The Registry of Standard Biological Parts is a library of DNA sequences combined with online characterization resources. The Registry has created a standard protocol for making segments of DNA compatible with all other segments of DNA, regardless of order or size. BioBrick is the term given to such segments of DNA, a term alluding to the fact that any number of bricks may be combined in any order to produce complex, unique systems. This is accomplished by standardizing the restriction enzymes used to surround BioBricks, as well as the plasmids used to transform them. For a graphical representation of of the process, please click here. A powerful online database provides information and characterization of all of the BioBricks in the Registry and uses the wiki format (the same one used in wikipedia) which encourages others to edit content directly on the page. Below are list of parts that were used/created for this project.
Parts used to characterize and build our final project
| Component | Part/Accession # | Base Pairs | Plasmid | Resistance | Well |
| RBS-34 | BBa_B0034 | 12 | pSB1A2 | Ampicillin | plate 1, 2M |
| Cph8 | BBa_I15010 | 2,238 | pSB2K3 | Kanamycin | N/A |
| RFP | BBa_J04051 | 720 | N/A | N/A | N/A |
| OmpR (E. coli) | BBa_K098011 | 720 | pSB1T3 | Tetracycline | N/A |
| OmpR (R. sphaeroides) | BBa_K227010 | 720 | N/A | N/A | N/A |
| Terminator | BBa_B0015 | 129 | pSB1AK3 | Ampicillin and Kanamycin | plate 1, 23L |
| RBS +OmpR(sph) + Terminator
includes prefix and suffix | sequence | 916 | pany-amp | Ampicillin | synthesized |
| OmpC promoter | BBa_R0082 | 108 | pSB1A2 | Ampicillin | plate 1, 16K |
| puc promoter | BBa_K227007 | 651 | pSB1k3 | Kanamycin | N/A |
| puc BA | BBa_K227006 | 375 | pSB1k3 | Kanamycin | N/A |
| puc B | BBa_K227005 | 156 | pSB1k3 | Kanamycin | N/A |
| puc A | BBa_K227004 | 165 | pSB1k3 | Kanamycin | N/A |
| OmpC promoter+BA | sequence | 539 | pany-kana | Kanamycin | synthesized |
| TetR repressible | BBa_J13002 | 74 | pSB1A2 | Ampicillin | plate 1, 13B |
| Green Fluorescent Protein | BBa_E0240 | 876 | pSB1A2 | Ampicillin | plate 1, 12M |
Plasmids used to create and characterize our project
| Plasmid | Base Pairs | Resistance | Copy Number |
| pSB1A2 | 2,079 | Ampicillin | high |
| pSB1K3 | 2,206 | Kanamycin | high |
| pSB1A3 | 2,157 | Ampicillin | high |
| pSB2K3 | 4,425 | Kanamycin | variable |
| pSB1T3 | 2,463 | Tetracycline | high |
| pSB1AK3 | 3,189 | Ampicillin and Kanamycin | high |
| pANY | |||
| pRK404 | |||
| pRKPLHT7 | Tetracycline | ||
| pRKCBC3 | Tetracycline |
Parts submitted to the Registry of Standard Biological Parts
| Part/Accession #Component | Component | Type | Base Pairs | Plasmid | Resistance |
| BBa_I15010 | Cph8 (resubmission) | Coding | 2,238 | plasmid | Resistance |
| BBa_K227004 | puc A | Coding | 165 | pSB1K3 | Kanamycin |
| BBa_K227005 | puc B | Coding | 156 | pSB1K3 | Kanamycin |
| BBa_K227006 | puc BA | Coding | 376 | pSB1K3 | Kanamycin |
| BBa_K227007 | puc promoter | Regulatory | 651 | pSB1K3 | Kanamycin |
| BBa_K227008 | ompC+PucBA (synthesized) | Composite | 539 | pSB1AT3 | Ampicilin,Tetracycline |
| BBa_K227009 | PucPromotor+GFP | Composite | 1377 | pSB1A2 | Ampicilin |
| BBa_K227010 | OmpR (sphaeroides) | Coding- not available | n/a | n/a | n/a |
| BBa_K227011 | RBS34+OmpR+Term (synthesized) | Composite | 916? | pSB1K3? | Kanamycin? |
| BBa_K227012 | RBS34+OmpR(sph)+Term+OmpC+PucB/A | Composite | ??? | pSB1K3 | Kanamycin |
| BBa_K227013 | ompC + GFP | Composite | 992 | plasmid | Resistance |
| BBa_K227014 | pucpro+pucBA | Composite | ??? | pSB1K3 | Kanamycin |
| BBa_K227015 | RBS34+OmpR(sph)+Term+OmpC+GFP | Composite | ??? | plasmid | Ampicilin |
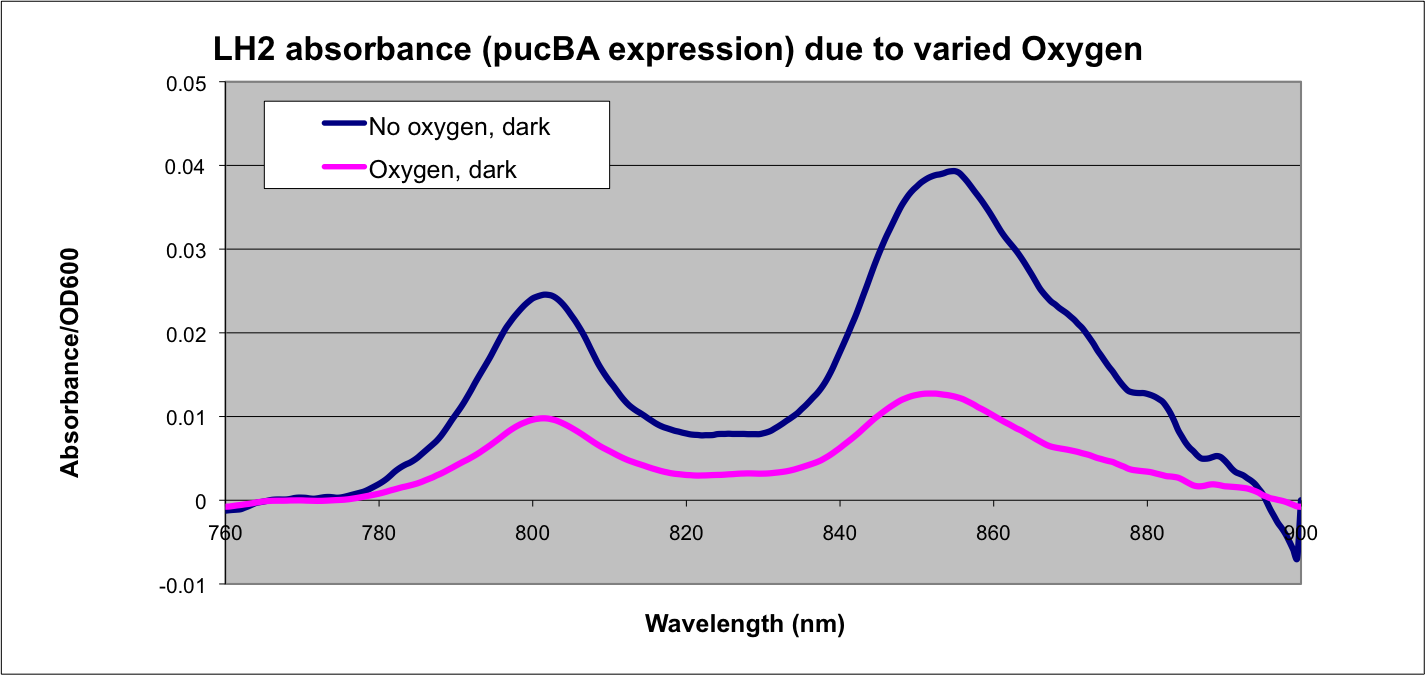
The first part of our characterization begins with the puc promoter, which promotes transcription of the LH2 pucB/A genes naturally in Rhodobacter sphaeroides. It is important that we are able to compare the transcription rate of the puc promoter in the natural system vs. our mutant system so that we can determine exactly how much efficiency is gained by adding a red light sensor. The absorption spectra of a DBCOmega mutant (LH2 deficient) transformed with pRKCBC3 containing the puc promoter and pucB/A genes will allow us to characterize the puc promoter under high and low oxygen conditions.
More absorption of light at the LH2 spectra peaks normalized to culture OD corresponds with more transcription and vis versa.
Method
The optical density of the cultures was recorded at 600nm, and each absorbance spectrum was normalized to OD600 by division by this value. Background subtraction of spectrophotometer data was performed in Origin 6.1 Software. A ten-point baseline was created by a "positive peak" algorithm then modified to approximate the scattering curve that falls as the inverse fourth power of wavelength.
Results
Modeling the Gene Regulatory Network
References
1. Alon, Uri. Introduction to systems biology and the design principles of biological networks. Boca Raton, FL: Chapman & Hall, 2006.
2. Bower, James M. Computational Modeling of Genetic and Biochemical Networks (Computational Molecular Biology). New York: M.I.T. PRESS, 2001.
3. System modeling in cellular biology from concepts to nuts and bolts. Cambridge, MA: MIT P, 2006.
Simulating a Bioreactor

|