Team:Groningen/Project/Accumulation
From 2009.igem.org
| Line 166: | Line 166: | ||
OpN+OpG = <span id="OpConcentration"></span> µM<br/> | OpN+OpG = <span id="OpConcentration"></span> µM<br/> | ||
ArsR = <span id="ArsRConcentration"></span> µM<br/> | ArsR = <span id="ArsRConcentration"></span> µM<br/> | ||
| - | <span style="color:#808080"> ArsD | + | <span style="color:#808080"> ArsD = <span id="ArsDConcentration"></span> µM<br/> </span> |
</dd> | </dd> | ||
Revision as of 12:36, 10 August 2009
[http://2009.igem.org/Team:Groningen http://2009.igem.org/wiki/images/f/f1/Igemhomelogo.png]
|
|---|
- Transport
- Accumulation
- Metal-sensitive Promoters
- Gas Vesicles
Introduction
TEXT
Once heavy metals have entered the cell it is key to keep them there. As these metals are toxic to cell survival in critical amounts evolution has provided us with biological detoxicification proteins such as [http://en.wikipedia.org/wiki/Metallothionein metallothioneins]. These proteins can aid us in our quest to accumulate a variaty of heavy metals as they bind to a wide range of metals including cadmium, zinc, mercury, copper, arsenic, silver, etc..
Metallothioneins
Metallothioneins are a class of low molecular-weight metal-binding proteins rich in cysteines residues. They are capable of binding a variety of heavy metals. And they have readily been used to create cell based systems for purification of contaminated water [http://www.ncbi.nlm.nih.gov/pubmed/9758654][http://www.ncbi.nlm.nih.gov/pubmed/18618649]. In addition to their wide application possibilities they also have the capacity to carry multiple metal ions at one time, in contrast to some other metalloproteins that carry them one-on-one[http://www.ncbi.nlm.nih.gov/pubmed/9579658]. Many forms of metallothioneins are known and their affinity for different metals has been investigated on several occasions, such as for cadmium[http://www.ncbi.nlm.nih.gov/pubmed/16890348], arsenic [http://www.ncbi.nlm.nih.gov/pubmed/16984198][http://www.ncbi.nlm.nih.gov/pubmed/15294789][http://www.ncbi.nlm.nih.gov/pubmed/18326684], mercuryhttp://www.ncbi.nlm.nih.gov/pubmed/9758654 1[http://www.ncbi.nlm.nih.gov/pubmed/9342882][http://www.ncbi.nlm.nih.gov/pubmed/17920767], nickel[http://www.ncbi.nlm.nih.gov/pubmed/12727263] or a combination of metalshttp://www.ncbi.nlm.nih.gov/pubmed/9579658 3[http://www.ncbi.nlm.nih.gov/pubmed/18313216]. Metal-protein complexes can be quantified using a fluorescent molecule[http://www.ncbi.nlm.nih.gov/pubmed/19133293].
Metals
- As--> ordered pP-1 with rh-MT (human MT gene for accumulation of As3+) [http://www.ncbi.nlm.nih.gov/pubmed/16984198].
- Vector properties unknown, the vector was found to be produced or so by GeneArt.
- Cu--> pBG68 with mymT (M. tuberculosis MT gene for Cu(I) accumulation)[http://www.ncbi.nlm.nih.gov/pubmed/18724363]
- Vector properties: pMB1 ori(20 copy nr.), M13 ori (? copy nr.), tagged with mxe-gyrA intein and chitin binding domain, produced from IPTG inducible T7 promoter (LacI also present).
- Zn--> pMHNR1.1 (a pET29a vector) with smtA (Cyanobacterial MT gene for Zn accumulation)[http://www.ncbi.nlm.nih.gov/pubmed/11493688].
Alternatives
Inclusion bodies[http://www.ncbi.nlm.nih.gov/pubmed/3297654]
(Bacterio)Ferritins
Phytochelatins
[http://www.wiley.com/legacy/products/subject/reference/messerschmidt_toc.html A list of opportunities]
Inhibitory characteristics?
Modelling
Arsenic ArsR
Below you can calculate how many grams of arsenic will be taken out of the water per cubic meter of cells. This extra weight raises the density of the cell and therefore lowers its capacity for buoyancy. Our preliminary results look very promising. Even under the assumption that the weight of the metal is added to the weight of the cells, without increasing their volume, we could add upto a hundred times the currently computed weight without having a large effect on the required fraction of gas vesicles (it will only go up from about 12.2% to 12.7%).
At this moment we use four different variables:
- Molecular weight of arsenic. Source: [http://en.wikipedia.org/wiki/Arsenic Arsenic page on Wikipedia]
- Millimol arsenic per kg of cell dryweight (note that this is equivalent to nmol/mg). Source: Kostal2004
- The proportion between the weight of a dry cell and a wet cell. Source: [http://redpoll.pharmacy.ualberta.ca/CCDB/cgi-bin/STAT_NEW.cgi CCDB Database]
- Cell density. Source: see our gas vesicle page.
|
awAs(III) = g/mol Mcell(dry) / Mcell(wet) = ρcell = kg/m3 |
As(III) intake per volume of cells = g/m3 = µmol/liter (TODO: check) |
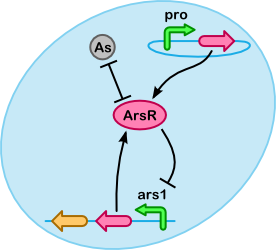
At a lower level arsenic accumulation can be described using reactions between ArsR, As(III) and the ars promoter. As shown in the figure on the right a number of different substances(/complexes) are involved. For our purposes it is especially important to determine what fraction of As(III) is unbound, if more As(III) is bound we can accumulate more.
The calculator below tries to compute the ratio between bound and unbound arsenic, specifically As(III), in the cell. See our Modelling page for detailed information on the constants/variables used and a derivation of the formulas. Note that the computations currently involve slightly more variables/constants than strictly necessary.
|
In conclusion:
- Even at the accumulation levels of Koster et al. the amount of arsenic accumulated in E. coli is so little that it shouldn't matter much for the buoyant density (which normally is about 1100kg/m3).
- If you substitute constitutive promotors for ars promotors you can see that it is clearly advantageous to use constitutive promotors (they give a much higher increase in accumulation).
Planning and requirements:
- Modelling
- Speed
- Metaliotheines concentration
- How often does the ArsR sensitive operator/operon occur in our E. coli?
- Lab
- Measurements
- Transport Assays
- Measure accumulation. By measuring before/after concentration metal with and without accumulation protein.
- Determine the dissociation constant of ArsR and As(III). (By measuring the ratio between bound and unbound ArsR?)
- It might be possible to do this with (tryptophan related) fluorescence (that is how it is done for ArsD in Chen1997). In the paper ArsD is purified, but if that's not feasible for us we might try to simply do it in living cells (and hope that ArsR both fluoresces enough and is produced enough to be measurable).
- Production rate of ArsR?
- Biobrick Bba_K129004
- Rest
- Measurements
Literature
- Brady et al.:[http://www.ncbi.nlm.nih.gov/pubmed/18618649 The use of hollow fiber cross-flow microfiltration in bioaccumulation and continuous removal of heavy metals from solution by Saccharomyces cerevisiae], Biotechnology and bioengineering (1994) 44(11);1362-1366
- Cadosch et al.: [http://www.ncbi.nlm.nih.gov/pubmed/19133293 Uptake and intracellular distribution of various metal ions in human monocyte-derived dendritic cells detected by Newport Green DCF diacetate ester] Journal of Neuroscience Methods (2009) 178(1);182-187
- Chang et al.:[http://www.ncbi.nlm.nih.gov/pubmed/9579658 Cysteine contributions to metal binding preference for Zn/Cd in the b-domain of metallothionein], Protein Engineering 1998 11(1);41–46
- Chen et al.: [http://www.ncbi.nlm.nih.gov/pubmed/9758654 Hg2+ removal by genetically engineered Escherichia coli in a hollow fiber bioreactor], Biotechnology progress (1998) 14(5);667-71
- Chen & Wilson: [http://www.ncbi.nlm.nih.gov/pubmed/9342882 Genetic engineering of bacteria and their potential for Hg2+ bioremediation], Biodegradation (1997) 8(2);97-103
- Deng et al.:[http://www.ncbi.nlm.nih.gov/pubmed/16890348 Cadmium removal from aqueous solution by gene-modified Escherichia coli JM109] Journal of hazardous materials (2007) 139(2);340-4
- Deng et al.: [http://www.ncbi.nlm.nih.gov/pubmed/17920767 Continuous treatment process of mercury removal from aqueous solution by growing recombinant E. coli cells and modeling study], Journal of hazardous materials (2008) 153(1-2);487-92
- Deng et al.: [http://www.ncbi.nlm.nih.gov/pubmed/12727263 Bioaccumulation of nickel from aqueous solutions by genetically engineered Escherichia coli], Water research (2003) 37(10);2505-11.
- Fowler: [http://www.ncbi.nlm.nih.gov/pubmed/3297654 Intracellular Compartmentation of Metals in Aquatic Organisms: Roles in Mechanisms of Cell injury], Environmental Health Perspectives (1987) 71;121-128
- Kao et al.: [http://www.ncbi.nlm.nih.gov/pubmed/18313216 Biosorption of nickel, chromium and zinc by MerP-expressing recombinant Escherichia coli], Journal of hazardous materials (2008) 158(1);100-106
- Ngu & Stillman: [http://www.ncbi.nlm.nih.gov/pubmed/16984198 Arsenic binding to human metallothionein], Journal of the American Chemical Society (2006) 128(38);12473-83.
- Singh et al.: [http://www.ncbi.nlm.nih.gov/pubmed/18326684 Highly Selective and Rapid Arsenic Removal by Metabolically Engineered Escherichia coli Cells Expressing Fucus vesiculosus Metallothionein] Applied and environmental microbiology (2008) 74(9);2924-7
 "
"