Team:MoWestern Davidson/project design
From 2009.igem.org
(→Predicting Effects of Suppressor tRNAs) |
(→Predicting Effects of Suppressor tRNAs) |
||
| Line 37: | Line 37: | ||
One of the most critical factors in choosing which suppressor tRNAs to use is the prevalence in the normal ''E. coli'' genome of the mRNA codon that the tRNA targets. In larger SAT problems, we do not want to use suppressor tRNAs whose mRNA codons occur relatively frequently in the reading frame of coding sequences, especially if the coding sequence is a gene for an essential protein in the cell. We wanted to obtain bioinformatic data to predict how much effect introducing suppressor tRNAs may have on ''E. coli''. We also chose to first construct a 1-SAT problem in ''E. coli'' - having a single 5-bp codon addition before our reporter gene, and its appropriate suppressor tRNA. We tested the effect of each suppressor tRNA separately to obtain biological data on the effects of these tRNAs. | One of the most critical factors in choosing which suppressor tRNAs to use is the prevalence in the normal ''E. coli'' genome of the mRNA codon that the tRNA targets. In larger SAT problems, we do not want to use suppressor tRNAs whose mRNA codons occur relatively frequently in the reading frame of coding sequences, especially if the coding sequence is a gene for an essential protein in the cell. We wanted to obtain bioinformatic data to predict how much effect introducing suppressor tRNAs may have on ''E. coli''. We also chose to first construct a 1-SAT problem in ''E. coli'' - having a single 5-bp codon addition before our reporter gene, and its appropriate suppressor tRNA. We tested the effect of each suppressor tRNA separately to obtain biological data on the effects of these tRNAs. | ||
<br/> | <br/> | ||
| - | [[Image:NoOccurences.png|500px]] | + | <center>[[Image:NoOccurences.png|500px]]</center> |
==Designing the Frameshift Mutation Leaders and Suppressor tRNAs== | ==Designing the Frameshift Mutation Leaders and Suppressor tRNAs== | ||
Revision as of 22:11, 28 July 2009

|
Introduction
Modularity... more time on one side where less building more interpreting. More time where more building and less interpreting.
To solve the SAT problem, we constructed reporter genes to express a specific phenotype if the E. coli solves the problem. To design these reporter genes, we constructed a continuum that models different levels of bacterial automation for computation. This allowed us to select the level of automation we desired and design reporter genes based off of that level. Engineering the reporter genes involved inserting a 5 base pair insertion that shifts the reading frame in translation, causing a nonsense protein. We then provided the cells with different suppressor tRNAs that may or may not suppress this insertion. If the insertion is suppressed, the reading frame will be restored and the protein will be expressed. These suppressor tRNAs represent the inputs of the SAT problem, and the 5 base pair insertions represent the clauses of the problem. In most designs the frame shift leader (FSL), which includes a start codon and the 5 base pair insertion, occurs at the beginning of the protein structure. Note that for every point on the bacterial automation scale, there are three inputs, representing the number of variables being evaluated for the particular SAT problem.
Automation Scale and FSL Design
1. Single Literal: In this approach, the reporter gene has a FSL of one literal, or 5 base pair insertion. The bacteria are given a set of tRNA variables as inputs to evaluate that literal, and if the bacteria receive that literal, then it will express a gene. This process is more of a screening to see if the design works. To evaluate the logical clauses and MAX SAT, we must look plates to see if the gene was expressed.
The design of the FSL in the reporter genes in this example is relatively simple. Immediately after the start codon, ATG, we insert a single the 5 base pair insertion. Different pairings of inputs, tRNA suppressors, and 5-mer reporter genes are mixed.
In the diagram shown, each column represents a clause and each row or literal represents an input
2. Single Clause: In this approach, the reporter gene has a FSL of one logical clause (a OR b), consisting of two 5 base pair insertions. Bacteria are given a set of tRNA variables as inputs to evaluate that clause. If the clause is satisfied and suppression occurs, then the gene will be expressed. In order to compute the problem we must then determine how many colonies expressed the gene, thus satisfying the clauses.
In this design, the FSL has two 5 base pair insertions. If one tRNA binds to either insertion, the reading frame is restored. However, these insertions are designed in such a manner that if one tRNA binds, another tRNA could not bind to the second 5 base pair insertion.
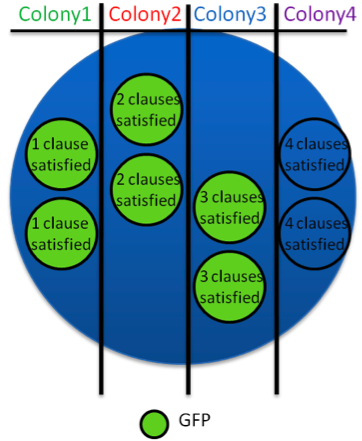
3. Automated Population with One Reporter Type: This approach uses 4 different FSLs ranging from 1 to 4 clauses (a or b) in the same reporter gene. These reporter genes are then divided into colonies representing the number of clauses (1, 2, 3, or 4) in their reporter gene. Bacteria are given a set of tRNA variables as inputs and evaluate the logical clauses and MAX SAT by reporting the gene expression. This set up allows us to determine the maximum number of clauses the tRNA is able to solve.
The FSL length and design varies according to the number of clauses inserted. However, the clauses are designed in the same way as in the previous single clause line. These single clauses are then strung together so that in order for the proper reading frame to be restored, exactly one 5 base pair insertion in each clause must be satisfied.
4. Automated Population with Individual Reporter Types: This approach is basically the same as the previous, except that the each clause has its own reporter gene. For example, the first clone tests for 1 clause satisfied and if satisfied, the clone will produce GFP. The second clone tests for 2 clauses satisfied and will produce RFP if satisfied. The third clone tests for 3 clauses satisfied and will produce Chloramphenicol resistance. The last clone tests for 4 clauses satisfied and will produce Tetracycline resistance.
5. Fully Automated Population with Individual Reporter Type in a Single Clone: In this approach, 4 different FSLs that test for at least 1, 2, 3, and 4 clauses satisfied are inserted in the beginning of different reporter genes used in a single clone. The first reporter gene tests for at least 1 clause satisfied, and if satisfied, the gene GFP will be expressed. The second reporter gene tests for at least 2 clauses satisfied and will express RFP if satisfied. The third reporter gene tests for at least 3 clauses satisfied and will produce Chloramphenicol resistance. Tetracycline resistance is the last reporter gene that tests for all 4 clauses satisfied. Each FSL design has the same SAT problem, (a OR b) AND (b OR c’) for example, encoded in it. Bacteria are given a set of tRNA variables as inputs and evaluate the logical clauses and MAX SAT.
Predicting Effects of Suppressor tRNAs
One of the most critical factors in choosing which suppressor tRNAs to use is the prevalence in the normal E. coli genome of the mRNA codon that the tRNA targets. In larger SAT problems, we do not want to use suppressor tRNAs whose mRNA codons occur relatively frequently in the reading frame of coding sequences, especially if the coding sequence is a gene for an essential protein in the cell. We wanted to obtain bioinformatic data to predict how much effect introducing suppressor tRNAs may have on E. coli. We also chose to first construct a 1-SAT problem in E. coli - having a single 5-bp codon addition before our reporter gene, and its appropriate suppressor tRNA. We tested the effect of each suppressor tRNA separately to obtain biological data on the effects of these tRNAs.

Designing the Frameshift Mutation Leaders and Suppressor tRNAs
We employed two separate approaches to design and construct our suppressor tRNAs and the frameshifted reporter proteins. The genomic sequence of the novel five nucleotide anticodon tRNAs were obtained from papers by Thomas J. Magliery, J. Christopher Anderson and Peter G. Schultz {Magliery et. al. & Anderson et. al.}.
We used synthesized single-stranded oligonucleotides to assemble the twelve suppressor tRNAs with flanking BioBrick prefix and suffix sticky ends. For more information on our tRNA designs, click here.
The reporter genes were designed and constructed by PCR-directed mutagenesis in order to add a frameshift mutation leader (FSL) to the beginning of the reporter gene sequence. For more information on our FSL designs click here.
 "
"