Team:Heidelberg/Project SaO
From 2009.igem.org
(→Outlook and Summary) |
|||
| Line 6: | Line 6: | ||
|width="650px" style="padding: 0 15px 15px 20px; background-color:#ede8e2"| | |width="650px" style="padding: 0 15px 15px 20px; background-color:#ede8e2"| | ||
__NOTOC__ | __NOTOC__ | ||
| - | + | ='''Outlook and Summary'''= | |
The urge in establishing manipulable eukaryotic systems has posed much pressure on the development of methods to help understand and characterize eukaryotic gene regulation. Those methods go beyond the already rather sophisticated methodology still being established in prokaryotes to investigate and thereafter engineer these cells as needed [[Team:Heidelberg/Project_SaO#References|[1]]]. For one thing, the design of promoters exclusively responsive to one transcription factor (TF) within eukaryotic cells could certainly help improve our understanding of the key components of one pathway or the other, while eliminating the cross-talk often observed with many naturally occurring promoters. Such promoters have often posed a challenge to researchers studying signal transduction in eukaryotic systems because of the different types of TFs that a single regulatory element can bind, and a single TF having multiple target regulatory regions [[Team:Heidelberg/Project_SaO#References|[2]]]. With the emerging of systematized research and attempts for modeling biological systems, the availability of data with minimal experimental variability and highly accurate experimental conditions has also contributed to the need for such finely-tuned promoters. Once such exclusive promoters could be available and methods for their characterization established, it is not so hard to imagine the revolutionary effect they could have on eukaryotic research. Some of many applications could be: | The urge in establishing manipulable eukaryotic systems has posed much pressure on the development of methods to help understand and characterize eukaryotic gene regulation. Those methods go beyond the already rather sophisticated methodology still being established in prokaryotes to investigate and thereafter engineer these cells as needed [[Team:Heidelberg/Project_SaO#References|[1]]]. For one thing, the design of promoters exclusively responsive to one transcription factor (TF) within eukaryotic cells could certainly help improve our understanding of the key components of one pathway or the other, while eliminating the cross-talk often observed with many naturally occurring promoters. Such promoters have often posed a challenge to researchers studying signal transduction in eukaryotic systems because of the different types of TFs that a single regulatory element can bind, and a single TF having multiple target regulatory regions [[Team:Heidelberg/Project_SaO#References|[2]]]. With the emerging of systematized research and attempts for modeling biological systems, the availability of data with minimal experimental variability and highly accurate experimental conditions has also contributed to the need for such finely-tuned promoters. Once such exclusive promoters could be available and methods for their characterization established, it is not so hard to imagine the revolutionary effect they could have on eukaryotic research. Some of many applications could be: | ||
{| style="background-color:#ede8e2;" | {| style="background-color:#ede8e2;" | ||
Latest revision as of 22:33, 21 October 2009
Outlook and SummaryThe urge in establishing manipulable eukaryotic systems has posed much pressure on the development of methods to help understand and characterize eukaryotic gene regulation. Those methods go beyond the already rather sophisticated methodology still being established in prokaryotes to investigate and thereafter engineer these cells as needed [1]. For one thing, the design of promoters exclusively responsive to one transcription factor (TF) within eukaryotic cells could certainly help improve our understanding of the key components of one pathway or the other, while eliminating the cross-talk often observed with many naturally occurring promoters. Such promoters have often posed a challenge to researchers studying signal transduction in eukaryotic systems because of the different types of TFs that a single regulatory element can bind, and a single TF having multiple target regulatory regions [2]. With the emerging of systematized research and attempts for modeling biological systems, the availability of data with minimal experimental variability and highly accurate experimental conditions has also contributed to the need for such finely-tuned promoters. Once such exclusive promoters could be available and methods for their characterization established, it is not so hard to imagine the revolutionary effect they could have on eukaryotic research. Some of many applications could be:
A high-level application of synthetic promoters that lies close at hand is the development of a "cell-based drug screening assay". Such an assay is based upon a cell line which is stably transfected with a multitude of promoters responsive to a variety of pathways. Each promoter would be linked to a unique output signal (see below). This cell line could be stimulated with a variety of drug candidates, and the molecular effect of each drug would directly be visualized. The availability of such a cell line would greatly accelerate the pace of drug discovery and pharmacology alike. We have created all the parts and knowledge required for such an assay, and would only need to assemble it.
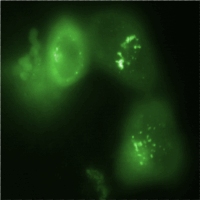
For output, we suggest using a variety of fluorescent proteins (with non-overlapping spectra) coupled to localization tags. We were able to provide two FPs (GFP and mCherry) as well as four localization sequences (1x Endoplasmic reticulum; 1x Nucleus; 2x Plasma membrane). We show that combining our FPs with our localization sequences works by using the [http://dspace.mit.edu/handle/1721.1/45139 BBb_2 proposal], and thus provide future users with the possibility to visualize at least 6 different promoters simultaneously. We also established methods for promoter measurement in eukaryotes utilizing microscopy, flow cytometry (FACS) and qRT-PCR. Promoter activity was not measured by microscopy before; we established this technique in order to be able to measure protein levels in different compartments and thus make the output legible. Not only to overcome challenges for promoter measurement, but also to lay the foundations for the proposed drug-screening assay as well as any other synthetic biology idea in mammalian cells, we emphasize the importance of stable cell line creation. At the end, we are proud to say that we have introduced many of the concepts, methods and tools that could serve as the basis for all other attempts in the engineering of eukaryotic gene regulatory systems. Not neglecting the need for further improvement, with such a collection of tools available, the ideas of selective protein and gene therapy, metabolic engineering, stem cell manipulation and better intracellular network modeling do not seem too far away. References[1] Venter M. Synthetic promoters: genetic control through cis engineering. Trends in Plant Science 12: 118-124 (2007). [2] Carey M., Smale S. T., Hughes H., Transcriptional Regulation in Eukaryotes: Concepts, Strategies and Techniques. New York:CSHL, p. 18-25 (2000). |
 "
"