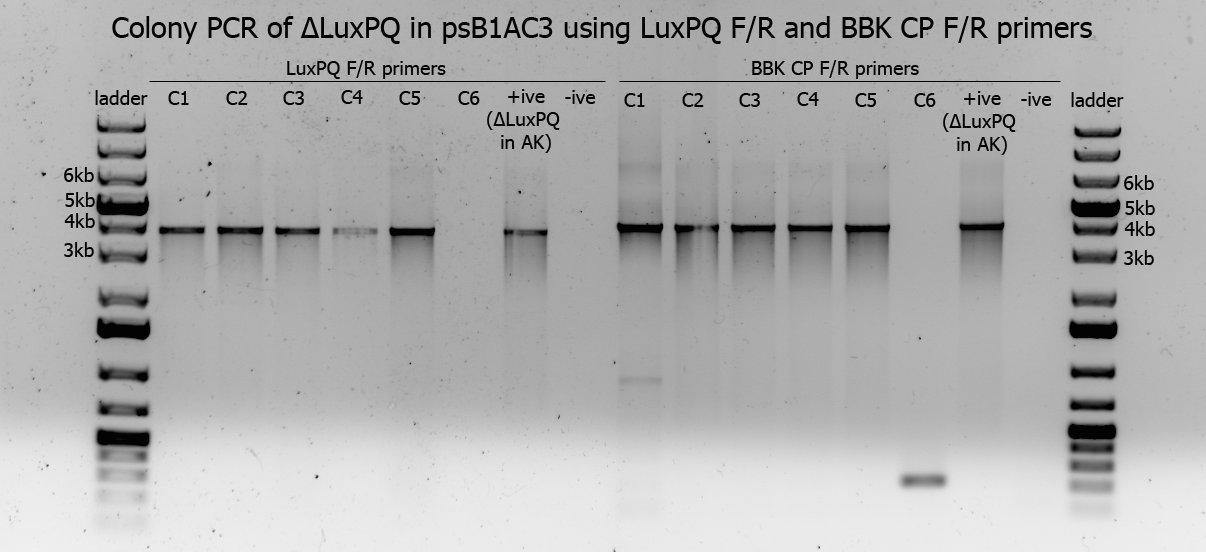|
|
| (2 intermediate revisions not shown) |
| Line 457: |
Line 457: |
| | | | |
| | Positive control = LuxPQ-B0015-R0040-LuxOU-B0015 in AK3 | | Positive control = LuxPQ-B0015-R0040-LuxOU-B0015 in AK3 |
| - | <br>
| |
| | <br> | | <br> |
| | PCR conditions | | PCR conditions |
| Line 535: |
Line 534: |
| | | | |
| | Result: the only bands that appeared on the gel were those of the ladders. Therefore, start new construction. | | Result: the only bands that appeared on the gel were those of the ladders. Therefore, start new construction. |
| - |
| |
| - | <br>
| |
| | <br> | | <br> |
| | <br> | | <br> |
| Line 733: |
Line 730: |
| | </table> | | </table> |
| | </html> | | </html> |
| - | <br>
| |
| | <br> | | <br> |
| | [[Image:Calgary_PQplasmid_switch_AC.png|700px]] | | [[Image:Calgary_PQplasmid_switch_AC.png|700px]] |
| - |
| |
| | <br> | | <br> |
| | Isolating plasmid from LuxPQ-B0015-R0040-LuxOU-B0015 (AC) and LuxPQ (in AC) | | Isolating plasmid from LuxPQ-B0015-R0040-LuxOU-B0015 (AC) and LuxPQ (in AC) |
| Line 847: |
Line 842: |
| | </html> | | </html> |
| | | | |
| - | <br>
| |
| - | <br>
| |
| | <br> | | <br> |
| | Construction of LuxPQ-B0015-R0040-LuxOU-B0015 by inserting B0015-R0040-LuxOU-B0015 (AK) into LuxPQ (in AC) | | Construction of LuxPQ-B0015-R0040-LuxOU-B0015 by inserting B0015-R0040-LuxOU-B0015 (AK) into LuxPQ (in AC) |
| | <br> | | <br> |
| - | <br>
| + | Purpose: To construct LuxPQ-B0015-R0040-LuxOU-B0015 by inserting B0015-R0040-LuxOU-B0015 (AK) into LuxPQ (in AC). This Restriction digest was set up overnight with the following sets of enzymes: XbaI and PstI (insert) and SpeI/PstI (recipient). |
| - | Purpose: To construct LuxPQ-B0015-R0040-LuxOU-B0015 by inserting B0015-R0040-LuxOU-B0015 (AK) into LuxPQ (in AC) | + | |
| - | <br>
| + | |
| - | Protocol
| + | |
| - | <br>
| + | |
| - | Recipient
| + | |
| - | <html>
| + | |
| - | <table x:str border=0 cellpadding=0 cellspacing=0 width=731 style='border-collapse:
| + | |
| - | collapse;table-layout:fixed;width:549pt'>
| + | |
| - | <col width=198 style='mso-width-source:userset;mso-width-alt:7241;width:149pt'>
| + | |
| - | <col width=205 style='mso-width-source:userset;mso-width-alt:7497;width:154pt'>
| + | |
| - | <col width=264 style='mso-width-source:userset;mso-width-alt:9654;width:198pt'>
| + | |
| - | <col width=64 style='width:48pt'>
| + | |
| - | <tr height=22 style='height:16.5pt;mso-yfti-firstrow:yes;mso-yfti-irow:0'>
| + | |
| - | <td height=22 class=xl25 width=198 style='height:16.5pt;width:149pt'>Recipient
| + | |
| - | 1</td>
| + | |
| - | <td class=xl26 width=205 style='width:154pt'>Recipient 2</td>
| + | |
| - | | + | |
| - | <td class=xl24 width=264 style='width:198pt'></td>
| + | |
| - | <td class=xl24 width=64 style='width:48pt'></td>
| + | |
| - | </tr>
| + | |
| - | <tr height=22 style='height:16.5pt;mso-yfti-irow:1'>
| + | |
| - | <td height=22 class=xl27 width=198 style='height:16.5pt;width:149pt'>2μL
| + | |
| - | REact 4</td>
| + | |
| - | <td class=xl28 width=205 style='width:154pt'>2μL REact 4</td>
| + | |
| - | <td colspan=2 class=xl24 style='mso-ignore:colspan'></td>
| + | |
| - | | + | |
| - | </tr>
| + | |
| - | <tr height=22 style='height:16.5pt;mso-yfti-irow:2'>
| + | |
| - | <td height=22 class=xl27 width=198 style='height:16.5pt;width:149pt'>0.75
| + | |
| - | μL SpeI</td>
| + | |
| - | <td class=xl28 width=205 style='width:154pt'>0.75 μL SpeI</td>
| + | |
| - | <td colspan=2 class=xl24 style='mso-ignore:colspan'></td>
| + | |
| - | </tr>
| + | |
| - | <tr height=22 style='height:16.5pt;mso-yfti-irow:3'>
| + | |
| - | | + | |
| - | <td height=22 class=xl27 width=198 style='height:16.5pt;width:149pt'>0.75
| + | |
| - | μL PstI</td>
| + | |
| - | <td class=xl28 width=205 style='width:154pt'>0.75 μL PstI</td>
| + | |
| - | <td colspan=2 class=xl24 style='mso-ignore:colspan'></td>
| + | |
| - | </tr>
| + | |
| - | <tr height=43 style='height:32.25pt;mso-yfti-irow:4'>
| + | |
| - | <td height=43 class=xl27 width=198 style='height:32.25pt;width:149pt'>2
| + | |
| - | μL of ∆LuxPQ in AK3 C2 [112.6ng/μL]</td>
| + | |
| - | | + | |
| - | <td class=xl28 width=205 style='width:154pt'>2 μL of ∆LuxPQ in AK3
| + | |
| - | C5 [115.3ng/μL]</td>
| + | |
| - | <td colspan=2 class=xl24 style='mso-ignore:colspan'></td>
| + | |
| - | </tr>
| + | |
| - | <tr height=22 style='height:16.5pt;mso-yfti-irow:5;mso-yfti-lastrow:yes'>
| + | |
| - | <td height=22 class=xl27 width=198 style='height:16.5pt;width:149pt'>14.5
| + | |
| - | μL ddH2O</td>
| + | |
| - | <td class=xl28 width=205 style='width:154pt'>14.5 μL ddH2O</td>
| + | |
| - | | + | |
| - | <td colspan=2 class=xl24 style='mso-ignore:colspan'></td>
| + | |
| - | </tr>
| + | |
| - | <tr height=21 style='height:15.75pt'>
| + | |
| - | <td height=21 colspan=4 class=xl24 style='height:15.75pt;mso-ignore:colspan'></td>
| + | |
| - | </tr>
| + | |
| - | <tr height=21 style='height:15.75pt'>
| + | |
| - | <td height=21 colspan=4 class=xl24 style='height:15.75pt;mso-ignore:colspan'></td>
| + | |
| - | </tr>
| + | |
| - | <tr height=21 style='height:15.75pt'>
| + | |
| - | | + | |
| - | <td height=21 colspan=4 class=xl24 style='height:15.75pt;mso-ignore:colspan'></td>
| + | |
| - | </tr>
| + | |
| - | <tr height=21 style='height:15.75pt'>
| + | |
| - | <td height=21 colspan=4 class=xl24 style='height:15.75pt;mso-ignore:colspan'></td>
| + | |
| - | </tr>
| + | |
| - | <tr height=21 style='height:15.75pt'>
| + | |
| - | <td height=21 colspan=4 class=xl24 style='height:15.75pt;mso-ignore:colspan'></td>
| + | |
| - | </tr>
| + | |
| - | <![if supportMisalignedColumns]>
| + | |
| - | | + | |
| - | <tr height=0 style='display:none'>
| + | |
| - | <td width=198 style='width:149pt'></td>
| + | |
| - | <td width=205 style='width:154pt'></td>
| + | |
| - | <td width=264 style='width:198pt'></td>
| + | |
| - | <td width=64 style='width:48pt'></td>
| + | |
| - | </tr>
| + | |
| - | <![endif]>
| + | |
| - | </table>
| + | |
| - | </html>
| + | |
| - | <br>
| + | |
| - | <br>
| + | |
| - | Insert
| + | |
| - | <html>
| + | |
| - | <table x:str border=0 cellpadding=0 cellspacing=0 width=731 style='border-collapse:
| + | |
| - | collapse;table-layout:fixed;width:549pt'>
| + | |
| - | <col width=198 style='mso-width-source:userset;mso-width-alt:7241;width:149pt'>
| + | |
| - | <col width=205 style='mso-width-source:userset;mso-width-alt:7497;width:154pt'>
| + | |
| - | <col width=264 style='mso-width-source:userset;mso-width-alt:9654;width:198pt'>
| + | |
| - | <col width=64 style='width:48pt'>
| + | |
| - | <tr height=22 style='height:16.5pt;mso-yfti-firstrow:yes;mso-yfti-irow:0'>
| + | |
| - | <td height=22 class=xl25 width=198 style='height:16.5pt;width:149pt'>2μL
| + | |
| - | REact 2</td>
| + | |
| - | <td class=xl27 width=205 style='width:154pt'></td>
| + | |
| - | | + | |
| - | <td class=xl24 width=264 style='width:198pt'></td>
| + | |
| - | <td class=xl24 width=64 style='width:48pt'></td>
| + | |
| - | </tr>
| + | |
| - | <tr height=22 style='height:16.5pt;mso-yfti-irow:1'>
| + | |
| - | <td height=22 class=xl26 width=198 style='height:16.5pt;width:149pt'>0.75
| + | |
| - | μL XbaI</td>
| + | |
| - | <td class=xl27></td>
| + | |
| - | <td colspan=2 class=xl24 style='mso-ignore:colspan'></td>
| + | |
| - | </tr>
| + | |
| - | | + | |
| - | <tr height=22 style='height:16.5pt;mso-yfti-irow:2'>
| + | |
| - | <td height=22 class=xl26 width=198 style='height:16.5pt;width:149pt'>0.75
| + | |
| - | μL PstI</td>
| + | |
| - | <td class=xl27></td>
| + | |
| - | <td colspan=2 class=xl24 style='mso-ignore:colspan'></td>
| + | |
| - | </tr>
| + | |
| - | <tr height=64 style='height:48.0pt;mso-yfti-irow:3'>
| + | |
| - | <td height=64 class=xl26 width=198 style='height:48.0pt;width:149pt'>2
| + | |
| - | μL of LuxPQ-B0015-R0040-LuxOU-B0015 in AK3 C1 [156.3ng/μL]</td>
| + | |
| - | | + | |
| - | <td class=xl27></td>
| + | |
| - | <td colspan=2 class=xl24 style='mso-ignore:colspan'></td>
| + | |
| - | </tr>
| + | |
| - | <tr height=22 style='height:16.5pt;mso-yfti-irow:4;mso-yfti-lastrow:yes'>
| + | |
| - | <td height=22 class=xl26 width=198 style='height:16.5pt;width:149pt'>12.5
| + | |
| - | μL ddH2O</td>
| + | |
| - | <td class=xl27></td>
| + | |
| - | <td colspan=2 class=xl24 style='mso-ignore:colspan'></td>
| + | |
| - | </tr>
| + | |
| - | | + | |
| - | <tr height=21 style='height:15.75pt'>
| + | |
| - | <td height=21 colspan=2 class=xl27 style='height:15.75pt;mso-ignore:colspan'></td>
| + | |
| - | <td colspan=2 class=xl24 style='mso-ignore:colspan'></td>
| + | |
| - | </tr>
| + | |
| - | <tr height=21 style='height:15.75pt'>
| + | |
| - | <td height=21 colspan=4 class=xl24 style='height:15.75pt;mso-ignore:colspan'></td>
| + | |
| - | </tr>
| + | |
| - | <tr height=21 style='height:15.75pt'>
| + | |
| - | <td height=21 colspan=4 class=xl24 style='height:15.75pt;mso-ignore:colspan'></td>
| + | |
| - | | + | |
| - | </tr>
| + | |
| - | <tr height=21 style='height:15.75pt'>
| + | |
| - | <td height=21 colspan=4 class=xl24 style='height:15.75pt;mso-ignore:colspan'></td>
| + | |
| - | </tr>
| + | |
| - | <tr height=21 style='height:15.75pt'>
| + | |
| - | <td height=21 colspan=4 class=xl24 style='height:15.75pt;mso-ignore:colspan'></td>
| + | |
| - | </tr>
| + | |
| - | <tr height=21 style='height:15.75pt'>
| + | |
| - | <td height=21 colspan=4 class=xl24 style='height:15.75pt;mso-ignore:colspan'></td>
| + | |
| - | | + | |
| - | </tr>
| + | |
| - | <![if supportMisalignedColumns]>
| + | |
| - | <tr height=0 style='display:none'>
| + | |
| - | <td width=198 style='width:149pt'></td>
| + | |
| - | <td width=205 style='width:154pt'></td>
| + | |
| - | <td width=264 style='width:198pt'></td>
| + | |
| - | <td width=64 style='width:48pt'></td>
| + | |
| - | </tr>
| + | |
| - | <![endif]>
| + | |
| - | | + | |
| - | </table>
| + | |
| - | </html>
| + | |
| - | <br>
| + | |
| - | <br>
| + | |
| - | Put in the incubator at 37ºC overnight.
| + | |
| | | | |
| | | | |
| Line 1,181: |
Line 1,012: |
| | <br> | | <br> |
| | <div class="heading"> | | <div class="heading"> |
| - | Descriptive Title of What You're Doing
| + | Changing Up the Wiki |
| | </div> | | </div> |
| | <br> | | <br> |
| | <div class="desc"> | | <div class="desc"> |
| | </html> | | </html> |
| - | WIKI CODING HERE
| + | Did regular update of blog onto different sections of the page, so that will keep being updated and it’s great that everybody has been keeping up with posting on the required days. |
| | + | <br><br> |
| | + | I fixed up the notebook template so it can be accessed easily from the header menu, and it now applies to all team members (not just the lab people). I’ve also edited the instructions for posting in it to include some more fundamental formatting requirements (with many helpful suggestions from Jamie). |
| | + | <br><br> |
| | + | TO DO: |
| | + | *more blog update inclusion |
| | + | *Edit Fahd’s news story about NUTV |
| | + | *Post video of NUTV |
| | + | *Edit Sponsor page |
| | + | *Teach Prima how to edit Acknowledgement page |
| | + | *edit content already there |
| | + | *add more to intros for human practices and second life |
| | + | <html> |
| | + | </div> |
| | + | <div class="heading"> |
| | + | Completion of DNA Extraction Activity in Second Life |
| | + | </div> |
| | + | <div class="desc"> |
| | + | </html> |
| | + | The DNA extraction activity is finally done, and it works! (I think). This officially makes me the slowest scripter ever D:< . I am still thinking of a way to make it easier to do. It is intuitive in that it gives you direct instructions, but right now, people can kind of jump in to the middle of it without doing the first bit (which I’m not sure is a good thing – it IS a long activity after all). |
| | + | <br><br> |
| | + | Following the DNA extraction activity, I’ve completed most of the instructions/description for the third lab mission. Tomorrow, I will be checking with Katie to make sure the alternative components fit in both the PCR and restriction digest activities to give the desired products (eg. PCR to make a linearized biobricked part). We will also be changing notecards and mission instructions to reflect the changes to these activities. |
| | + | <br><br> |
| | + | Speaking of notecards, I’m editing the one I wrote for DNA extraction. I’ve started writing up the instructions for the first lab mission (bacterial transformation) – it’s a ridiculous task (INVISIBLE BACTERIA :D), but I thought it would be fun. This was inspired by seeing Katie make a thing you can wear on your avatar to make you mostly invisible- we might be able to give something like that as a prize? |
| | | | |
| | <html> | | <html> |
| | + | <br> |
| | </div> | | </div> |
| | </td> | | </td> |
|
|
CAROL
Finishing Modeling Paper
- Finished writing modeling paper that summarizes what we will be doing once the circuit is completed. The experiments should not take long to accomplished and these results can be tabulated in Matlab quickly.
- Plated the week-long ligation of the promoter library in both XL Gold Ultra competent cells and Top 10 cells, but there is no growth.
- A reason that we thought of that might have caused minimal colonies is the phosphatase step. The 32 primers that we have synthesized does not have 5' phosphates. If we phosphatase the vector, that might result in no ligation.
- Re-started restriction enzyme digestion on vector pCS26 with XhoI and BamHI at 37oC for 4 hours. Then I ligated the promoter with the vector with Quick Ligase for 5 minutes. Then I transformed the plasmids into both XL Gold Ultracompetent cells and Top 10 cells. Incubate at 37oC for 16 hours.
|
|
|
CHINMOYEE
Descriptive Title of What You're Doing
|
|
|
EMILY
Construction of J13002-LuxOD47E-B0015
- Did a construction digest of the J13002-LuxOD47E construct and B0015 terminator with XbaI/ PstI/ SpeI enzymes.
- Transformation into TOP10 cells and plated on AC plates.
- Hopefully things will go smoothly from here and my circuit will be fully-contructed by Thursday!
Ethics/ Outreach
- Today I talked with Fahd, Stefan and Mandy about our Ethics paper as well as our conference. Mandy has a location in mind and she want Fahd and I to log into Second Life and explore the ethics area a bit. Mandy had a chance to look at our paper first thing this morning and she made some good suggestions for areas to add on to the introduction, bringing in some of the stuff that we learned about in the webinar. I spent a large part of today re-writing that part and I will have a copy of the rough draft of our paper by tonight!
|
|
|
FAHD
Marketing and Ethics for July 27th 2009
Today, I concentrated my energies at marketing our iGEM project and looking at the ethical aspect of our project. Here is what I did today:
1) I got a call from David Corriveau. He is the vice president for engineering and devlopment at Albian Sands Energy Inc. He was extremely impressed by our work and wanted to know more about the possible bio-remediation aspect of our project. I sent him an e-mail detailing about our project this year. I also asked Patrick to write me an attractive paragraph about second life and I also wrote up a para about ethics and the potential applications of our project. I combined all this information into a iGEM package which I ran thru some of my team members and then sent it off (e-mail).
2) I helped emily with the ethics paper; I helped her write the Bioterrorism and the Preactionary Framework.
3) I had contacted Ron Hall on Friday. Ron Hall is a UofC Alumni and currently works at Focus Corporation (O&G company, geomatics). Today I sent him a sponsorship package, our june's newsletter and our proposal and i will follow up with him this week.
4) I downloaded our iGEM NUTV news coverage and converted into a suitable format for everyone to view. I also wrote up description for our iGEM NUTV coverage for our WIKI.
5) I discussed fundraising ideas with my team mates.
6) I did Reasearch on the following comapnies:
-Fraiser Milner Casgrain LLP
-Graymont Western Canada Inc.
7) I wrote a rejection thank you letter for APotex Canada and asked them if i could have some contacts who would be able to sponsor us.
|
|
|
IMAN
Descriptive Title of What You're Doing
|
|
|
JAMIE
Descriptive Title of What You're Doing
|
|
|
JEREMY
Colony PCR of LuxPQ-B0015-R0040-LuxOU-B0015 construct in psB1AC3 using BBK CP F/R and LuxPQ F/LuxOU R primers
Colony PCR of LuxPQ-B0015-R0040-LuxOU-B0015 construct in psB1AC3 using BBK CP F/R and LuxPQ F/LuxOU R primers
Purpose: To verify the presence of LuxPQ-B0015-R0040-LuxOU-B0015 construct in psB1AC3.
Protocol
| |
MM1 (6x) (μL) |
MM2 (6x) (μL) |
| 10X PCR buffer minus MgCl2 |
30 |
30 |
| 10mM dNTPs |
6 |
6 |
| 50mM MgCl2 |
9 |
9 |
| F primer |
6 (BBK CP F) |
6 (LuxPQ F) |
| R primer |
6 (BBK CP R) |
6 (LuxOU R) |
| ddH2O |
241.8 |
241.8 |
| pTaq |
1.2 |
1.2 |
|
|
|
Positive control = LuxPQ-B0015-R0040-LuxOU-B0015 in AK3
PCR conditions
| # of cycles |
Temp (ºC) |
Time |
| 1 |
94 |
6 min |
| 36 |
94 |
30sec |
| 55 |
45sec |
| 72 |
6min 20sec |
| 1 |
72 |
10min |
| |
4 |
hold |
|
|
|
Result: the only bands that appeared on the gel were those of the ladders. Therefore, start new construction.
Colony PCR of ∆LuxPQ in psB1AC3 using BBK CP F/R and LuxPQ F/R primers
Purpose: To verify the presence of ∆LuxPQ in psB1AC3 (to verify a successful plasmid switch)
Protocol
| |
MM1 (8x) (μL) |
MM2 (6x) (μL) |
| 10X PCR buffer minus MgCl2 |
40 |
40 |
| 10mM dNTPs |
8 |
8 |
| 50mM MgCl2 |
12 |
12 |
| F primer |
8 (LuxPQ F) |
8 (BBK CP F) |
| R primer |
8 (LuxPQ R) |
8 (BBK CP R) |
| ddH2O |
322.4 |
322.4 |
| pTaq |
1.6 |
1.6 |
|
|
|
Positive control = ∆LuxPQ in psB1AK3
PCR conditions
| # of cycles |
Temp (ºC) |
Time |
|
| 1 |
94 |
6 min |
|
| 36 |
94 |
30sec |
|
| 55 |
45sec |
|
| 72 |
6min 20sec |
|
| 1 |
72 |
10min |
|
| |
4 |
hold |
|
|
|
|
|
|
|
|
|
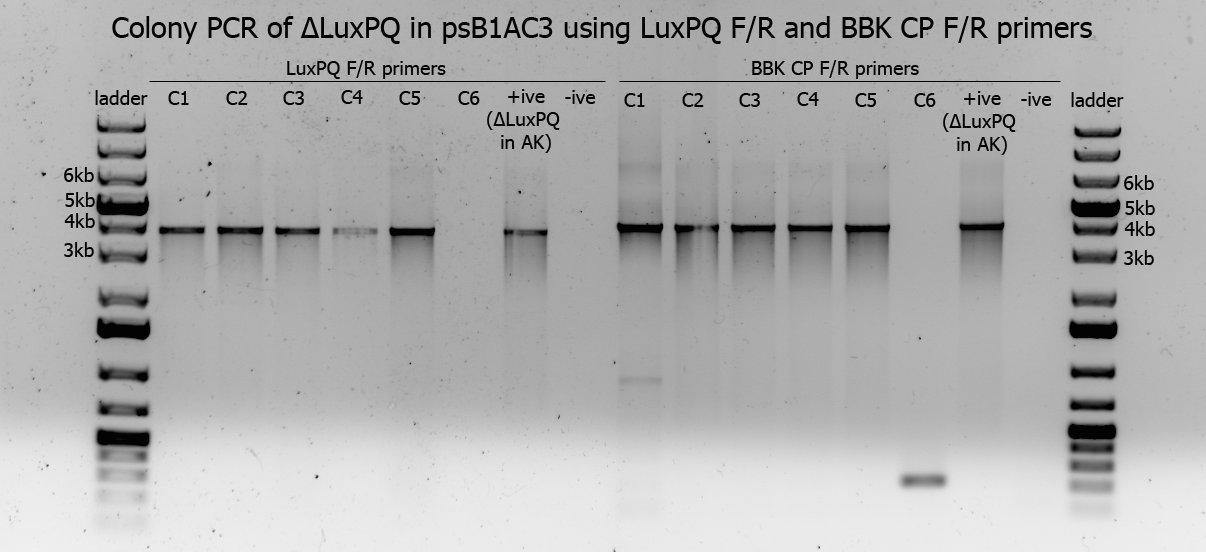
Isolating plasmid from LuxPQ-B0015-R0040-LuxOU-B0015 (AC) and LuxPQ (in AC)
Purpose: isolate and measure concentrations of pure plasmids.
| DNA |
260/280 |
260/230 |
Conc. [ng/μL] |
| ∆LuxPQ
in AC3 C1 |
1.83 |
2.23 |
146.8 |
| ∆LuxPQ
in AC3 C2 |
1.81 |
2.19 |
112.6 |
| ∆LuxPQ
in AC3 C3 |
1.8 |
2.27 |
145.8 |
| ∆LuxPQ
in AC3 C4 |
1.78 |
2.09 |
96.6 |
| ∆LuxPQ
in AC3 C5 |
1.81 |
2.25 |
115.3 |
| ∆LuxPQ
in AC3 C6 |
1.79 |
2.17 |
44.8 |
| PQ-B-R-OU-B
in AC3 C1 |
1.76 |
1.86 |
29.4 |
| PQ-B-R-OU-B
in AC3 C2 |
1.59 |
1.44 |
20.1 |
| PQ-B-R-OU-B
in AC3 C3 |
1.7 |
2.27 |
16.3 |
| PQ-B-R-OU-B
in AC3 C4 |
1.71 |
1.79 |
17 |
|
|
|
|
Construction of LuxPQ-B0015-R0040-LuxOU-B0015 by inserting B0015-R0040-LuxOU-B0015 (AK) into LuxPQ (in AC)
Purpose: To construct LuxPQ-B0015-R0040-LuxOU-B0015 by inserting B0015-R0040-LuxOU-B0015 (AK) into LuxPQ (in AC). This Restriction digest was set up overnight with the following sets of enzymes: XbaI and PstI (insert) and SpeI/PstI (recipient).
|
|
|
KATIE
Continuing to Progress in the Lab
I had to change the order script within the restriction digest tubes since everything needed to be added backward, which turned out to be a problem with my inventory function, which I now have adding inventory names to a list in the opposite order so that it is now correct. I completed the movement scripts I have been working on within the restriction digest tubes so they will move to the water bath, the heating block and then the insert tube moves to the incubator and I think that maybe I will make it wait there for a while until the avatar goes to get it as well as placing it on a timer so if someone does not go to retrieve it will return to its original position anyway and just not give anything and reset the whole script.
Bacteria colony note cards have been made to give when the texture has been changed on the plates, when they are empty, they are inactive, just needed to determine what permission I will need to set on these note cards so that they cannot be copied to inventory, cannot be modified but are still able to be read although not when contained within an object.
I am changing names within construction sites, but decided to keep some of the ones I was going to change general instead of part information, which will now be within the note cards, which people can look at after it is given. I would really like to add more genes to the biobrick construction activity so then I may ask what biobricked part they would like to perform restriction digest on, which provides more of a variety of genes that could be used (I think we will stick with the same promoter/rbs and terminator since it lessens time spent performing restriction digest on similar parts, but people may create circuits that serve different functions).
I have planned out how to begin the lagging strand simulation, which will have to use detection or collision in order to be successful since it will detect (DNA polymerase that is) when it comes across a RNA primer it will begin adding nucleotides. Once the polymerase has traveled down the strand it will signal to I will try to get the replication out into the open at the base of the levels since I have just been using the lab and then taking everything into my inventory.
I will also be changing the PCR machine a bit for the activities so that it may use specific primer you will apparently be given in a starter folder as well as adding a product you may receive.
|
|
|
KEVIN
Sequencing result of Pqrr4+I13500 (GFP+RBS)
Purpose of this sequencing was to verify the presence of both Pqrr4 and I13500 for Emily to test her mutant circuit.
I have received the sequencing result today. At first, I was disappointed because the sequencing that was done with Forward primer had a perfect match with expected Pqrr4 sequence, but only matched 96% of I13500, and as for the sequencing done with Reverse primer, it had perfect match with the expected I13500 sequence, but only matched 99% of Pqrr4. However, because polymerases make mistakes after about 1000bp, this often occurs. So I decided to align the Forward sequence and the reverse sequence.
The following is the result:
Query: Pqrr4+I13500 Forward
Subject: Pqrr4+I13500 Reverse
>lcl|789
Length=1034
Score = 1663 bits (900), Expect = 0.0
Identities = 916/923 (99%), Gaps = 6/923 (0%)
Strand=Plus/Minus
Query 128 CATGAAACTAGAGCCCCGCACACTTGCGGGGCTTTTTAATTTTGAATTTCTTTCTTATTA 187
||||||||||||||||||||||| |||||||| |||||||||||||||||||| |||||
Sbjct 1034 CATGAAACTAGAGCCCCGCACAC-TGCGGGGC-TTTTAATTTTGAATTTCTTT-NTATTA 978
Query 188 AAACGCCATTTTTCTGATAAATGTATTAGTAGCAATGCGCATGGTGGCATATTTGCATCA 247
|||||||| |||||||||||||||||||||||||||||||||||||||||||||||||||
Sbjct 977 AAACGCCA-TTTTCTGATAAATGTATTAGTAGCAATGCGCATGGTGGCATATTTGCATCA 919
Query 248 TTTTGCATTTTGCAAATGCGATTTGCAAAATGCGTGCTCAATAAAGCACCAATATGCATC 307
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
Sbjct 918 TTTTGCATTTTGCAAATGCGATTTGCAAAATGCGTGCTCAATAAAGCACCAATATGCATC 859
Query 308 AGGATCGAAGAAAAAAGGCGTTTTTAAAAGTTGGCACGCATCGTGCTTTATACAGATACT 367
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
Sbjct 858 AGGATCGAAGAAAAAAGGCGTTTTTAAAAGTTGGCACGCATCGTGCTTTATACAGATACT 799
Query 368 AGAGAAAGAGGAGAAATACTAGATGCGTAAAGGAGAAGAACTTTTCACTGGAGTTGTCCC 427
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
Sbjct 798 AGAGAAAGAGGAGAAATACTAGATGCGTAAAGGAGAAGAACTTTTCACTGGAGTTGTCCC 739
Query 428 AATTCTTGTTGAATTAGATGGTGATGTTAATGGGCACAAATTTTCTGTCAGTGGAGAGGG 487
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
Sbjct 738 AATTCTTGTTGAATTAGATGGTGATGTTAATGGGCACAAATTTTCTGTCAGTGGAGAGGG 679
Query 488 TGAAGGTGATGCAACATACGGAAAACTTACCCTTAAATTTATTTGCACTACTGGAAAACT 547
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
Sbjct 678 TGAAGGTGATGCAACATACGGAAAACTTACCCTTAAATTTATTTGCACTACTGGAAAACT 619
Query 548 ACCTGTTCCATGGCCAACACTTGTCACTACTTTCGGTTATGGTGTTCAATGCTTTGCGAG 607
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
Sbjct 618 ACCTGTTCCATGGCCAACACTTGTCACTACTTTCGGTTATGGTGTTCAATGCTTTGCGAG 559
Query 608 ATACCCAGATCATATGAAACAGCATGACTTTTTCAAGAGTGCCATGCCCGAAGGTTATGT 667
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
Sbjct 558 ATACCCAGATCATATGAAACAGCATGACTTTTTCAAGAGTGCCATGCCCGAAGGTTATGT 499
Query 668 ACAGGAAAGAACTATATTTTTCAAAGATGACGGGAACTACAAGACACGTGCTGAAGTCAA 727
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
Sbjct 498 ACAGGAAAGAACTATATTTTTCAAAGATGACGGGAACTACAAGACACGTGCTGAAGTCAA 439
Query 728 GTTTGAAGGTGATACCCTTGTTAATAGAATCGAGTTAAAAGGTATTGATTTTAAAGAAGA 787
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
Sbjct 438 GTTTGAAGGTGATACCCTTGTTAATAGAATCGAGTTAAAAGGTATTGATTTTAAAGAAGA 379
Query 788 TGGAAACATTCTTGGACACAAATTGGAATACAACTATAACTCACACAATGTATACATCAT 847
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
Sbjct 378 TGGAAACATTCTTGGACACAAATTGGAATACAACTATAACTCACACAATGTATACATCAT 319
Query 848 GGCAGACAAACAAAAGAATGGAATCAAAGTTAACTTCAAAATTAGACACAACATTGAAGA 907
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
Sbjct 318 GGCAGACAAACAAAAGAATGGAATCAAAGTTAACTTCAAAATTAGACACAACATTGAAGA 259
Query 908 TGGAAGCGTTCAACTAGCAGACCATTATCAACAAAATACTCCAATTGGCGATGGCCCTGT 967
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
Sbjct 258 TGGAAGCGTTCAACTAGCAGACCATTATCAACAAAATACTCCAATTGGCGATGGCCCTGT 199
Query 968 CCTTTTACCAGACAACCATTACCTGTCCACACA-TCTGCCCTTTCGAA-GATCCCAACGA 1025
||||||||||||||||||||||||||||||||| |||||||||||||| |||||||||||
Sbjct 198 CCTTTTACCAGACAACCATTACCTGTCCACACAATCTGCCCTTTCGAAAGATCCCAACGA 139
Query 1026 AAAGAGAGACCACATGGTCCTTC 1048
|||||||||||||||||||||||
Sbjct 138 AAAGAGAGACCACATGGTCCTTC 116
From it, we can see that the Forward sequence has mismatches towards the bottom, and Reverse sequences has mismatches near the beginning, which makes sense. Because the two sequences match perfectly in the middle, I can assume that I have the right plasmid in these cells.
Next step for this would be to make these cells competent again for Emily and Vicki to put their mutant circuits into it.
|
|
|
MANDY
Changing Up the Wiki
Did regular update of blog onto different sections of the page, so that will keep being updated and it’s great that everybody has been keeping up with posting on the required days.
I fixed up the notebook template so it can be accessed easily from the header menu, and it now applies to all team members (not just the lab people). I’ve also edited the instructions for posting in it to include some more fundamental formatting requirements (with many helpful suggestions from Jamie).
TO DO:
- more blog update inclusion
- Edit Fahd’s news story about NUTV
- Post video of NUTV
- Edit Sponsor page
- Teach Prima how to edit Acknowledgement page
- edit content already there
- add more to intros for human practices and second life
Completion of DNA Extraction Activity in Second Life
The DNA extraction activity is finally done, and it works! (I think). This officially makes me the slowest scripter ever D:< . I am still thinking of a way to make it easier to do. It is intuitive in that it gives you direct instructions, but right now, people can kind of jump in to the middle of it without doing the first bit (which I’m not sure is a good thing – it IS a long activity after all).
Following the DNA extraction activity, I’ve completed most of the instructions/description for the third lab mission. Tomorrow, I will be checking with Katie to make sure the alternative components fit in both the PCR and restriction digest activities to give the desired products (eg. PCR to make a linearized biobricked part). We will also be changing notecards and mission instructions to reflect the changes to these activities.
Speaking of notecards, I’m editing the one I wrote for DNA extraction. I’ve started writing up the instructions for the first lab mission (bacterial transformation) – it’s a ridiculous task (INVISIBLE BACTERIA :D), but I thought it would be fun. This was inspired by seeing Katie make a thing you can wear on your avatar to make you mostly invisible- we might be able to give something like that as a prize?
|
|
|
PATRICK
Flaws and Fixes
Retrospective Notebook: This entry was not written on this day, but derived later from working notes I made that day.
Seeing the biobrick simulator systems in action in the [http://igemcalgary.blogspot.com/2009/07/patrick-and-biobrick-simulator.html blog video] I put together was an excellent way to point out its flaws!
Made one simple but huge upgrade to all of the proteins that bind DNA: caused them to glow briefly when the binding event is accepted by the DNA. No more banging that RNAP on the DNA wondering whether it will take, or whether it wasn't close enough yet! Now it lets you know, as soon as you see your RNAP light up you can stop dragging it and let it get to business!
Also, decided today to scrap multimeric proteins, despite all the work I'd done on them. Putting your TetR together before you can use it is just a pain, transcribing/translating it twice before you can use it is a pain, representing larger protein complexes accurately in SL is nearly impossible, and it all adds very little of value to the simulation. Removing this functionality was surprisingly easy to do without introducing any new bugs, and the unbind screen in the HUD still remains useful for getting your now single object proteins to detach from DNA.
The only reason that I might resuse the type of binding I had before, is if I have time to introduce cofactors to the sim. It only makes sense for IPTG to associate with LacI, for example, and this would add quite a lot of power to the simulation.
|
|
|
PRIMA
Sponsorship deals continued and hotel searches in Boston
Today:
Called Company 1- said they’re not sponsoring any outside projects this year. But they’ll keep us in mind for next year. Wrote a thank you letter and asked for contacts.
Emailed John Winter (Account Manager) for Corning Life Sciences and he’s giving us filtered pipet tips and conical tubes (total of approx $1300)……so they are our bronze sponsors
Called Company 2 – left a message follow up tomorrow
Called Company 3– apparently my last message was sent to the right person but since he never got back to me I got his personal email address and sent him (marketing VP) the sponsorship pkg follow up July 29
Emailed Canadian society of Life Sciences for sponsorship …follow up July 29
Called Company 4 – left a msg follow up tomorrow
Research and emailed the following oil & gas industries:
- Prairie Mines and Royalty Ltd – related to iGEM through pipelines and biofilms
- North American Construction Group – related to iGEM through pipelines and biofims
- Northwest Hydraulics Consultants – cleaning water…
Wrote a thank you letter to Jon’s dad and got the rest of the marketing team to look it over and gave it to Jon. Then I looked up some T-shirts companies and looked up deals.
Tomorrow
- Follow-up with Sigma Aldrich tomorrow- currently looking over sponsorship pkg,
- Call some of the companies I found earlier about T-shirts and try to get discounts/quotes
- Talk about next mass bake sale w/ team
- Talk about other fundraising activities w/ team
|
|
|
STEFAN
GFP zone
Today I added a new "exhibit" to the underwater kingdom!
What is it? Extracting GFP from a jellyfish and putting it in a
bacteria, which then glows green.
How do you do it? Simply click the jellyfish to receive the protein.
Then, open the inventory of the bacteria, put the GFP inside, click on
it and watch it glow!
Why would I do this? This will help people get used to using the SL
inventory system because it will be required in the lab portion of our
island. At the same time, it will be a visual representation of how
cool it is to extract this stuff from animals and put it to use in
bacteria.
Problems: Once I put the GFP inside the glowing bacteria stays like
that. I've been using a sleep script for it to turn off but that
doesn't eliminate the original GFP in the inventory so if I left it
like this the GFP would build up inside. This would not be optimal, so
I've obtained a script from Katie (she also helped me with the
inventory script) to delete the protein inside. I hope to get this
working by tomorrow.
In other news, I brainstormed a few pathways for the Synthetic Kingdom
so people don't get lost and wonder around like Brownian motion. I
also visited other islands in order to get an idea of how to structure
the Help Island the people who come to our sim teleport on. Soon, I
will have a rough draft of the narrative and path people follow (the
only thing I have decided on is glowing platforms. They are
ESSENTIAL).
|
|
|
VICKI
Materials&methods and results section work
Both are complete (save the culture conditions for the bacteria in the Materials/Methods), although I think that they’ll require substantial editing and reorganisation. I had to take extra time to prepare my sequence results in nice form with the biobrick sites highlighted, which took longer than it was supposed to. I’m sending you the PDF now and I’ll convert it to Word later tonight.
|

 "
"