Team:Groningen/Notebook/14 August 2009
From 2009.igem.org
(→Transporters) |
(→Transporters) |
||
| Line 66: | Line 66: | ||
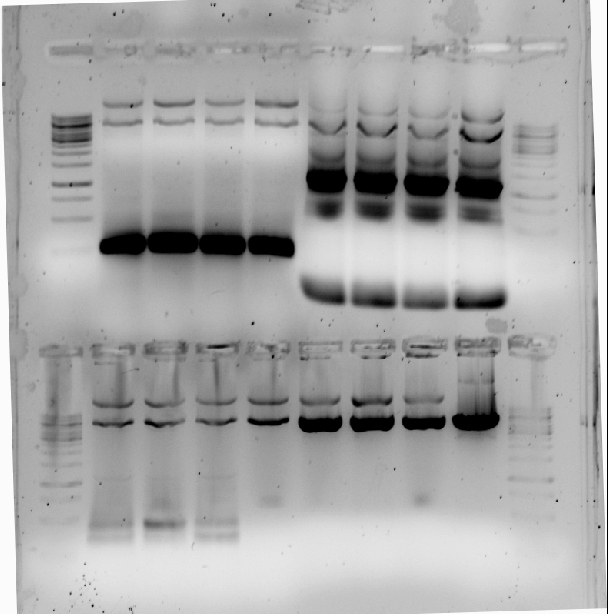
In the first 4 lanes PCR2 (261bp) is created from HmtA-AC3 plasmid with PstI 's left in place. In lane 5 till 8 PCR1 (1153bp)is made from HmtA-AC3 plasmid with PstI 's left in place to make mega primers to mutate the restriction sites. | In the first 4 lanes PCR2 (261bp) is created from HmtA-AC3 plasmid with PstI 's left in place. In lane 5 till 8 PCR1 (1153bp)is made from HmtA-AC3 plasmid with PstI 's left in place to make mega primers to mutate the restriction sites. | ||
| - | In the lower quarter is a PCR on HmtA-AC3 plasmid with PstI 's left in place with VF2 VR | + | In the lower left quarter is a PCR on HmtA-AC3 plasmid with PstI 's left in place with VF2 VR and failed. |
| - | quarter is a XbaI restriction control on plasmids 4,5,6 and 8 not completely digested but indicating the construct is correct. | + | The second lower quarter is a XbaI restriction control on plasmids 4,5,6 and 8 not completely digested but indicating the construct is correct. |
| - | [[Image:F102471_2009-08-14_12hr_03min_PCR1_PCR2_Dynazyme_XbaI.jpg]] | + | [[Image:F102471_2009-08-14_12hr_03min_PCR1_PCR2_Dynazyme_XbaI.jpg|thumb|PCR2, PCR1 Dynazyme XbaI]] |
===Metal Accumulation=== | ===Metal Accumulation=== | ||
Revision as of 14:39, 14 August 2009
Wet
GVP Cluster
- → DONE make glycerol stocks for pSB1AC3-BBa_J23106-BBa_I750016 and pSB1AC3-BBa_J23109-BBa_I750016 (two o.n. cultures of each are grown)
- → DONE isolate plasmid J61002-BBa_J23100, J61002-BBa_J23106, and J61002-BBa_J23109 from o.n. cultures
- → DONE make a planning for next week
Plates
Showed single colony growth on plates with pSB3K3-H/L-GVP plasmids, and were stored in the fridge for future preculture growth.
- → The plates with pSB1AC3-H-GVP plasmid showed no growth, and were stored at 37°C for an additional 4 hours to be sure. Still no growth after addition incubation at 37°C, can be due to high energy cost of vesicle production resulting in the death of the cells!
Over Night Cultures
- → All cultures showed growth, and can be used for plasmid isolation and glycerol stocks. From each culture 800uL was added to 300uL 87% glycerol and frozen in liquid nitrogen. Stocks were stored at position 21 to 24 in the -80°C fridge.
Plasmid Purification
Plasmid isolation was performed on the cultures of E.coli TOP10 containing plasmids BBa_J61002 with high, medium and low constitutive promoters with the "Sygma-Aldrich™ GenElute™ Plasmid Miniprep Kit".
- From each tube 8mL of culture was collected in a 2.0mL cup, and the cells were pelleted by centrifugation for 1 min. at max. speed and the supernatant discarded.
- Plasmids were eluted with 50μL MQ and stored in the fridge
Concentration
| Plasmid | Conc. ng/μL | 260/280 | 260/230 | -20 box (michael | Restriction Control |
| BBa_J61002-BBa_J23100 (high+RFP) | 288.1 | 1.90 | 2.33 | D-8 | No |
| BBa_J61002-BBa_J23106 (med.+RFP) | 162.2 | 1.93 | 2.31 | D-9 | No |
| BBa_J61002-BBa_J23109 (low+RFP) | 114.2 | 1.93 | 2.28 | D-10 | No |
Transporters
After 2 weeks of PCR-ing plasmid isolations there is a product worth wile to send for sequencing. 4,5,6, and 8
In the first 4 lanes PCR2 (261bp) is created from HmtA-AC3 plasmid with PstI 's left in place. In lane 5 till 8 PCR1 (1153bp)is made from HmtA-AC3 plasmid with PstI 's left in place to make mega primers to mutate the restriction sites.
In the lower left quarter is a PCR on HmtA-AC3 plasmid with PstI 's left in place with VF2 VR and failed. The second lower quarter is a XbaI restriction control on plasmids 4,5,6 and 8 not completely digested but indicating the construct is correct.
Metal Accumulation
- ligate MymT in pSB1AC3
- Purify mymT-RBS-pre/suff from gel using Roche Gel Extraction kit
- Elute in 25uL TE buffer (pH8)
- Digest mymT-RBS-pre/suff with EcoRI and Spe I
- Ligate with pSB1AC3/EcoRI-SpeI
Vectors
Dry
|
|
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
|
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
 "
"