Team:Imperial College London/M1
From 2009.igem.org
(→Module 1 Contents) |
|||
| Line 142: | Line 142: | ||
| - | <html><center><a href="https://2009.igem.org/Team:Imperial_College_London/M1/EnzymeDelivery"><img style="vertical-align:bottom;" width="20%" src="http://i691.photobucket.com/albums/vv271/dk806/II09_Homepageimage3.png"></a><a href="https://2009.igem.org/Team:Imperial_College_London/M1/PeptideDelivery"><img style="vertical-align:bottom;" width=" | + | <html><center><a href="https://2009.igem.org/Team:Imperial_College_London/M1/EnzymeDelivery"><img style="vertical-align:bottom;" width="20%" src="http://i691.photobucket.com/albums/vv271/dk806/II09_Homepageimage3.png"></a><a href="https://2009.igem.org/Team:Imperial_College_London/M1/PeptideDelivery"><img style="vertical-align:bottom;" width="18%" src="https://static.igem.org/mediawiki/2009/e/ea/II09_Homepageimage3.png"></a><a href="https://2009.igem.org/Team:Imperial_College_London/M1/Genetic"><img style="vertical-align:bottom;" width="20%" src="http://i691.photobucket.com/albums/vv271/dk806/II09_geneticcircuit1.png"></a><a href="https://2009.igem.org/Team:Imperial_College_London/Wetlab/Results#Module_1"><img style="vertical-align:bottom;" width="20%" src="http://i691.photobucket.com/albums/vv271/dk806/II09_Wetlabmainimage9.png"></a><html><a href="https://2009.igem.org/Team:Imperial_College_London/M1/Modelling"><img style="vertical-align:bottom;" width="20%" src="http://i691.photobucket.com/albums/vv271/dk806/II09_Drylabmainimage6.png"></a><center></html> |
</center> | </center> | ||
<html><table border="0" style="background-color:transparent;" width="100%"> | <html><table border="0" style="background-color:transparent;" width="100%"> | ||
Revision as of 02:45, 22 October 2009

Contents |
Module 1: Protein Production
Protein Production
Overview
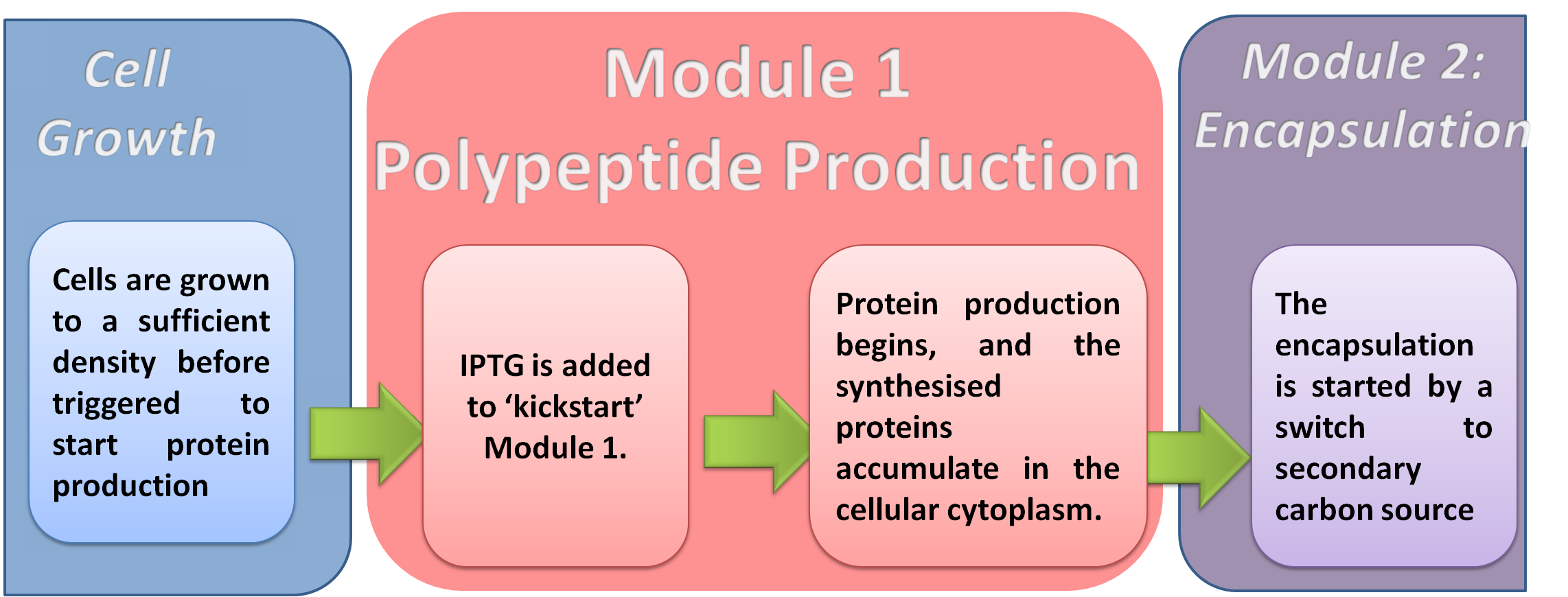
The E.ncapsulator has been designed to produce and deliver protein biopharmaceuticals to the intestine. Module 1 encompasses the protein production phase of the system. The E.ncapsulator is engineered to allow the production of any protein or polypeptide.
Rationale
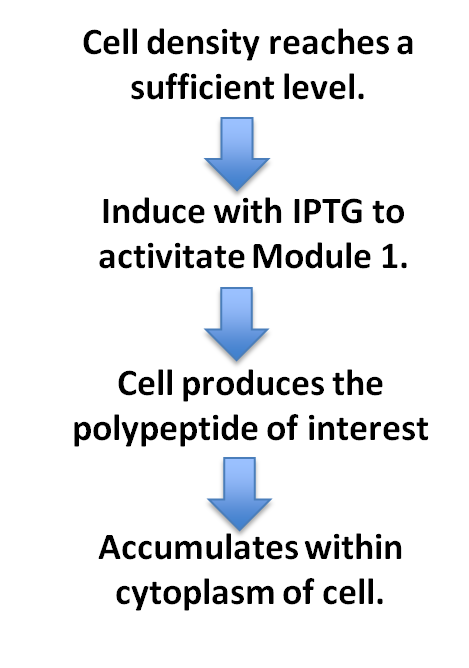
In order to perform this function successfuly, the polypeptide (amino acid polymer) must be synthesised at a rate that will be sufficient to facilitate its accumulation inside the cytoplasm of the cell. With our generic design, it is possible to synthesise any polypeptide, so we have a thoroughly reusable system.
 About the difference between enzymes and peptides.
About the difference between enzymes and peptides.
Theory
Engineering a cell to produce a protein
DNA is transcribed into mRNA which is in turn translated into protein. By knowing the amino acid sequence of the polypeptide of interest, we work backwards and convert this into a DNA sequence coding for production of the protein.
In our case this has been developed further by optimising the DNA sequence for E.coli. By combining this coding sequence the other necessary genetic components, we can engineer the chassis to manufacture the protein.
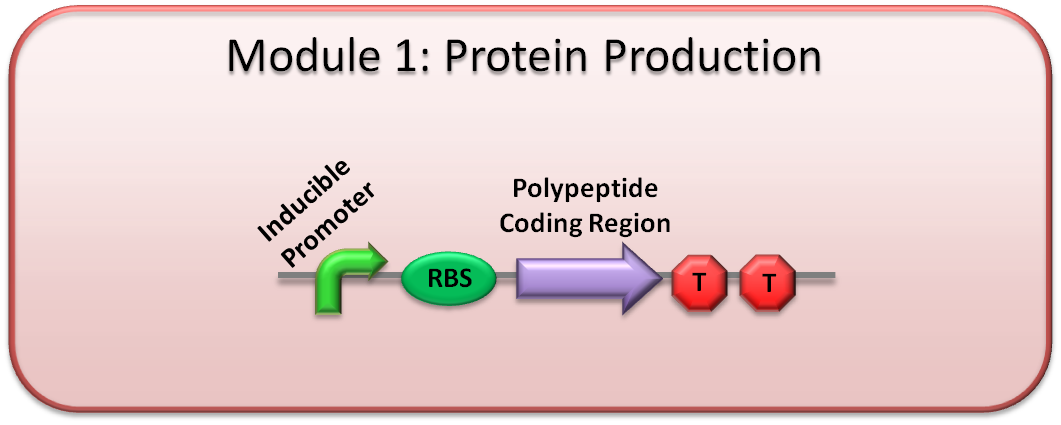
Genetic circuit
This is the remaining part of the Module 1 genetic circuit
Polypeptide Showcase
To demonstrate The E.ncapsulator's versatility, we have chosen to showcase it with both enzymes and peptides. These two classes of polypeptide have very different properties that we have considered and catered for in The E.ncapsulator's design.
Enzyme Production
One of the challenges involved in enyzme production is the need for resistance to the proteases found within the small intestine. Although The E.ncapsulator is capable of delivering the enyzmes safely through the harsh environment in the stomach, upon release into the small intestine, the cells would be susceptible to breakdown by the proteases naturally found in the gut enviroment. For this reason, we have used protease resistant forms of the enyzmes to be produced.We have chosen two enzymes to showcase The E.ncapsulator's protein production module. These are:
- Cellulase - an enzyme that breaks down the tough fibrous molecule cellulose into cellobiose. In delivering cellulase to the gut we aim to increase the nutritional value of food consumed.
 About cellulase and our proposed solution
About cellulase and our proposed solution
- Phenylalanine Hydroxylase - an enzyme that breaks down the amino acid phenylalanine into tyrosine. In delivering this enzyme to the gut we aim to provide a treatment to those suffering from the metabolic condition, phenylketonuria.
 About the genetic condition phenylketonuria, PAH and how we modified it
About the genetic condition phenylketonuria, PAH and how we modified it
Peptide Production
The delivery and production of short chain peptides is a different challenge altogether. All peptides when synthesised always start with the amino acid methionine. If synthesised directly, this can mean that the peptide no longer has the same bioactivity. The body naturally has a mechanism by which larger polypeptides are degraded into smaller functional peptides.
Using this mechanism, we have designed a universal adapter for short chain peptide production and delivery, by which any peptide can be produced and delivered to the gut.
 About our unique strategy for universal manufacture of peptides
About our unique strategy for universal manufacture of peptides
To demonstrate this, we have chosen to showcase a short chain peptide, opiorphin:
- Opiorphin - is a small pentapeptide (5 amino acids) that is naturally produced by the body, and plays a role in pain relief and as an anti-depressant. By delivering this small peptide into the gut, we hope to offer a natural alternative to addictive drugs.
 About opiorphin, and the remarkable effects of this small pentapeptide
About opiorphin, and the remarkable effects of this small pentapeptide
Results
Wet Lab
We have planned protocols for the testing of each of our enzymes of interest, [http://partsregistry.org/wiki/index.php?title=Part:BBa_K200028 BBa_K200028] and [http://partsregistry.org/wiki/index.php?title=Part:BBa_K200007 BBa_K200007] (see Wet Lab Protocols). However, up till now, we have been unable to ligate them into a testable construct.
Dry Lab
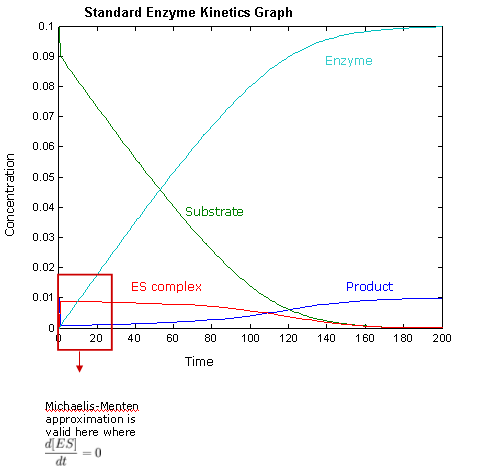
Our Dry Lab team focused on analysing enzyme kinetics theories, such as the well-know Michaelis Menten kinetics. One of the main aims of the dry lab was to characterise the dynamics of the system, making sure enzyme delivery was suitable.
Enzyme kinetic simulations were run to model our system. These are some of the general findings:
- For lower values of [S0], the rate of formation of the product is directly proportional to the amount of [S0]. However, for higher values of [S0], rate of formation of product saturates at a maximum rate.
- Michaelis-Menten approximation only holds for very small values of [E0].
- Increasing K3 values will increase rate of formation of product.
And finally:
- Michaelis-Menten assumption can be applied to our system because for us, KM >> [E0]
Conclusion
Engineering of bacteria to produce proteins is by no means a new development. However, due to the way in which we have designed our chassis to manufacture a protease resitant strains of the enzymes, as well as the unique method for the production and delivery of small peptides, we believe that The E.ncapsulator offers a novel approach to polypeptide production.
Project Tour


Module 1 Contents





 "
"