adh2 from S. cerevisiae
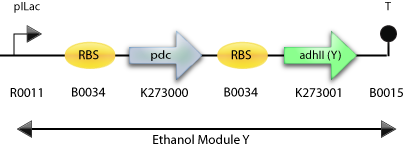
: The Final Construct for Synechocystis sp PCC6803 with adh2 from S. cerevisiae
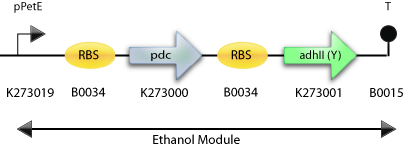
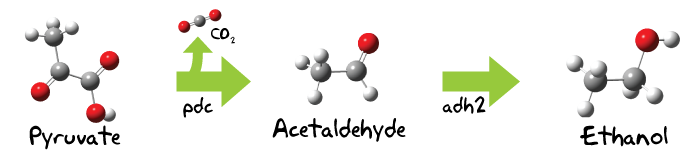
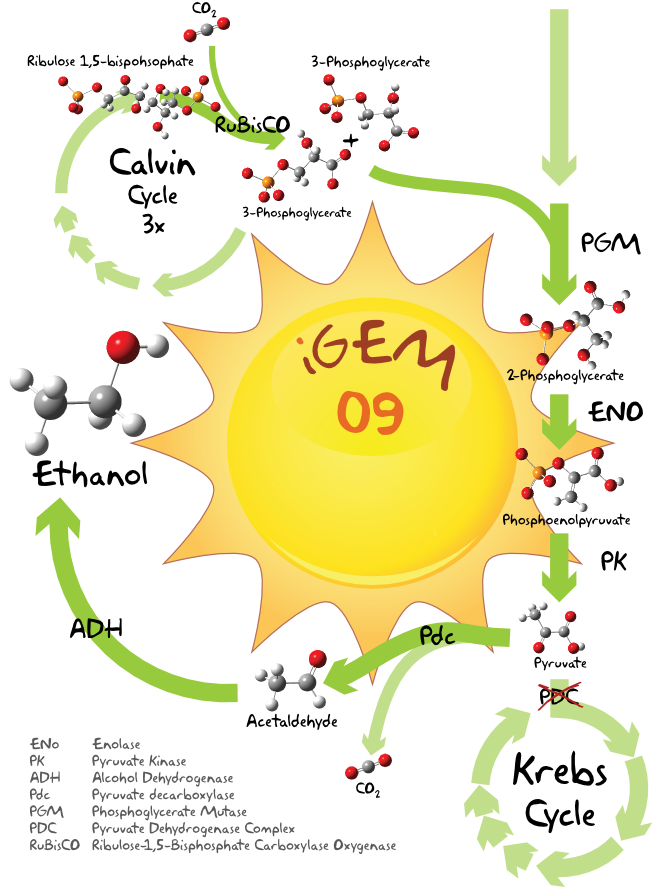
The Pathway
Here some theory...
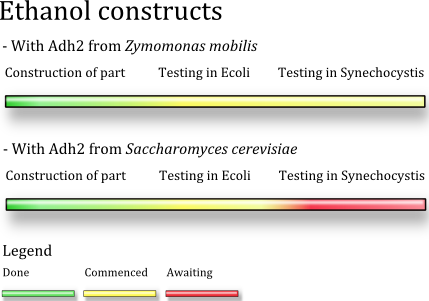
Progress
as of 2009-1018
We have completed the construction of both the construct variants and they are currently being tested in E.coli employing the pLac promoter. The Z.mobilis variant has been transformed to to Synechocystis and is currently growing to pick able size on plate, this takes about one week. The Yeast construct is lagging behind the Z.mobilis variant due to some failed transformations.
After transformation to Synechocystis it takes approximately two weeks till the culture has grown to enough volume to be used in tests.
 "
"