Team:Imperial College London/PPS
From 2009.igem.org
JamesField (Talk | contribs) |
(→Peptide Processing) |
||
| (9 intermediate revisions not shown) | |||
| Line 1: | Line 1: | ||
| + | {{Imperial/09/TemplateTop}} | ||
=Peptide Processing= | =Peptide Processing= | ||
To enable <b><i>The E.ncapsulator</i></b> to produce peptides without inhibiting their functionality, we have developed a removable linker region that starts with the amino acid methionine. Following the release of the peptide in the intestine, a naturally occuring enzyme (enteropeptidase) removes the linker region from the front of the peptide releasing the remainder in its functional form. The use of this linker region can be extended to any polypeptide that does not begin with methionine. | To enable <b><i>The E.ncapsulator</i></b> to produce peptides without inhibiting their functionality, we have developed a removable linker region that starts with the amino acid methionine. Following the release of the peptide in the intestine, a naturally occuring enzyme (enteropeptidase) removes the linker region from the front of the peptide releasing the remainder in its functional form. The use of this linker region can be extended to any polypeptide that does not begin with methionine. | ||
| - | |||
| - | This | + | [[Image:Opiorphin.png|left|400px]] |
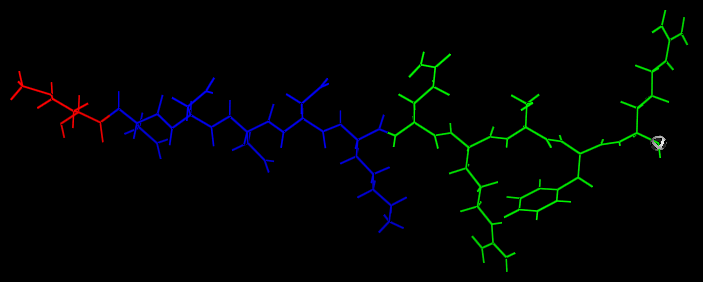
| + | This image shows the universal template for peptide production. The methionine start is shown in RED, the protease recognition site is shown in BLUE and opiorphin is shown in GREEN. The RED & BLUE sections make up the removable linker region cleaved in the gut. | ||
| + | <br><br><br><br><br><br> | ||
| + | <html><center><a href="https://2009.igem.org/Team:Imperial_College_London/M1/PeptideDelivery"><img width=150px src="https://static.igem.org/mediawiki/2009/e/e2/II09_Back2Pep.png"></a></center> | ||
| + | </html> | ||
| + | <br><br> | ||
| + | |||
| + | {{Imperial/09/TemplateBottom}} | ||
Latest revision as of 18:37, 12 October 2009

Peptide Processing
To enable The E.ncapsulator to produce peptides without inhibiting their functionality, we have developed a removable linker region that starts with the amino acid methionine. Following the release of the peptide in the intestine, a naturally occuring enzyme (enteropeptidase) removes the linker region from the front of the peptide releasing the remainder in its functional form. The use of this linker region can be extended to any polypeptide that does not begin with methionine.
This image shows the universal template for peptide production. The methionine start is shown in RED, the protease recognition site is shown in BLUE and opiorphin is shown in GREEN. The RED & BLUE sections make up the removable linker region cleaved in the gut.

 "
"