Team:EPF-Lausanne/Results/EDS
From 2009.igem.org
We run an equilibration of 80ns using 32 processors on the dark state (2v0u). It took a month of computation to have this result.
Here is a movie over the trajectory file.
Dark State
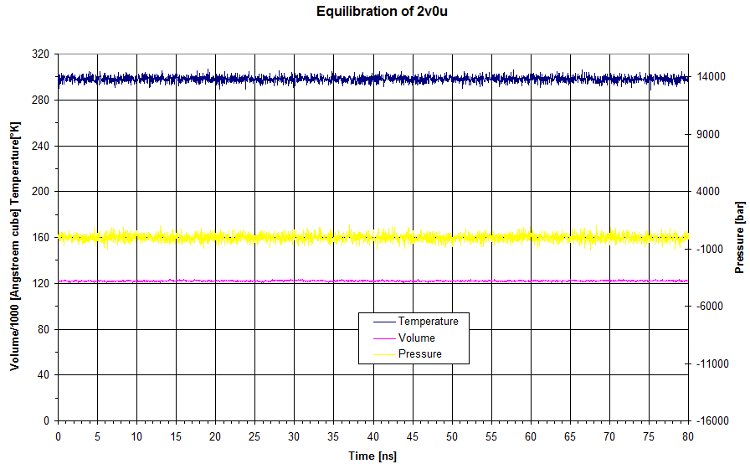
Here we look at the output to check input parameters.
The raw data for the equilibration match what we set for the NPT. Pressure and temperature are kept constant using namd dynamic. The volume is quite constant as well.
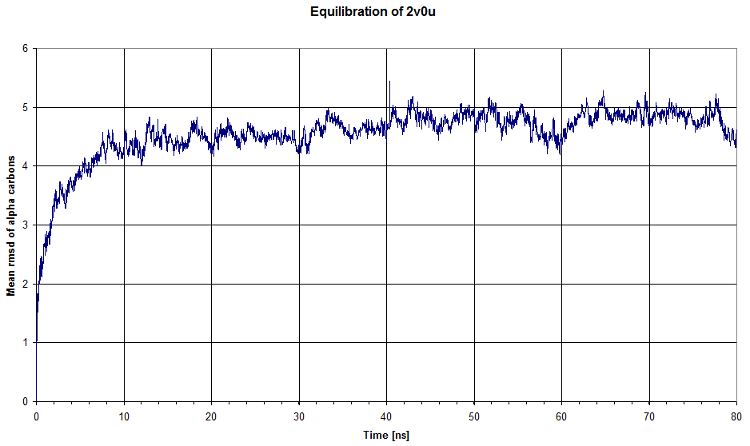
Then we computed the evolution of the rmsd compared to the first timestep of equilibration. We see that there is a plateau after ~40ns, which means that our system's energy is reaching a minimum. That's clearly what we expected.
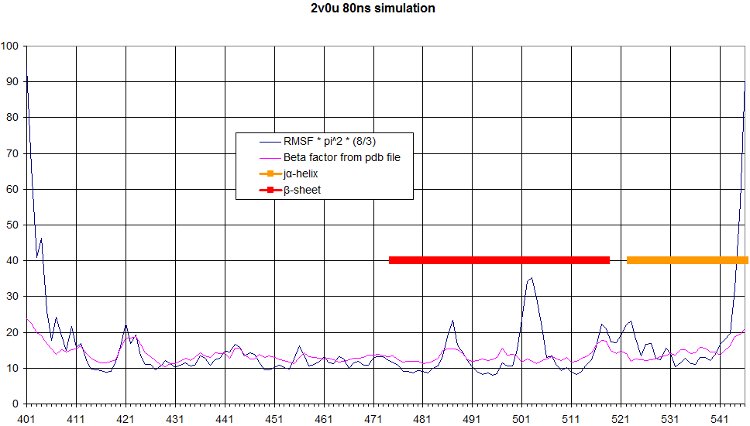
The comparison of the RMSF to the beta factor measured during crystallography of all alpha carbons over the simulation is a nice validation of our simulation. We get quite similar curves, with some differences at one end of the protein. We see in the movie that this part moves a lot.
Here we computed the oscillation of RMSF in function of residue number, and we highlight the interesting part of our protein, namely the beta sheet and the alpha helix.
Analysis of the simulation
We have organized our analysis on 2 main ideas:
- Find a structural change in the Jα helix based on the simulation using namd.
- Find residues showing different comportment in dark and light state
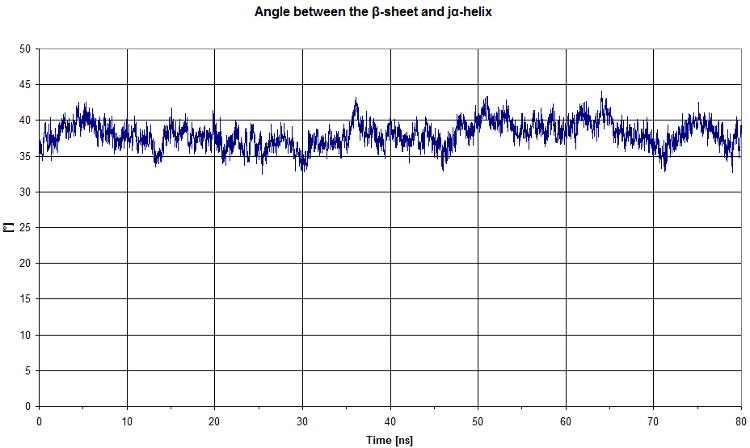
First, we start by looking at the angle between the beta sheet and the Jα helix.
Click here to see the code used in VMD to get angle data
We get a quite constant value. It will be more interesting to compare this graph to the light state.
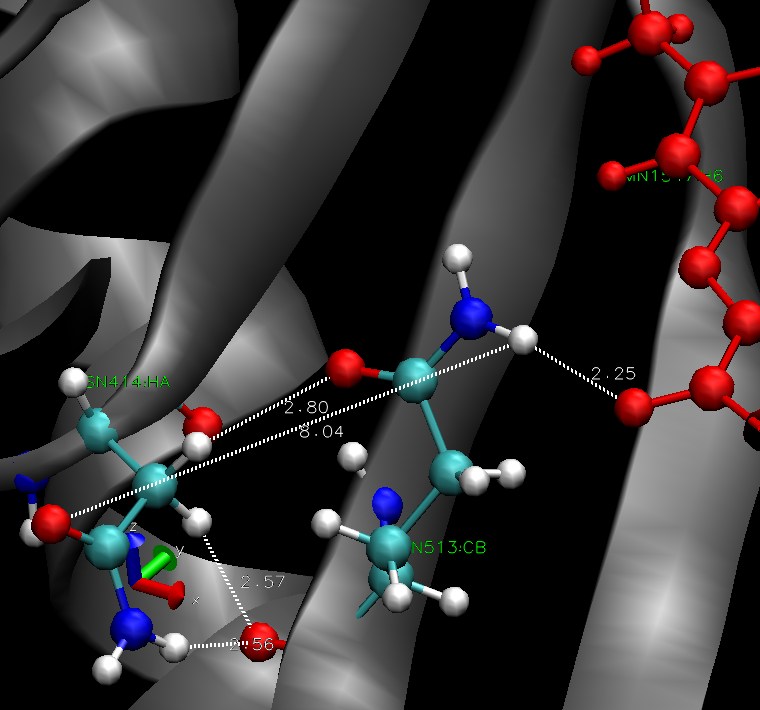
The Jα helix is stabilized by h-bonds to the beta sheet. These bonds are supposed to be disrupted by the conformational change in the dark state. The residues 513 seems to be involved in stabilisation of FMN through hydrogen bonds. We hope it is linked to beta sheet, more precisely the residue 414. There is a picture of the situation, residue ASN 414 is on the left (beta sheet), GLN513 in the middle and the FMN is in red. All the hydrogen bonds we investigated over the simulation are pictured.
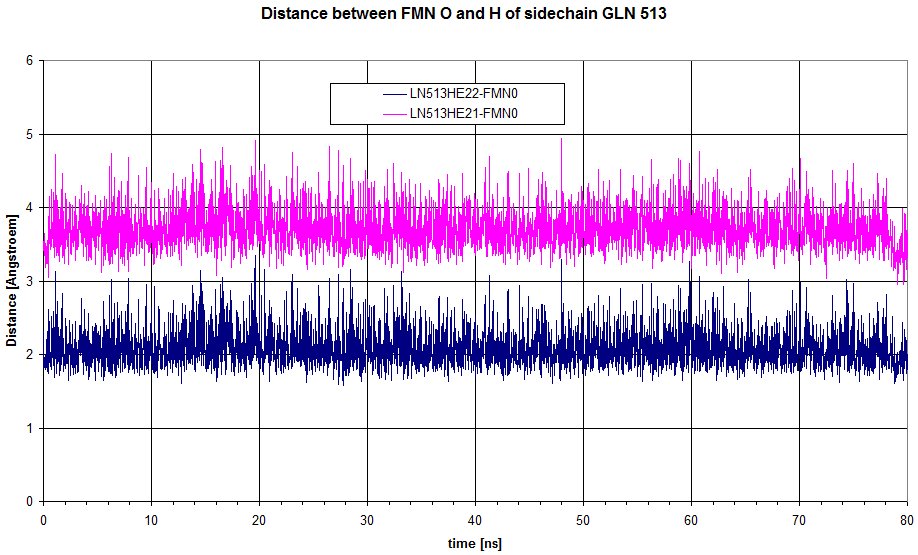
Here is a plot of the distance between the 2 hydrogens from sidechain of GLN513 to the oxygen of FMN. HE22 is definitely involved in an hydrogen bond, but doesn't move enough to loose the interaction.
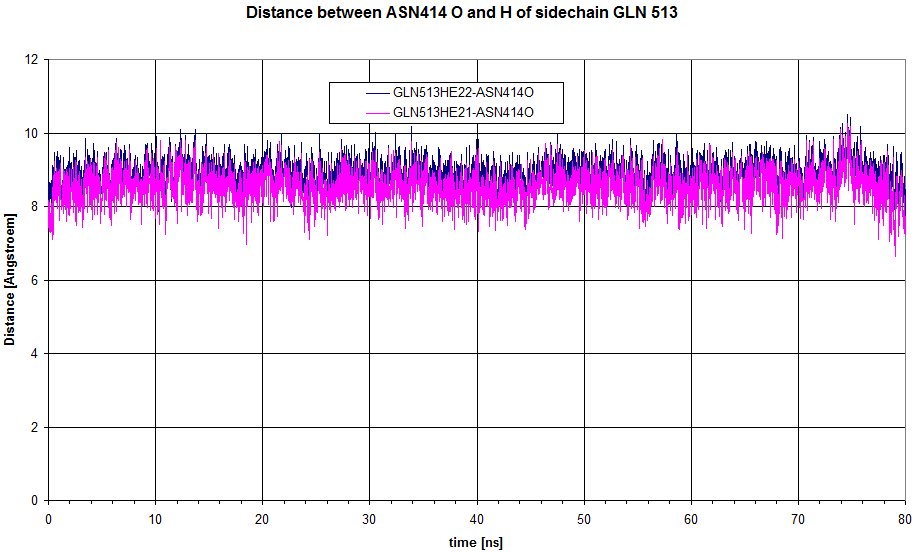
We can have a look at the distance between O of sidechain of ASN 414 to Hs of GLN513.
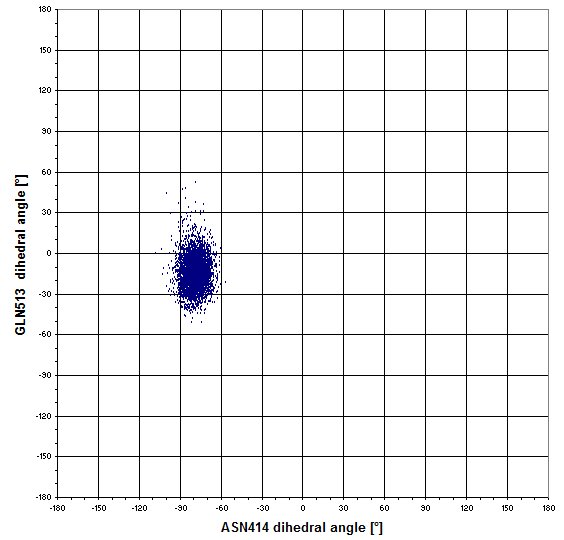
Maybe the sidechains of GLN513 moves in regard the the position of ASN414? -> no, there is a single combination on the next graph.
An interesting residue to study in the dark state is the residue n° 450, which is the cystein that reacts with the cofactor. Here we plot the dihedral angle of this residue to see how many time the cystein point toward the FMN:
Here we plot the distance between the sulfur atom of the cystein and the FMN carbon which is attacked by the sulfur atom upon light activation:
Here is the distance:
Some useful distances
The Jα helix is anchored to the β structure by two H-bond networks the first involving Lys533 (Jα) and the couple Glu475, Gln497 (β structure), the second involving Lys413 (Jα) and Thr535 (β structure). Here we plot the H-bond distances (two in the former and one in the latter).
- Bond between Gln497 and Lys533 in dark state
- Bond between Gln475 and Lys533 in dark state
 "
"