Team:Washington/Project/Display
From 2009.igem.org
Display System
|
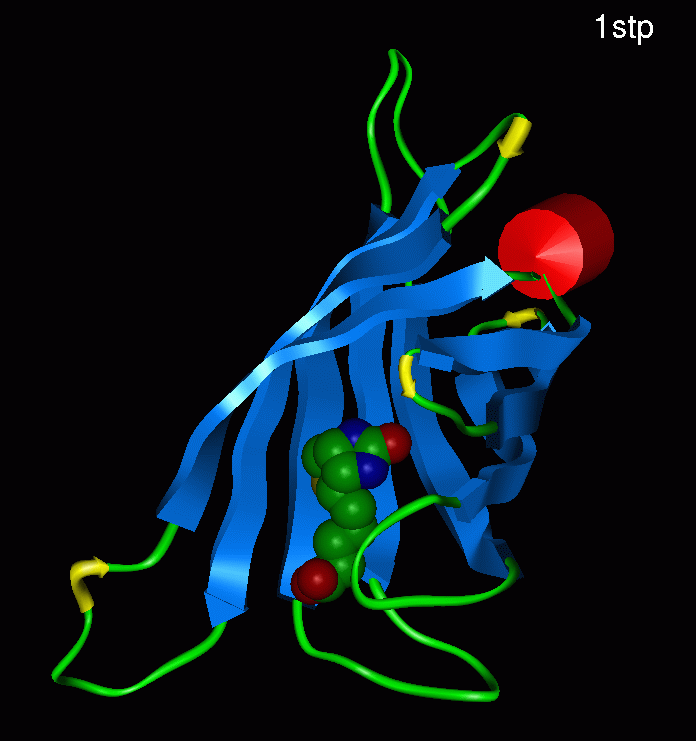
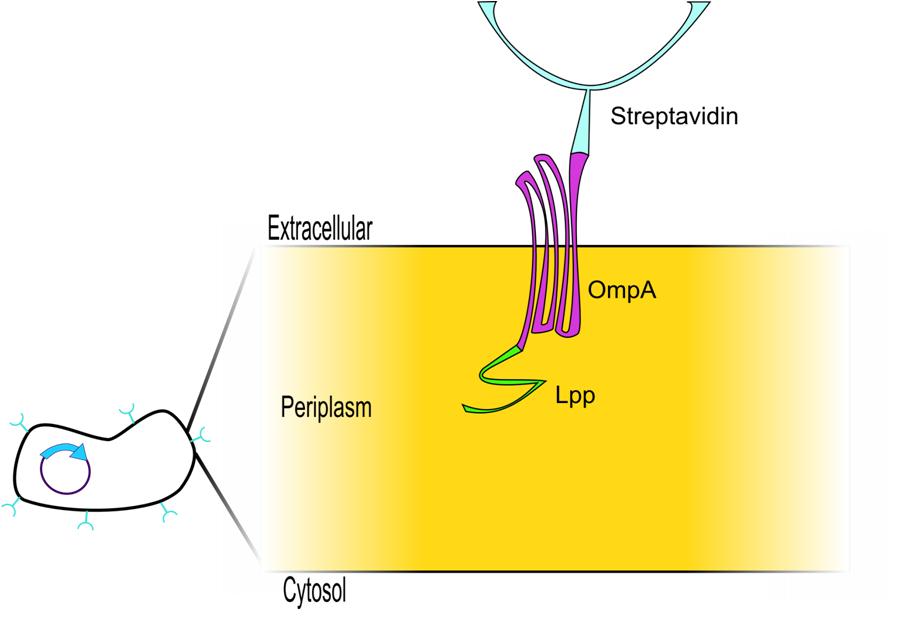
BackgroundOur system hinged on finding a protein which could bind other proteins to the outside of the cell, but whose interaction with these proteins was weak enough to be disrupted by another small molecule. Streptavadin presented itself as a logical choice because it can bind biotin (a small molecule) as well as a nanotag sequence which can be added easily to any protein sequence [1]. The streptavadin biotin complex is shown below [11].
Now we had the parts that we wanted. After looking at the work Harvard did on these parts, we decided to do the following set of tests.
ExperimentsWesternThe goal of this experiment was to make sure that the proteins were being expressed in our cell lines, and also to make sure that they were the correct length. This was crucial to ascertain before we moved on and began to test the parts. Even though we had the individual parts sequenced confirmed we needed to make sure that the proteins were being expressed correctly in the cell. Since all our the Harvard parts are conveniently his tagged this means that we could use a his tag antibody to detect the presence of these proteins in our western [8].
ResultsFile:Western.jpg Waiting for image from Ingrid The expected values of each of these proteins are shown in the table below:
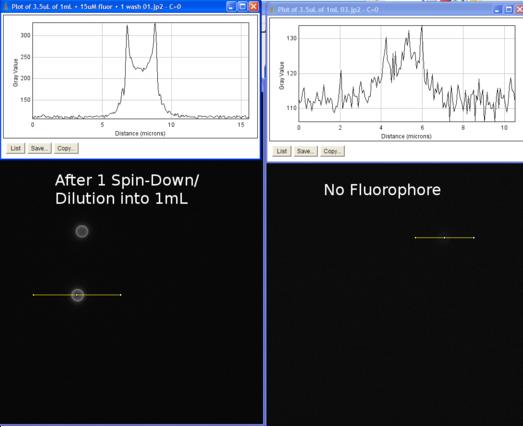
This data showed that indeed our parts did come out at the expected lengths. The varying brightness of the bands does not indicate the strength of expression because the protein amount was not normalized before it was inserted into the gel. This was an important conclusion, which proved to us that these proteins were indeed being expressed by the cell. Now we could move on to check to see if the proteins were being expressed on the surface, and if they were functional. To do this we did a microscope assay. Microscope AssayThe goal of this experiment was to see if we could confirm the presence of streptavadin on the outside of our cell. We wanted to do this using a biotyntalated flourophore. We expected that this flourophore would bind to the streptavadin on the outside of the cell in such a high level that we would be able to detect it under a high powered florescence microscope. The resulting concentration of florescence if high enough, would result in a halo of florescence around the cells. As a positive control we used streptavadin coated beads which were roughly the same size (with respect to volume) as our cells (except spherical).
Data
Flow CytometryThe goal of this experiment was to see how the biotin bonded to streptavadin over a whole population of cells. Where as the Microscope experiment looked at a few localized cells, we set the cytometer to look 50,000 cells and read the resulting florescence. Our positive control was again the strepatavadin coated beads. The cytomtery would be able to read the flouresence of each individual particle passing through it, which would allow us to get much more accurate reading, and look at a much larger sample size than we could under the microscope. The results of this experiment are described below.
Datahttp://soslab.ee.washington.edu/igem/2009/images/f/f9/Hists.png File:2009-08-07 51s.png Our positive control showed nice separation between the beads with flourophore and the beads without flourophore. This confirmed what we saw under the microscope, and also indicated that our assay was working. However this data showed us that we did not have any appreciable binding occurring with any of the ompA-sterptavadin parts to biotin at the global level. All our the cells have low levels of florescence which are comparable to those of the beads. ConclusionIn conclusion we were able to shown that the proteins were being expressed in the cell by a western blot. We were unable however to see any sort of binding between the biotyntalated flourophore and the cells expressing these proteins. This meant that our display system was likely not binding biotin correctly. We hypothesized this could have been do to streptavadin not having enough room to form tetramers on the surface (which is its native state)[10]. Our solutions to theses problems are described in the future directions section. References
|
 "
"