ULB/13 August 2009
From 2009.igem.org
(Difference between revisions)
| Line 352: | Line 352: | ||
*Bacteria with the plasmid with RBS (BBa_B0034) growed. | *Bacteria with the plasmid with RBS (BBa_B0034) growed. | ||
*The ligation of the promoter-RBS-plasmid (chloramphenicol resistant) didn't work: the reason could be the use of the protocol of manual assembly, who told us to leave league 10 minutes at room temperature, then deactivate the enzymes, insted of 24 hours by the T4 ligase guide. | *The ligation of the promoter-RBS-plasmid (chloramphenicol resistant) didn't work: the reason could be the use of the protocol of manual assembly, who told us to leave league 10 minutes at room temperature, then deactivate the enzymes, insted of 24 hours by the T4 ligase guide. | ||
| - | |||
*Verify that the bricks are now in the medium of our extractions: Digestion of the promoter and promoter + RBS by restriction enzymes EcoRI and Spel, RBS + RFP and the terminator by XbaI and PstI. | *Verify that the bricks are now in the medium of our extractions: Digestion of the promoter and promoter + RBS by restriction enzymes EcoRI and Spel, RBS + RFP and the terminator by XbaI and PstI. | ||
*We made a PCR with primers for ccdA program received at 55 ° C then a gel migration. | *We made a PCR with primers for ccdA program received at 55 ° C then a gel migration. | ||
Latest revision as of 21:13, 21 October 2009
| ||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| ||||||||||||||||||||||||||||||||||||||||||||||||||
| ||||||||||||||||||||||||||||||||||||||||||||||||||
Manipulation on plasmides
- Bacteria with the plasmid with RBS (BBa_B0034) growed.
- The ligation of the promoter-RBS-plasmid (chloramphenicol resistant) didn't work: the reason could be the use of the protocol of manual assembly, who told us to leave league 10 minutes at room temperature, then deactivate the enzymes, insted of 24 hours by the T4 ligase guide.
- Verify that the bricks are now in the medium of our extractions: Digestion of the promoter and promoter + RBS by restriction enzymes EcoRI and Spel, RBS + RFP and the terminator by XbaI and PstI.
- We made a PCR with primers for ccdA program received at 55 ° C then a gel migration.
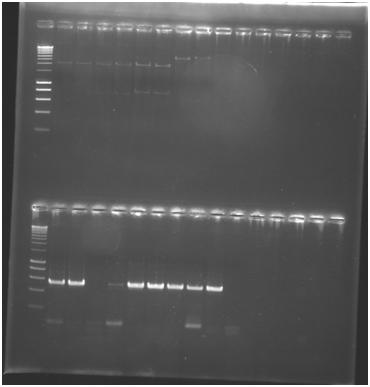
Results
- Verification of bricks:4 DNA primers "normal", white primers "normal", 4 DNA primers reverse, white reverse primers.
- PCR: all tubes have succeeded except the one control sample which is out of place (4th position instead of 6th).
</div>
 "
"