Wiki/Team:Warsaw/igem project.htm
From 2009.igem.org
m |
|||
| Line 1: | Line 1: | ||
{{WarHead|headimg=2009.igem.org/wiki/images/1/1d/WarLogoProject.jpg}} | {{WarHead|headimg=2009.igem.org/wiki/images/1/1d/WarLogoProject.jpg}} | ||
| + | |||
| + | |||
| + | __TOC__ | ||
==Aims of the project== | ==Aims of the project== | ||
Revision as of 21:43, 2 July 2009
 |
||
|
Aims of the projectOne of the most important challenges in the field of modern medicine is to invent the efficacious anticancer therapy. The gene therapy appears to be the most effective strategy for metastasis remission without dangerous side effects, which are common disadvantages in the case of both radioand chemotherapy. Use of bare DNA particles is very inefficient, therefore new DNA delivery system is needed. The most common vectors used in genetic therapy are genetically altered viruses. However they generate problems that have not been solved yet. An integration into the host genome may lead to unpredictable sideeffects and increases cancer risk. Moreover this type of vector could cause acute immune response or symptoms of viral infection. Artificial systems of DNA or RNA delivery such as liposomes or alkaline polymers are characterized by low effectiveness and high toxicity. The main aim of this project is to design a model system based on genetically modified Escherichia coli bacteria able to invade eukaryotic cells. This approach is the new alternative for established for years techniques. It has most of advantages of previously known viral vectors and, importantly, it does not induce negative changes in the patient’s genome. The created bacterial system will be applied to two problems. The first one is to create special bacterial strain, which is able to conjugate with the mitochondrion within the cytoplasm. It will enable effective DNA delivery in scientific or therapeutic approaches. The second strategy is to induce apoptosis of tumor cells by means of the proapoptotic proteins secreted to the cytoplasm by bacteria that exist within the cell. Theoretical basisEntrance of bacteria into eukaryotic cellsMany bacterial species are able to invade an eukaryotic cell. One of the crucial proteins for this process is invasin. It is capable of selective interaction with integrins, which are present on eukaryotic membrane. It triggers signalling cascade indispensable to start endocytosis [9]. This interaction occurs via the invasin N-terminal domain, which is able to promote effective endocytosis of bacteria that synthesize invasin by cells normally unable to undertake phagocytosis [3]. Majority of the enteropathogenic bacteria are incapable to divide within the phagosome, but after leaving the phagosome they can proliferate rapidly in the cytoplasm. Escape from the phagosome is possible because of hemolysin which causes membrane permeabilisation and phagosome disruption [16]. It was presented previously that transfection with gene encoding hemolysin is sufficient to enable Bacillus subtilis and Escherichia coli to escape phagosomal vesicle [1]. Mitochondrial transformationElectron transport chain function can be impaired by a variety of defects acting on the DNA level: not only point mutation and local deletions in genes encoding proteins of the chain, but also deletions of huge fragments or even whole mitochondrial genome (mtDNA). Correct mitochondrion contains 2 to 10 copies of mtDNA. Defects in mitochondrial DNA result in severe medical disorders especially in tissues which consume large amount of energy like nervous and muscle tissue. Effects of this impairments are neuroand myopathies [20]. The standard therapy based on bypassing interrupted pathways does not apply under mentioned circumstances. Administration of enzyme cofactors or other metabolically active substances is not effective and may be applied only as supplementary therapy. We are forced to seek novel treatment strategies to cure patients suffering from mitochondrial disorders [7]. Currently the most commonly used method of mitochondrial transformation is the "gene gun" approach which is based on delivery of metal beads coated with DNA into the cell. This method is mostly limited to plants and yeast and has low therapeutic potential [7]. Current mammalian mitochondrial transformation techniques like electroporation are also limited. Whole process takes place ex vivo and causes irreversible mitochondrion destructions which make its efficient reintroduction to the cell almost impossible. The conjugation between bacteria and mitochondria free from these side effects. Moreover, this phenomenon has been proven to act in vitro. Conjugative plasmid had been placed in mitochondrion, where the expression of encoding genes occurred [22]. More recently a conjugation between two strains of bacteria within the cytoplasm of a mammalian cell has been observed . These findings suggest that conjugation between a bacterium and a mitochondrion is feasible [10]. Bacterial systems of protein secretionBacteria in natural environment use diverse, effective transport and secretion systems. In gram negative bacteria like E. coli common are type I secretion systems e.g. well characterized hemolysin secretion system. Hemolysin is responsible for pathogenicity of some E. coli strains. Systems using fusion of protein of interest to hemolysin C-terminus result in secretion of the protein into the growth medium[6] The same method can be use to secret bacterial proteins into the eukaryotic cell. It is currently used by designing new generation vaccines[5]. Induction of apoptosis by p53Mitochondrial outer membrane permabilization MOMP is one of the main programmed cell death mechanisms. The process is regulated by PTPc complex and Bcl2 family proteins [17]. Many cancer types involve disruption of this process therefore inducing MOMP in the tumour cells is regarded as potential therapeutic solution. Factors inducing MOMP include p53 protein. It is mutated in almost half of the cancer cases. Protein p53 by acting as a transcription regulator in the nucleus arrests the cell cycle and induce apoptosis [18]. In the cytoplasm p53 interacts with mitochondrial membrane and can enter the mitochondrium. It has been proven that these processes induce apoptosis and do not influence the cell cycle. In the mitochondrium p53 is an effector protein that activates soluble (cytochrom c, Smac) and insoluble (AIF, EndoG) apoptototic factors. Moreover interaction of p53 and Bax protein in vitro leads to permabilisation of lipid bilayer analogous to mitochondrial membrane. [12] Because p53 is inactive in tumour cells many research groups focused on designing gene therapies based on p53 gene delivery. Most frequently adenoviruses and eukaryotic plasmids were used. The research results confirmed increased effectiveness of chemotherapeutic drugs after introduction of p53 gene copy into the cell [4]. It has been shown that p53 with the mitochondrial leader sequence is enough to induce apoptosis in mammalian cells. The fusion protein Lp53 with mitochondrial leader sequence from ornithine transcarbamylase was used. In terms of apoptosis induction fusion protein was almost as effective as standard p53. Previous research did not include delivery of p53 into tumour cells by bacterial vectors. Secretion of p53 into the cytoplasm by bacteria has an advantage over gene therapy because it does not disrupt genome integrity. Results of p53 integration with host genome are not fully characterized yet, but too high level of p53 is shown to accelerate ageing. Our project proposes a method to induce apoptosis without arresting the cell cycle. Cell cycle arrest decreases the sensitivity of cancer cells to chemotherapeutic agents, because they can affect only proliferating cells [14]. Detailed research projectIntroduction
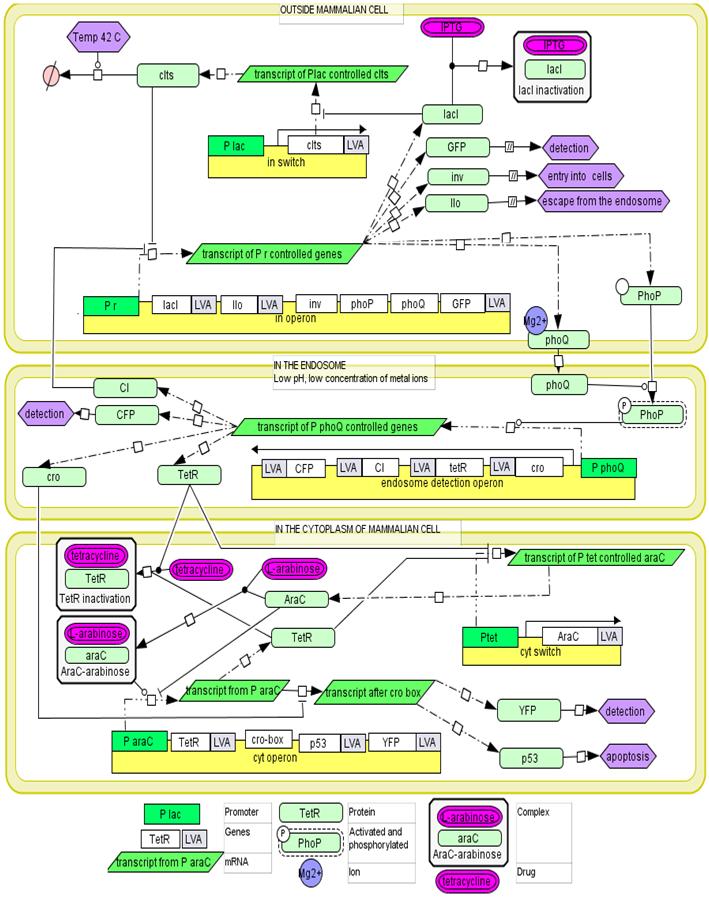
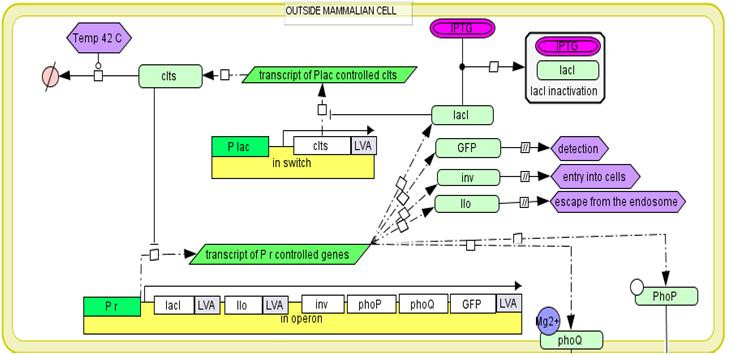
Invasion operon
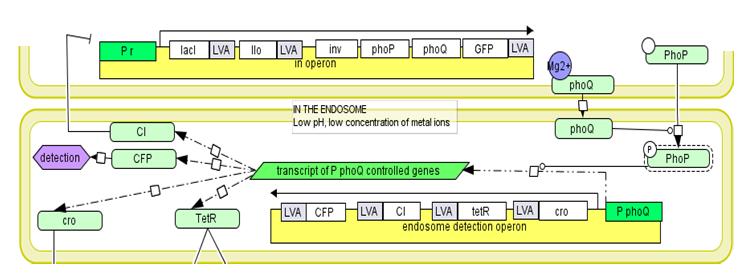
Endosome detection operon
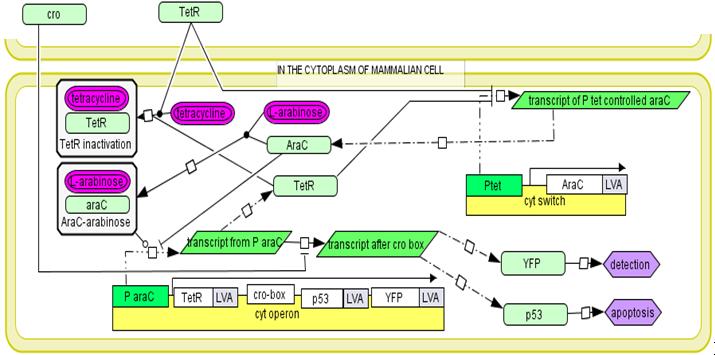
The cytoplasmic operon.
The mechanism of proposed regulatory system.In the medium the expression of all genetic circuits is inactive. After temperature increase to 42°C the system is activated by thermal deactivation of cI. Genes from invasion operon are transcribed. They maintain their own expression, cause internalisation of E. coli by mammalian cells and enable escape from endosome. When bacterial cell is inside the endosome the endosome detection module is activated by phoP/phoQ system. In trans expression of regulatory genes cI and tetR deactivates invasion operon and activates cytoplasmic operon. However its genes are not expressed as long as bacterial cell is inside endosome due to cro protein activity. When bacterium escapes from the endosome, the transcription from endosome detection operon is repressed and cytoplasmic operon genes are expressed. Conjugation with mitochondria.To place the copy of the mitochondrial genome into bacterial cell the pBACrNESd plasmid will be used. It is built basing the Bacterial Artificial Chromosome. Replication origin (ori R6K), kanamycin resistance gen (kanR) and replication complex genes (repE, parA, parC) from pBACrNESd plasmid will be fused with origin of transfer (oriTF) from natural conjugative F plasmid. Unique restriction site will be introduced to the mitochondrial genome by PCR reaction and the described gene construct will be inserted into mitochondrion. To enable conjugation proteins encoded on RP4 plasmid (Tc, MuKm, Tn7) will be placed in the cytoplasmic operon. These genes are responsible for pillus formation and transport of single stranded DNA. The cell with this system will conjugate with any structure surround by lipid bilayer. We will prepare mitochondrial DNA deficient cells using procedure described in [2]. The occurrence of conjugation will be confirmed by PCR reaction specific for sequences added to mitochondrial DNA. The quality of the DNA will be determined by analysis of restriction patterns. The expression of mitochondrial genes will be confirmed by semiquantitative time PCR [2]. Secretion of p53 into the cytoplasmTo efficiently transport p53 to the eukaryotic cells the endogenic E. coli secretion system responsible for secretion of hemolysin by uropathogenic strains will be used. It is composed of three transport proteins HlyB, HlyD and TolC [10]. TolC is present in the genome of the lab strain derived from K12 strain. To ensure efficient secretion hlyB and hlyD have to be introduced into the genome and proapoptotic protein has to be fused with 60 amino acid long C-terminal sequence of hemolysin. [5]. Directing bacterial proteins to mammalian mitochondria.We need to be able to direct to the mammalian mitochondria some of the proteins secrete by bacteria into the cytoplasm of the host cell. It is going to be archived by fusion of mitochondrial leader sequence from human protein Suv3 to the N-terminus of proteins. It has been shown that 40 amino acids long N-terminal sequence is enough for effective transport of Suv3 through the mitochondrial membrane. [21]. First the efficiency of the process will be assessed by detection of Red Fluorescent Protein with confocal microscopy. The red fluorescence should be visible in the mitochondria of the cells invaded by bacteria expressing RFP fused with mitochondrial leader sequence. Then the ability to induce apoptosis by mitochondrially directed p53 will be investigated. Apoptosis induction will be confirmed by flow cytometry [8]. In addition we will use confocal microscopy to monitor the process. In both cases cells will be stained with propidium iodide and annexin V.
REFERENCES
|
 "
"