Team:Heidelberg/HEARTBEAT database
From 2009.igem.org
(→HEARTBEAT: Rational design of promoter sequences) |
(→HEARTBEAT: Rational design of promoter sequences) |
||
| Line 18: | Line 18: | ||
** [[Team:Heidelberg/HEARTBEAT_database#An in vivo Test of HEARTBEAT predicted sequences|An ''in vivo'' Test of HEARTBEAT predicted sequences]] | ** [[Team:Heidelberg/HEARTBEAT_database#An in vivo Test of HEARTBEAT predicted sequences|An ''in vivo'' Test of HEARTBEAT predicted sequences]] | ||
*[[Team:Heidelberg/HEARTBEAT_database#Discussion|Discussion]] | *[[Team:Heidelberg/HEARTBEAT_database#Discussion|Discussion]] | ||
| + | **[[Team:Heidelberg/HEARTBEAT_database#Transcription factors have a spatial binding preference|Transcription factors have a spatial binding preference]] | ||
| + | **[[Team:Heidelberg/HEARTBEAT_database#Discussion|Discussion]] | ||
**[[Team:Heidelberg/HEARTBEAT_database#Automatization of promoter sequence generation|Automatization of promoter sequence generation]] | **[[Team:Heidelberg/HEARTBEAT_database#Automatization of promoter sequence generation|Automatization of promoter sequence generation]] | ||
Revision as of 01:34, 22 October 2009
HEARTBEAT: Rational design of promoter sequencesContents
IntroductionIn the field of Synthetic Biology mathematical models based on bioinformatic methods are mainly used in order to describe the systems behavior of the involved synthetic constructs. To conduct the orchestra of these parts systems biologists try out different modeling approaches for example analytical or stochastic modeling depending on the underlying system. The introduced parameters in these models are naturally fitted to experimental data acquired in vivo [1]. But the interests in synthetic biology will not only be concerned about systems of existing parts, the focus of research in systems biology will rather be in the design of totally rational assembled constructs not described in nature so far. For that purpose we need models which predict and optimize the properties of these in silico designed constructs. In order to enable this modeling approach it is useful to define a part as a basic biological function encoded as genetic material ([http://blip.tv/file/548184 listen to Drew Endy, President of the BioBricks Foundation]). Starting from this abstract concept we propose to devide up the "parental" part into smaller functional units which we define “subparts” that exhibit their own biological function. However, this function can only be exerted if all subparts are exactly assembled together in the “parental” part. If all system related parameters for every subpart are known, one will be able to model the characteristics of the "parental" part. In this work we propose a model which describes the rational design of a certain promoter of interest. Furthermore, the model is able to predict the functional outcome of the created sequence and it can be used to improve the input sequence, both based on fuzzy logic modeling (HEARTBEAT fuzzy network (FN)). For the development of the model understanding of the functionality and composition of a promoter was essential: it consists of a core-promoter comprising one or more subparts which can work as the binding site for the basal transcription machinery, i.e. RNA polymerase, TATA-box and the Inr-element. Transcription factors regulating the transcriptional activity interact preferentially with the proximal promoter [2]. The intensity of the regulation is further defined by the position of the transcription factor binding site (TFBS) relative to the transcriptional start site (TSS), the binding affinity of the TF to a particular binding motive and also by synergistic effects evolving from interactions between the simultaneously binding of multiple Tfs. [3],[4] In addition the spacer sequences are directly influencing the promoter strength [5]. Based on these assumptions the optimal synthetic promoter sequence can be designed using the following parameters (Fig. 1):
In the following articles we propose a standardized strategy to specify these parameters with computerized methods based on HEARTBEAT. We will further explain the manual design of promoter sequences using these parameters for SREBP and VDR. Initital measurements show the potential of HEARTBEAT-designed sequences to function in vivo (see results). On the basis of our observations, we claim that our way of combining rational generation and experimental validation of synthetic constructs provides a novel and effective strategy in synthetic biology.
MethodsOur promoter sequence was defined to be 1000 bp upstream of the TSS. The [http://genome.ucsc.edu/cite.html| UCSC Genome Browser] [6] provides reference sequence and working draft assemblies for a large collection of genomes including the human genome. We derived the provided sequences containing the 1000 bases upstream of annotated TSS for each RefSeq gene [7]. Upon this pre-selection, we further narrowed our choice of promoter set by selection of distinct pathways. KEGG ([http://www.genome.jp/kegg/Kyoto| Kyoto Encyclopedia of Genes and Genomes]) [8], [9], [10] provides a database of biological systems, consisting of several building blocks including for instance genes and proteins (KEGG GENES) or hierarchies and relationships of various biological objects (KEGG BRITE). Here, KEGG PATHWAY which comprises molecular interaction and reaction networks for metabolism, various cellular processes and human diseases was of particular interest. From this collection of molecular wiring maps, we chose all physiologically relevant pathways for our project, thereby ruling out tissue or other highly specific pathways, like olfactory / taste transduction as well as several pathways related to human diseases. PromotersweepOne of the most challenging problems in bioinformatics remains the computerized localization of TFBS and the transcriptional start sites (TSS) as well as the determination of the core promoter. Many of the available motive discovery tools exhibit the problem of extended false positive predictions. The Promotersweep web-tool improves the accuracy of this analysis by combining a vaste number of different algorithms and methods simultaneously [11]. It integrates information from three homology databases (EnsEMBL Compara [12], NCBI HomoloGene [13],DoOP database [14]), five promoter databases(EPD [15], DBTSS [16]), six sequence motive identification tools (e.g. Gibbs MotifSampler [17]) and two matrix profile databases (Jaspar Core Library [18], Transfac ProfessionalLibrary [19]) to identify and annotate TFBS. The Promotersweep pipeline is started by entering a sequence, chosen between human or mouse as origin. Initially a homology search is performed by using different BLAST algorithms. As a result orthologous promoter regions are deduced from EnsEMBL, Homologene or DoOP, respectively. Subsequently, several motive discovery tools determine shared motives of orthologous or co-regulated sequences. In the last step each TFBS is identified and evaluated with the help of the Transfac - and the Jaspar Core library. Every identified TFBS is classified as weak, conserved or reliable - according to the similarity of the predictions of the different algorithms. In Fig. 2 the result for the Heat shock cognate 71 kDa protein (NM_153201) promoter is shown. For this promoter four different binding sites were discovered. Three of them were classified as reliable and one as conserved. For each hit, the output of Promotersweep contains the position of the motive relative to the TSS. So far we were able to analyse 4395 different promoter sequences, which hold 29966 TFBS in total. The HEARTBEAT-databaseIn order to retrieve the information computed by promotersweep as fast as possible we decided to develop a database structure based on MySQL (My systems query language) [20]. MySQL is one of the most popular relational database management systems. It offers not only a language to set up a hierarchical database but also an interface for easy manipulation of data. The advantage of MySQL is its very intuitive command language and the table structure which helps to minimize redundant data. Simple queries are written in a “SELECT - FROM - WHERE” format. With SELECT value all requested columns are specified. The FROM value calls the corresponding table and WHERE allows a more accurate selection. For our database the average query duration is below 200 ms. This enabled us to provide a fluent online access of HEARTBEAT through the HEARTBEAT GUI. Our data is stored in the tables “Main_Info” and “Gene_Info”. Main_Info contains all necessary data to define the location, binding motive and quality of a TFBS, whereas Gene_Info offers additional information for the gene as well as several gene annotations, where the TFBS is located on. In table 1 and 2 the table structure is shown for Main_Info and Gene_Info. Table 1: Database structure: Main Info
Table 1: Database structure: Gene Info
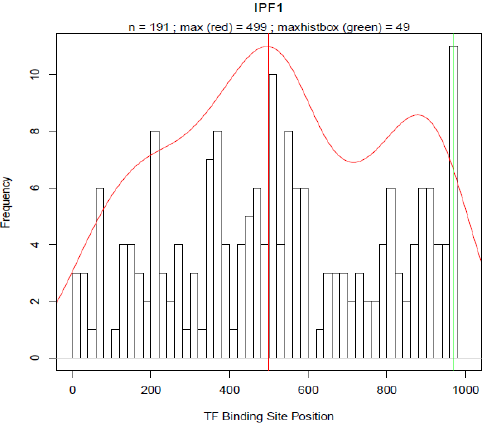
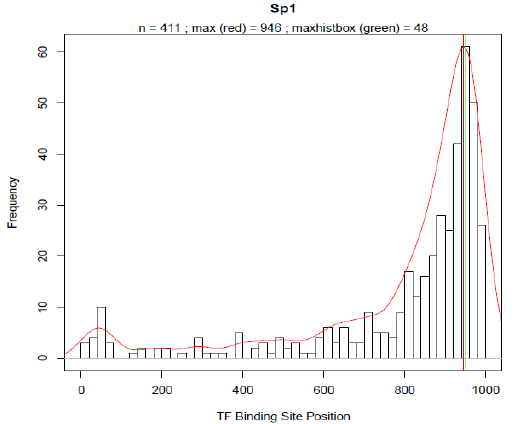
Promoter designFor the rational design of a responsive promoter construct several preliminary considerations have to be done. The first question which needs to be addressed concerns the inducibility of the pathway of interest. Preceding experiments revealed that for VDR (vitamin D receptor) as well as for SREBP (sterol regulatory element binding protein), convenient conditions and treatments exist under which each pathway can be exclusively activated without killing the chassis, that is in our studies the transfected cells. For further reference about induction conditions refer to Material and Methods. After deciding what kind of TFBS is to be included appropriate consensus motives where TFs would presumably bind on had to be selected. Reliable consensus motives can be deduced from matrices provided in the TRANSFAC database. In case of several different binding matrices, we chose the longest motive which contains the most definite bases [19]. Focusing from now on only on VDR and SREBP we created the frequency distributions of the TFBS occurrence for both TFs based on HEARTBEAT (see Fig. 3 and 4). As mentioned above the basic assumption of our model is that most transcription factors exhibit a spatial preference for binding to the DNA which includes not only the binding sequence and the distance to the TSS but also the mutual distance between potential TF pairs. Based on this concept we specified distance of the pdf-maxima to the TSS for both VDR and SREBP. Subsequently the binding motive is embedded into the artificial promoter around the position where the majority of binding sites are located in natural promoters. With this idea we created first a series of synthetic promoters in which we differed only the number of binding motives positioned around the pdf-maxima. For all sequences designed for VDR, see HEARTBEAT_VDR.pdf. For all sequences designed for SREBP, see HEARTBEAT_SREBP.pdf. With a second series of artificially designed promoter sequences we tried to answer to what extend further auxiliary TFBS affect the binding activity of VDR and SREBP. Therefore we plotted the frequencies of all TFBS which are co-occuring when VDR or SREBP is present in a natural promoter sequence as well (see Fig. 5 and 6). For SREBP ZF5 was also present with a relative frequency of 60%. In case of VDR, AP-2 co-exists in 48% and WT1 in 54% of all VDR-promoters. In the following we proceeded in analogy to series one. We created a variety of sequences where we included TFBS in proximity to the pdf-maximum of their frequency distribution besides the VDR and SREBP binding sites. Depending on the number of species of TFBS we distinguish between a blue (1 TFs), green (2 TFs) and orange (3 different TFs) series. Finally all spacer sequences were filled with a random sequence with A:T and C:G content ratios being equal. To make sure that our sequences are as specific as possible we iteratively checked and modified our sequence with the Transfac match tool [21] until no other TFBS expect for our chosen ones were detected. Additionally we tested the sequence for every restriction site used in any Biobrick standard. Finally we added a HindIII at the 5' end and a SpeI restriction site at the 3' end to allow for cloning the construct into the reporter plasmid. ResultsHEARTBEAT analyzes transcription factor binding preferencesFor the statistical analysis we plotted the absolute frequency of occurrence for each TF-binding site in a histogram against the position relative to the TSS where the TSS is located at base 1001 -1003. Each bin comprises 20 bases analogous to different low resolution approaches which analysed the spatial distribution of TFBS with a sliding window of 20-25 bp [22], [3], [23]. From 356 different TFBS for which Transfac contains at least one binding matrix 144 TFBS occurred at least within 50 from 4390 natural promoters. TFBS with less than 50 counts were removed from the selection and not considered for further analysis. In Fig. 7-10 the spatial distributions of Sp1, AP-2, IPF1 (Insulin promoter factor 1) and Kid3 binding sites are shown. The red solid line represents the re-scaled probability density function (pdf). We introduced this function for two reasons. On the one hand the pdf is more robust with respect to outliers than a normal histogram. On the other hand we used the rescaled area under the curve between a shifting frame of 20 bases as a measurement for the significance of a particular TFBS occurrence. The vertical red line in each plot defines the maximum of the pdf. Around the respective base position the majority of binding motives are located within the natural promoters. The maximum of the pdf will serve in the following as the position where binding sites are introduced into our rational designed promoter sequences. An in vivo Test of HEARTBEAT predicted sequences SREBP-responsive sequences designed as outlined above were transfected in HeLa cells and SREBP was induced as described. Promoter activity was then preliminarily analyzed by TECAN (automated fluorescence plate reader). Sequence HB9 showed significant induction (Fig. 11) .
DiscussionIn predicting a functional promoter sequence, we followed one of the basic principles of synthetic biology: Studying nature quantitatively and applying the knowledge thus gained to the de novo creation of biological entities. Four points require further discussion:
Transcription factors have a spatial binding preferenceUpon statistical analysis of over 4000 promoter sequences, we discovered 90 out of 356 TFBS distributions with one significant peak reflecting the spatial occurrence of a particular binding motive. 54 distributions contain two equally high local maxima. For the remaining 212 transcription factors less than 50 TFBS could be detected by Promotersweep and hence were not in the scope of our analysis. Transcription factors have been shown before to have certain spatial binding preferences [3], [4], but no analysis based upon multiple databases and gene orthologies was ever conducted. Our method therefore minimizes false positives, and still, our results confirm existing findings. Improving HEARTBEAT databaseAs noted above, for 212 TFs, less than 50 TFBSs could be detected. In order to overcome this problem in the future we plan to accomplish the screening of the entire genome and to systematically expand our promoter screening by including genomes from different mammalian species. With the increased variety of input sequences we hope to decrease the influence of false positive hits in our statistics. Furthermore it could also help to understand the 54 distributions with multiple maxima and answer the question about the occurance of multiple potential binding sites of one single TF within a given promoter sequence. To make HEARTBEAT even more universal we plan to screen a broader range of the natural promoters which will comprise the sequence downstream of the TSS as well, as it is a well known fact that TF bind to introns and 5' UTR also [24]. In parallel we want to focus on promoters involved in disease related pathways which promises deeper insight in the differences in their molecular regulation. In vivo test of predicted sequences shows that functional promoter can be predicted by HEARTBEAT Figure 12: HEARTBEAT generates functional promoter sequences. Sequence HB 8 and 9 are inducible under appropriate conditions . Sequences marked red were introduced as a negative control Of all the designed sequences, HB8 and HB9 showed activity under appropriate conditions(compare Fig. 11, Results). These sequences are highly similar (see Fig. 12 for how SREs were placed) since they have Transcription Factor Binding Sites under both peaks of the distribution with at least one auxiliary Sp1 element. The other sequences either lack the second SREBP binding site or the Sp1 binding site. Sequences having another type of putative SREBP response element are not functional. We thus showed that the sequences which most closely reflect the expected distribution also work best! These results point out that promoters can be predicted by HEARTBEAT. It also becomes clear that from the study of synthetic promoters, very useful information about gene regulation can be deduced. In order to obtain the information it requires elaborate models, which we started to develop. In the combination with RA-PCR, we provide two very powerful tools for the generation of synthetic promoters. We believe that especially by combining the two methods (see discussion M-RA-PCR), we are able to create any promoter desired in the foreseeable future, thereby accelerating the progress of virotherapy development and fundamental research alike.
Automatization of promoter sequence generationIn this work, the design of promoter sequences was an ad-hoc process which took an entire day. We therefore developed a GUI which assists users with sequence design and automates certain step. Tools such as this will make promoter design a very easy task in the future. References[1] BCCS-Bristol: Best Model 2008 [2] Heintzman N. D. & Ren B. The gateway to transcription: identifying, characterizing and understanding promoters in the eukaryotic genome. Cellular and Molecular Life Science 64: 386-400 (2007). [3] Vardhanabhuti, S., Wang, J. & Hannenhalli, S. Position and distance specificity are important determinants of cis-regulatory motifs in addition to evolutionary conservation. Nucl Acid Res 35(10): 3203-3213 (2007). [4] Yokoyama, K. D., Ohler, U. & Wray, G. A. Measuring spatial preferences at fine-scale resolution identifies known and novel cis-regulatory element candidates and functional motif-pair relationships. Nucl Acid Res 37(13): e92 (2009) [5] Ellis T., Wang X. & Collins J. J. Diversity-based, model-guided construction of synthetic gene networks with predicted functions. Nature Biotechnology 27: 465-471 (2009). [6] Kent W. J., Sugnet C. W., Furey T. S., Roskin K. M., Pringle T. H., Zahler A. M. & Haussler D. The human genome browser at UCSC. Genome Res. 12(6): 996-1006 (2002). [7] Kuhn R. M., Karolchik D., Zweig A. S., Wang T., Smith K. E., Rosenbloom K. R., Rhead B., Raney B. J., Pohl A., Pheasant M., Meyer L., Hsu F., Hinrichs A. S., Harte R. A., Giardine B., Fujita P., Diekhans M., Dreszer T., Clawson H., Barber G. P., Haussler D. & Kent W. J. The UCSC Genome Browser Database: update 2009. Nucleic Acids Res. 37(Database issue): D755-61 (2009). [8] Kanehisa M., Araki M., Goto S., Hattori M., Hirakawa M., Itoh M., Katayama T., Kawashima S., Okuda S., Tokimatsu T. & Yamanishi, Y. KEGG for linking genomes to life and the environment. Nucleic Acids Res. 36 (Database issue): D480-4 (2008). [9] Kanehisa M., Goto S., Hattori M., Aoki-Kinoshita K.F., Itoh M., Kawashima S., Katayama T., Araki M. & Hirakawa M. From genomics to chemical genomics: new developments in KEGG. Nucleic Acids Res. 34 (Database issue): D354-7 (2006). [10] Kanehisa M. & Goto S. KEGG: Kyoto Encyclopedia of Genes and Genomes. Nucleic Acids Res. 28(1): 27-30 (2000). [11] del Val C., Pelz O., Glatting K.-H., Barta E. & Hotz-Wagenblatt, A. PromoterSweep: a tool for identification of transcription factor binding sites. Theor. Chem. Acc. DOI 10.1007: s00214-009-0643-8 (2009). [12] [http://apr2006.archive.ensembl.org/info/software/compara/index.html Ensembl Compara (database)] Hubbard T., Andrews D., Caccamo M., Cameron G., Chen Y., Clamp M., Clarke L., Coates G., Cox T., Cunningham F., Curwen V., Cutts T., Down T., Durbin R., Fernandez-Suarez X. M., Gilbert J., Hammond M., Herrero J., Hotz H., Howe K., Iyer V., Jekosch K., Kahari A., Kasprzyk A., Keefe D., Keenan S., Kokocinsci F., London D., Longden I., McVicker G., Melsopp C., Meidl P., Potter S., Proctor G., Rae M., Rios D., Schuster M., Searle S., Severin J., Slater G., Smedley D., Smith J., Spooner W., Stabenau A., Stalker J., Storey R., Trevanion S., Ureta-Vidal A., Vogel J., White S., Woodwark C. & Birney E. Ensembl. Nucleic Acids Res. 33 (Database issue): D447-D453 (2005). [13] [http://www.ncbi.nlm.nih.gov/homologene NCBI HomoloGene (database)] [14] [http://doop.abc.hu/ DoOP (database)] Barta E., Sebestyén E., Pálfy T. B., Tóth G., Ortutay C. P. & Patthy L. DoOP: Databases of Orthologous Promoters, collections of clusters of orthologous upstream sequences from chordates and plants. Nucleic Acids Res. 33 (Database issue): D86-D90 (2004). [15] [http://www.epd.isb-sib.ch/ EPD (database)] Schmid C. D., Praz V., Delorenzi M., Périer R. & Bucher P. The Eukaryotic Promoter Database EPD: the impact of in silico primer extension. Nucleic Acids Res. 32 (Database issue): D82-5 (2004). [16] [http://dbtss.hgc.jp/ DBTSS (database)] Wakaguri H., Yamashita R., Suzuki Y., Sugano S., Nakai K. DBTSS: database of transcription start sites, progress report 2008. Nucleic Acids Res.36(Database issue): D97-101 (2008) [17] [http://bayesweb.wadsworth.org/gibbs/gibbs.html Gibbs MotifSampler Lawrence (database)] C. E., Altschul S. F., Boguski M. S., Liu J. S., Neuwald A.F., Wootton J. C. Detecting subtle sequence signals: a Gibbs sampling strategy for multiple alignment. Science 262:208-214(1993) [18] [http://jaspar.genereg.net/ Jaspar Core Library (database)] Sandelin A., Alkema W., Engstrom P., Wasserman W. W., Lenhard B. JASPAR: an open-access database for eukaryotic transcription factor binding profiles. Nucleic Acids Res. 32(Database issue): D91-4 (2004). [19] Matys V., Kel-Margoulis O. V., Fricke E., Liebich I., Land S., Barre-Dirrie A., Reuter I., Chekmenev D., Krull M., Hornischer K., Voss N., Stegmaier P., Lewicki-Potapov B., Saxel H., Kel A. E. & Wingender E. TRANSFAC® and its module TRANSCompel®: transcriptional gene regulation in eukaryotes. Nucleic Acids Res. 34 (Database issue): D108-110 (2006). [20] [http://www.mysql.com/?bydis_dis_index=1 Mysql] [21] Kel A. E., Gössling E., Reuter I., Cheremushkin E., Kel-Margoulis O. V. & Wingender E. MATCH: A tool for searching transcription factor binding sites in DNA sequences. Nucleic Acids Res. 31: 3576-3579 (2003). [22] Hannenhalli S. & Levy S. Predicting transcription factor synergism. Nucleic Acids Res. 30: 4278-84 (2002). [23] FitzGerald P. C., Shlyakhtenko A., Mir A. A. & Vinson C. Clustering of DNA sequences in human promoters. Genome Res. 14: 1562-74 (2004). [24] Alberts B., Johnson A., Walter P. & Lewis J. Molecular Biology of the Cell. 5th edition, 2008. Garland Science, Chapter 6
|
 "
"