Team:Waterloo/Modeling
From 2009.igem.org
Dookehster (Talk | contribs) (→Conclusion) |
Dookehster (Talk | contribs) (→Software) |
||
| (18 intermediate revisions not shown) | |||
| Line 3: | Line 3: | ||
===Abstract=== | ===Abstract=== | ||
| - | + | The primary goal of our software was to model integrase mediated DNA rearrangement. After our software was capable of simulating a biological system to steady state, our secondary goal was to be able to generate the initial reactants needed to arrive at a given end state. It was found that our brute force approach could not be used to achieve the second goal as the simulation would run out of hardware resources. | |
===Introduction=== | ===Introduction=== | ||
| Line 11: | Line 11: | ||
The first stage of the modeling project was therefore to formally codify the reaction rules employed by the design team with the aim of applying computational power to the problem. The predominant challenge faced at this stage was to abstract the concept of DNA strands into a computationally workable form along with developing mathematically rigorous definitions of the behaviours of reaction sites. | The first stage of the modeling project was therefore to formally codify the reaction rules employed by the design team with the aim of applying computational power to the problem. The predominant challenge faced at this stage was to abstract the concept of DNA strands into a computationally workable form along with developing mathematically rigorous definitions of the behaviours of reaction sites. | ||
| - | Formally, the grand object of the modeling project was the determination of a finite deterministic sequence of ''att'' sites and their enclosed operators that would allow one to predictably insert into a chromosome | + | Formally, the grand object of the modeling project was the determination of a finite deterministic sequence of ''att'' sites and their enclosed operators that would allow one to predictably insert into a chromosome an arbitrary number of desired sequences. |
| - | The form that the solution could take, we postulated would be a sequence of two or three plasmids such that each would contain at least one matching set of ''att'' sites with the addition of several incomplete ''att'' sites. | + | The form that the solution could take, we postulated, would be a sequence of two or three plasmids such that each would contain at least one matching set of ''att'' sites with the addition of several incomplete ''att'' sites. |
| - | There were two general approaches used in | + | There were two general approaches used in modeling. Software development toward a top-down (inductive, brute force) solver has finished. Characterization of the algorithm underpinning the solver, however, revealed that the problem is NP-hard at least, and NP-complete at worst. An auxiliary approach was attempted whereupon we tried to map the sequence problem onto a mathematical problem with known solutions. |
===Software=== | ===Software=== | ||
| - | In order to run the solver, we | + | It was decided early in the design stages of the mathematical simulation that representing strands of DNA as strings of nucleotides was highly inefficient. Instead, each functional block is represented as a single token, as shown in table 1. |
| + | |||
| + | {| class="wikitable" border="1" style="width:100%;" | ||
| + | |+ Table 1: Token Mapping | ||
| + | |- | ||
| + | ! Sequence | ||
| + | ! Corresponding Token | ||
| + | |- | ||
| + | | style="width:50%;" |P [attP site is composed of POP'] | ||
| + | | P | ||
| + | |- | ||
| + | | P' | ||
| + | | Q | ||
| + | |- | ||
| + | | B [attB site is composed of BOB'] | ||
| + | | B | ||
| + | |- | ||
| + | | B' | ||
| + | | D | ||
| + | |- | ||
| + | | Chromosomal Origin | ||
| + | | 0 or 9 | ||
| + | |- | ||
| + | | Plasmid Origin | ||
| + | | 1 to 8 | ||
| + | |- | ||
| + | | Nucleotides | ||
| + | | A, C, G, T | ||
| + | |- | ||
| + | | Selection Genes | ||
| + | | Counter Selectable Markers: M, N | ||
| + | Selectable Markers: I, J, K, L | ||
| + | |- | ||
| + | | Sequence of Interest | ||
| + | | X, Y, Z | ||
| + | |- | ||
| + | | Complements | ||
| + | | Lowercase characters become uppercase and the vice versa. For origins, subtract the digit to be complemented from nine (ie. 0 and 9 are complements). For nucleotides, A/T and G/C remain complemented pairs. | ||
| + | |} | ||
| + | |||
| + | Note that nucleotides are still individually represented by tokens, even though most sequences have abstract tokens. This is necessary because the behavior of the att sites is dependent on the core dinucleotide and the most useful representation of the dinucleotide is simply a pair of nucleotides. | ||
| + | |||
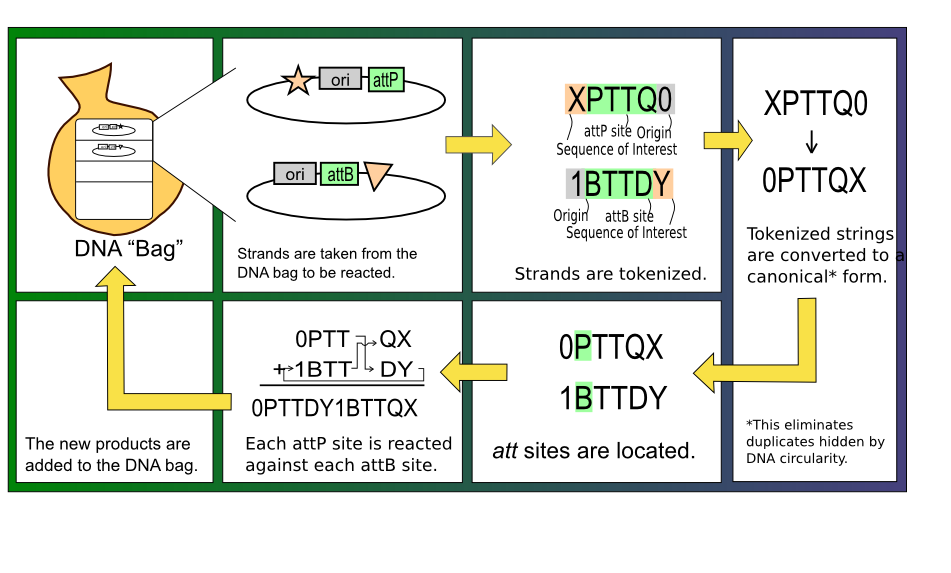
| + | With this mapping of DNA sequences to characters completed, the algorithm to react two strands of DNA together is straightforward. First, the P and B tokens and their complements must be found, indicating the location of potential att sites. The validity of these att sites are then checked by ensuring that two nucleotides and a corresponding P' or B' sequence are present in the correct orientation. At this stage, each valid attP site can be reacted with any valid attB site by starting at one of the sites and iterating through the string of tokens until the other site is reached. Depending on the orientations of the two sites and whether two strands or a single molecule are involved in the reaction, the result may be an excision, an inversion, or a merge. The figure below gives a visual representation of this process. | ||
| + | |||
| + | [[Image:RecombinatronProcess.PNG|frame|center|The partial control flow of the simulation; filtering steps are not included.]] | ||
| + | |||
| + | In order to run the solver, we made several assumptions. First, as we did not know which combinations of sequences with ''att'' sites would be part of the solution, we assumed that any product in our search space was a valid sequence for the next generation of reactions. We further assumed that any plasmid with valid ''att'' sites and complementary operators was capable of self reacting and also of reacting with any other plasmid in the history of the modeled cell. | ||
| + | |||
| + | Filters were set to regularly delete DNA strands that did not meet selection criteria. The frequency of the filtering was based on intuitive notions of the number of operations that could occur before stable strands were produced, where each operation consists of one reaction of a pair of att sites. These frequency values were originally set conservatively high at 30 operations for filtering based on the number of origins present on a strand and 255 operations for filtering based on selectable markers. In later testing, the frequency of filtering was drastically increased in an attempt to remove unnecessary products and improve the simulation's performance. | ||
| + | |||
| + | Lastly, because of the exponential behaviour of the search space, we assumed that the smallest solution that exists can be found within the search space generated after reacting 10E7 plasmids. This assumption was made in order to have sane parameters for termination. | ||
===Math=== | ===Math=== | ||
| - | An ancillary branch | + | An ancillary branch of investigation arose out of the necessity to tend to the exponential behaviour of the problem. There may exist some math that inherently facilitates the modeling and solving of this problem. We explored maths that mainly dealt with topology (knot theory) and functional reasoning (lambda theory, combinatory calculus) but finally could not identify a good candidate as a scaffold to our solution. |
===Results=== | ===Results=== | ||
| - | To test the program, we ensured that it was capable of doing cassette exchange. In cassette exchange, a plasmid has a gene | + | To test the program, we ensured that it was capable of doing cassette exchange. In cassette exchange, a plasmid has a gene of interest flanked by two ''attB'' sites and the chromosome has a marker flanked by two ''attP'' sites. After enzyme-mediated recombination, the gene of interest should be in the chromosome, where the flanking sites will be changed to ''attL'', and the marker will be in the plasmid, where the flanking sites will be changed to ''attR''. No other products are possible after this reaction has run to completion. Our program was able to correctly perform the recombination and emulate the selection process on various types of selectable media. |
| - | One of our team members | + | One of our team members postulated a potentially-viable recursive recombination system. We converted this into a format usable by the program and ran it. Unfortunately, the program ran out of memory before running to steady state. After several rounds of optimization and running the program on a computer with 12GB of RAM provided by Dr. Moreno's lab at Wilfrid Laurier University, we were still unable to run the program to steady state. |
The combinatorial explosion of reaction products of the integrase reaction was far greater than anticipated. Even increasing the selection pressure beyond the point of biological possibility failed to control the combinatorial explosion. | The combinatorial explosion of reaction products of the integrase reaction was far greater than anticipated. Even increasing the selection pressure beyond the point of biological possibility failed to control the combinatorial explosion. | ||
| Line 35: | Line 85: | ||
===Conclusion=== | ===Conclusion=== | ||
| - | Due to the sheer number of permutations that would have occurred in our | + | Due to the sheer number of permutations that would have occurred in our biological system, the initial assumptions upon which we built our software were incorrect. That is, given our hardware resources, it was not feasible for a brute force algorithm to reach steady state. |
| - | From the beginning until our actual simulations, emphasis was placed on algorithm design rather than choice of language or underlying implementations of data structures. Because it had been empirically proven that our given solution was not sufficient, a C++ | + | From the beginning until our actual simulations, emphasis was placed on algorithm design rather than choice of language or underlying implementations of data structures. Because it had been empirically proven that our given solution was not sufficient, a re-implementation in C++ was initiated to maximize efficiency given a finite amount of computing power. However, at the time of writing, this has yet to be finished. We hope that the C++ version will significantly reduce our algorithm's running time, but given the combinatorial explosion in the Python version, this may not be the case. If so, effort must be put into devising a new algorithm rather than into optimizing the program. |
| - | Steps | + | Steps that may have led to our simulation's reaching steady state include: |
| - | + | <ul> | |
| - | + | <li>Optimizing on the lowest level from start to finish.</li> | |
| - | + | <li>Use of distributed computing techniques.</li> | |
| - | + | <li>Use of a field programmable gate array (FPGA) rather than a conventional computing solution.</li> | |
| + | <li>More exploration into devising a non-brute force algorithm.</li> | ||
| + | </ul> | ||
| - | Following the Jamboree, the modeling team is likely to seek | + | Following the Jamboree, the modeling team is likely to seek solutions in two avenues: optimizing our current algorithm, be it in Python or otherwise, and continuing to search non-obvious rigorous methods to determine a steady state. |
Latest revision as of 00:46, 22 October 2009
Contents |
Abstract
The primary goal of our software was to model integrase mediated DNA rearrangement. After our software was capable of simulating a biological system to steady state, our secondary goal was to be able to generate the initial reactants needed to arrive at a given end state. It was found that our brute force approach could not be used to achieve the second goal as the simulation would run out of hardware resources.
Introduction
The mathematical modeling component of this year's project consisted of a computational simulation of DNA recombination as mediated by the ΦC31 integrase enzyme. The necessity for this simulation arose directly from challenges faced by the design team in its attempts to create a recursively repeatable technique for inserting sequences of interest onto chromosomes. Specifically, it was noted that manually examining the possible results of interactions between DNA strands quickly becomes infeasible due to the number of potential reaction pathways.
The first stage of the modeling project was therefore to formally codify the reaction rules employed by the design team with the aim of applying computational power to the problem. The predominant challenge faced at this stage was to abstract the concept of DNA strands into a computationally workable form along with developing mathematically rigorous definitions of the behaviours of reaction sites.
Formally, the grand object of the modeling project was the determination of a finite deterministic sequence of att sites and their enclosed operators that would allow one to predictably insert into a chromosome an arbitrary number of desired sequences.
The form that the solution could take, we postulated, would be a sequence of two or three plasmids such that each would contain at least one matching set of att sites with the addition of several incomplete att sites.
There were two general approaches used in modeling. Software development toward a top-down (inductive, brute force) solver has finished. Characterization of the algorithm underpinning the solver, however, revealed that the problem is NP-hard at least, and NP-complete at worst. An auxiliary approach was attempted whereupon we tried to map the sequence problem onto a mathematical problem with known solutions.
Software
It was decided early in the design stages of the mathematical simulation that representing strands of DNA as strings of nucleotides was highly inefficient. Instead, each functional block is represented as a single token, as shown in table 1.
| Sequence | Corresponding Token |
|---|---|
| P [attP site is composed of POP'] | P |
| P' | Q |
| B [attB site is composed of BOB'] | B |
| B' | D |
| Chromosomal Origin | 0 or 9 |
| Plasmid Origin | 1 to 8 |
| Nucleotides | A, C, G, T |
| Selection Genes | Counter Selectable Markers: M, N
Selectable Markers: I, J, K, L |
| Sequence of Interest | X, Y, Z |
| Complements | Lowercase characters become uppercase and the vice versa. For origins, subtract the digit to be complemented from nine (ie. 0 and 9 are complements). For nucleotides, A/T and G/C remain complemented pairs. |
Note that nucleotides are still individually represented by tokens, even though most sequences have abstract tokens. This is necessary because the behavior of the att sites is dependent on the core dinucleotide and the most useful representation of the dinucleotide is simply a pair of nucleotides.
With this mapping of DNA sequences to characters completed, the algorithm to react two strands of DNA together is straightforward. First, the P and B tokens and their complements must be found, indicating the location of potential att sites. The validity of these att sites are then checked by ensuring that two nucleotides and a corresponding P' or B' sequence are present in the correct orientation. At this stage, each valid attP site can be reacted with any valid attB site by starting at one of the sites and iterating through the string of tokens until the other site is reached. Depending on the orientations of the two sites and whether two strands or a single molecule are involved in the reaction, the result may be an excision, an inversion, or a merge. The figure below gives a visual representation of this process.
In order to run the solver, we made several assumptions. First, as we did not know which combinations of sequences with att sites would be part of the solution, we assumed that any product in our search space was a valid sequence for the next generation of reactions. We further assumed that any plasmid with valid att sites and complementary operators was capable of self reacting and also of reacting with any other plasmid in the history of the modeled cell.
Filters were set to regularly delete DNA strands that did not meet selection criteria. The frequency of the filtering was based on intuitive notions of the number of operations that could occur before stable strands were produced, where each operation consists of one reaction of a pair of att sites. These frequency values were originally set conservatively high at 30 operations for filtering based on the number of origins present on a strand and 255 operations for filtering based on selectable markers. In later testing, the frequency of filtering was drastically increased in an attempt to remove unnecessary products and improve the simulation's performance.
Lastly, because of the exponential behaviour of the search space, we assumed that the smallest solution that exists can be found within the search space generated after reacting 10E7 plasmids. This assumption was made in order to have sane parameters for termination.
Math
An ancillary branch of investigation arose out of the necessity to tend to the exponential behaviour of the problem. There may exist some math that inherently facilitates the modeling and solving of this problem. We explored maths that mainly dealt with topology (knot theory) and functional reasoning (lambda theory, combinatory calculus) but finally could not identify a good candidate as a scaffold to our solution.
Results
To test the program, we ensured that it was capable of doing cassette exchange. In cassette exchange, a plasmid has a gene of interest flanked by two attB sites and the chromosome has a marker flanked by two attP sites. After enzyme-mediated recombination, the gene of interest should be in the chromosome, where the flanking sites will be changed to attL, and the marker will be in the plasmid, where the flanking sites will be changed to attR. No other products are possible after this reaction has run to completion. Our program was able to correctly perform the recombination and emulate the selection process on various types of selectable media.
One of our team members postulated a potentially-viable recursive recombination system. We converted this into a format usable by the program and ran it. Unfortunately, the program ran out of memory before running to steady state. After several rounds of optimization and running the program on a computer with 12GB of RAM provided by Dr. Moreno's lab at Wilfrid Laurier University, we were still unable to run the program to steady state.
The combinatorial explosion of reaction products of the integrase reaction was far greater than anticipated. Even increasing the selection pressure beyond the point of biological possibility failed to control the combinatorial explosion.
Conclusion
Due to the sheer number of permutations that would have occurred in our biological system, the initial assumptions upon which we built our software were incorrect. That is, given our hardware resources, it was not feasible for a brute force algorithm to reach steady state.
From the beginning until our actual simulations, emphasis was placed on algorithm design rather than choice of language or underlying implementations of data structures. Because it had been empirically proven that our given solution was not sufficient, a re-implementation in C++ was initiated to maximize efficiency given a finite amount of computing power. However, at the time of writing, this has yet to be finished. We hope that the C++ version will significantly reduce our algorithm's running time, but given the combinatorial explosion in the Python version, this may not be the case. If so, effort must be put into devising a new algorithm rather than into optimizing the program.
Steps that may have led to our simulation's reaching steady state include:
- Optimizing on the lowest level from start to finish.
- Use of distributed computing techniques.
- Use of a field programmable gate array (FPGA) rather than a conventional computing solution.
- More exploration into devising a non-brute force algorithm.
Following the Jamboree, the modeling team is likely to seek solutions in two avenues: optimizing our current algorithm, be it in Python or otherwise, and continuing to search non-obvious rigorous methods to determine a steady state.
 "
"