Team:UNIPV-Pavia/Project/Results
From 2009.igem.org
| Line 24: | Line 24: | ||
</tr> | </tr> | ||
<tr> | <tr> | ||
| - | <td> {{UNIPV-Pavia/Results}} | + | <td> {{UNIPV-Pavia/Results}} |
| - | + | ||
| - | + | ||
=Experimental results= | =Experimental results= | ||
| Line 128: | Line 126: | ||
=Project conclusions and perspectives= | =Project conclusions and perspectives= | ||
| + | |||
| + | </td> | ||
| + | </tr> | ||
| + | </table> | ||
Revision as of 19:20, 20 October 2009

|
|
|
Results & Conclusions | |||||||||||
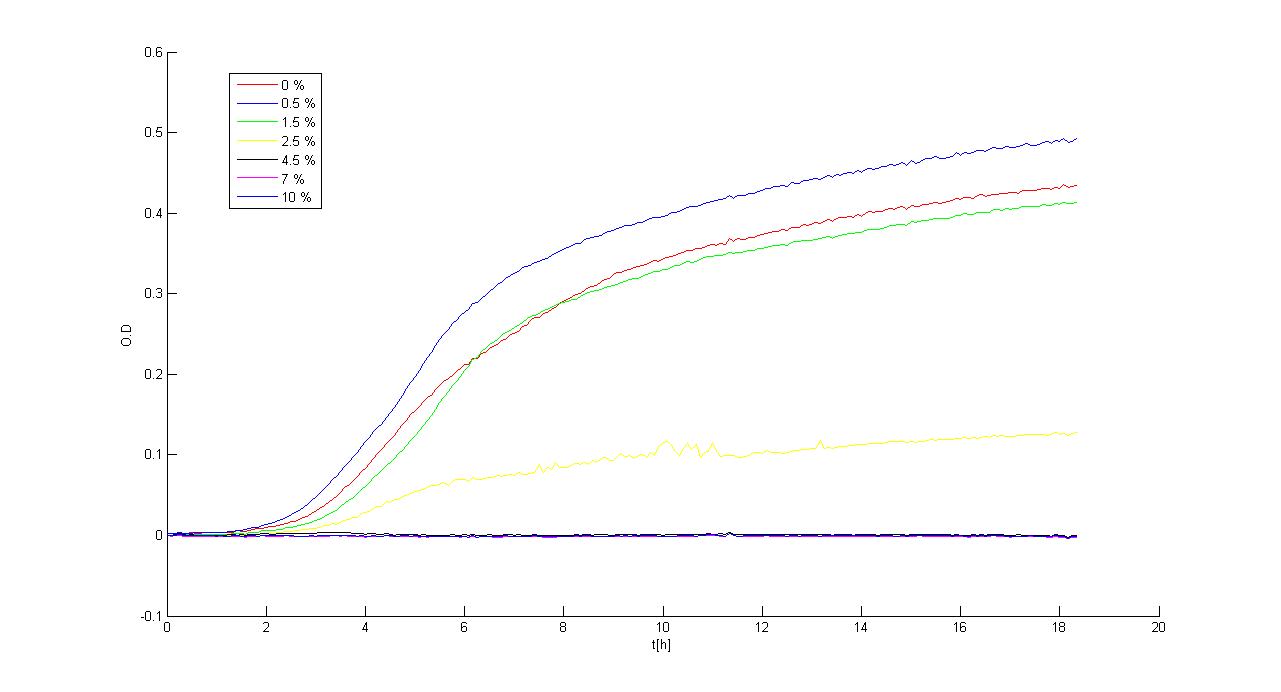
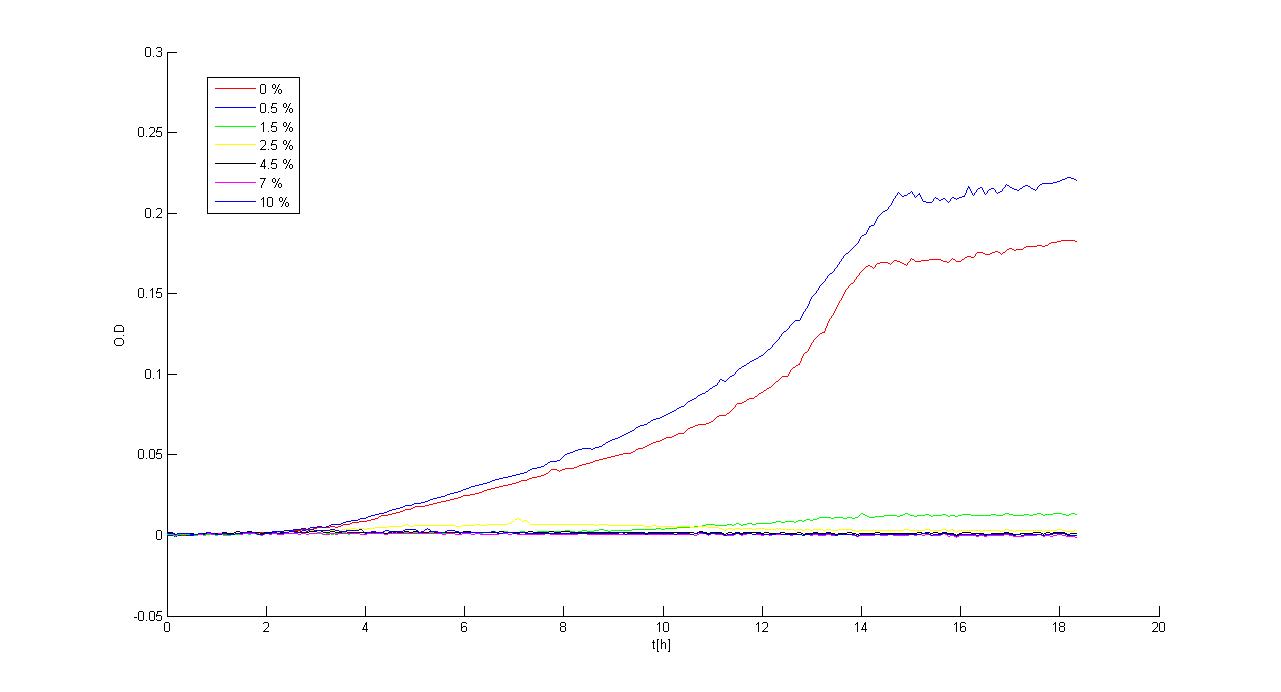
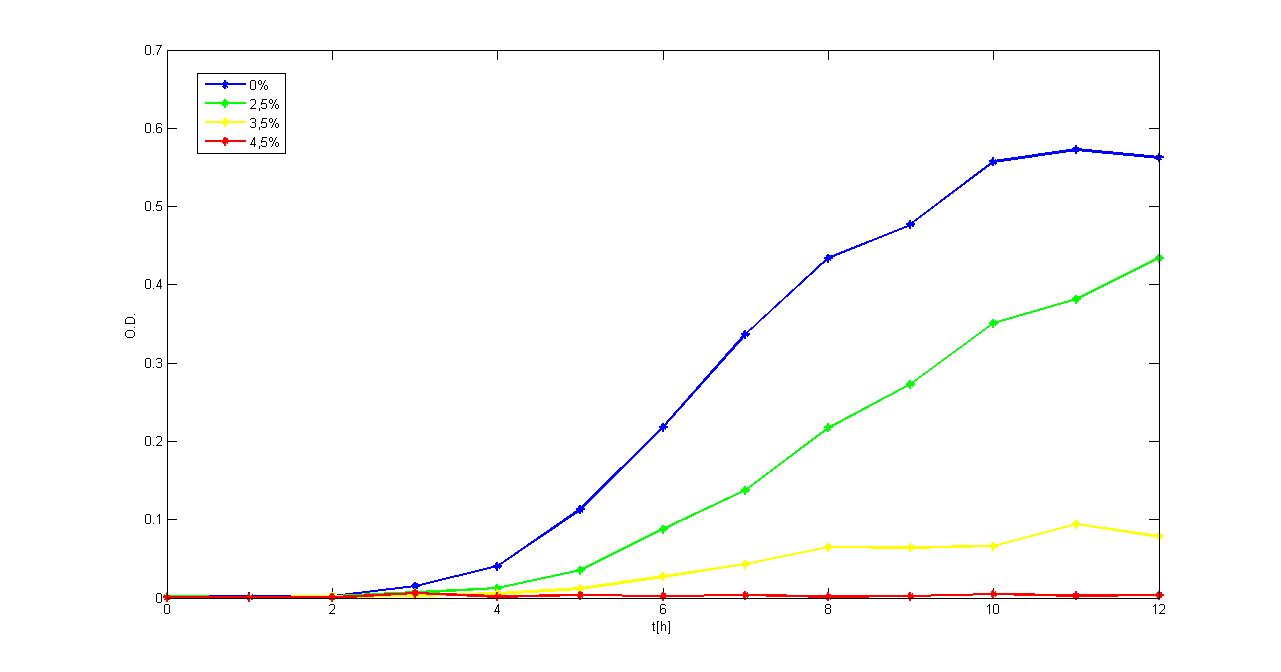
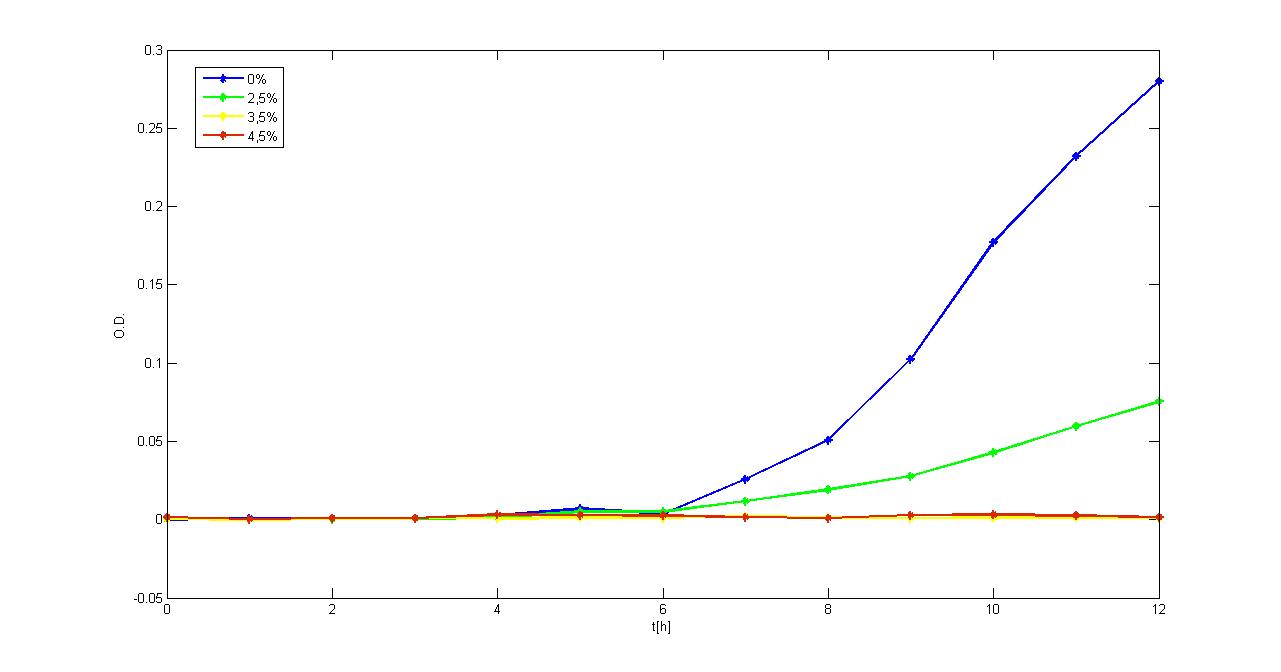
Experimental resultsIn this section a summary of "Ethanol? Whey not!" project results is reported. For more information you can visit Parts Characterization section. Ethanol fermentationSurvival of E. coli TOP10 in ethanolDifferent concentrations of ethanol have been added to LB and M9 supplemented with glycerol selective media and E. coli TOP10 (bearing BBa_B0030 BioBrick) survival, when grown in a microplate reader (37°C, automatic protocol) and in a 50 ml falcon tube (37°C, 220 rpm), has been tested.
Of the concentrations we tested, in the microplate reader bacteria can grow at concentrations up to 2.5%w/v (LB) and 1.5%w/v (M9), while in the 50 ml falcon tube they can grow at concentrations up to 3.5%w/v (LB) and 2.5%w/v (M9). In general, in the falcon tube bacteria grow better than in the microplate (see Measurement section for details) and this is confirmed by these data: it seems that the threshold concentration of ethanol is higher for growth in falcon tube than in the microplate (for example, at 2.5%w/v in M9, bacteria can survive in the falcon tube, but not in the microplate). Ethanol productionA synthetic ethanol-producing operon () has been built up by our team and has been assembled downstream of several promoters or devices:
These strain's most important phenotype qualitative characteristics can be summarized as follows:
All the other details can be found on Parts Characterization page, as well as in Registry page.
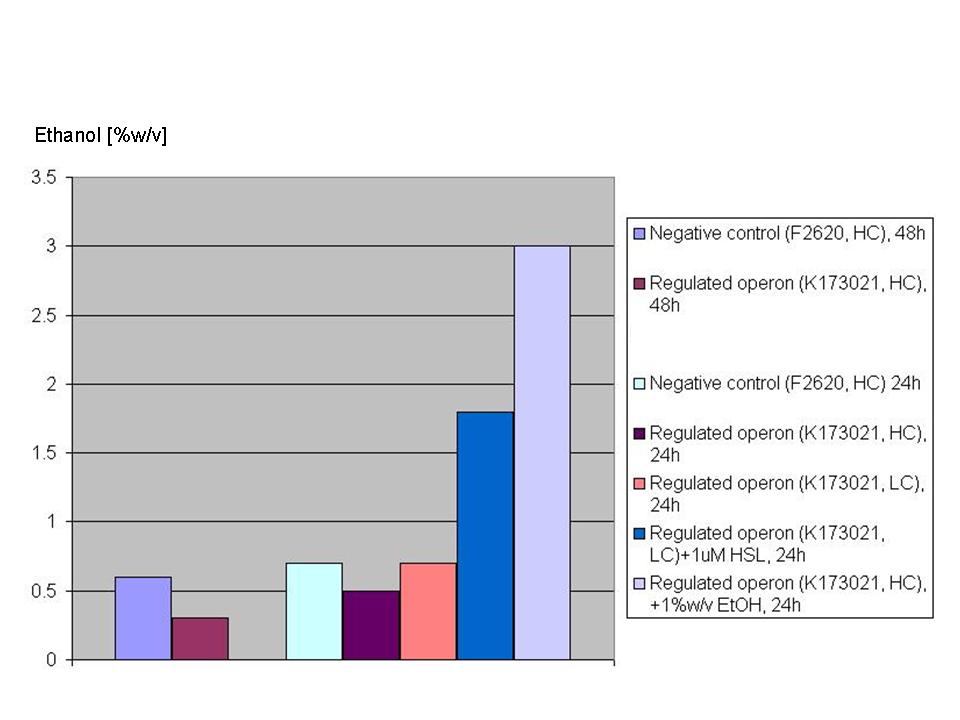
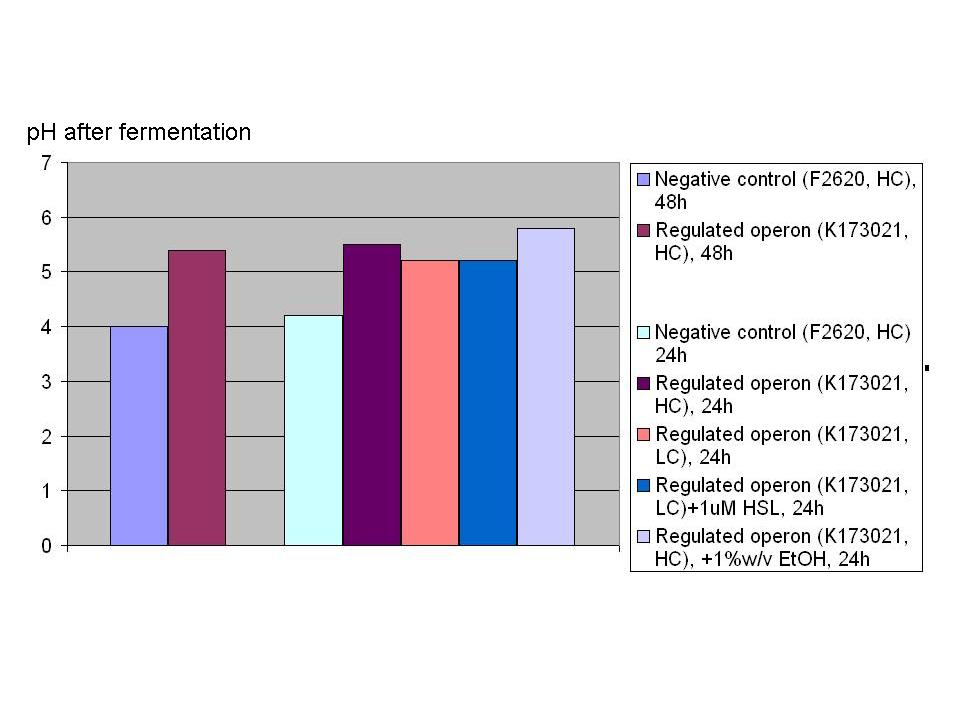
These strain's most important phenotype quantitative characteristics can be summarized as follows: Growth of strains bearing:
has been monitored in selective LB + 10% glucose starting from an inoculum in the microplate reader (figure below).
The main fermentation results can be summarized as follows:
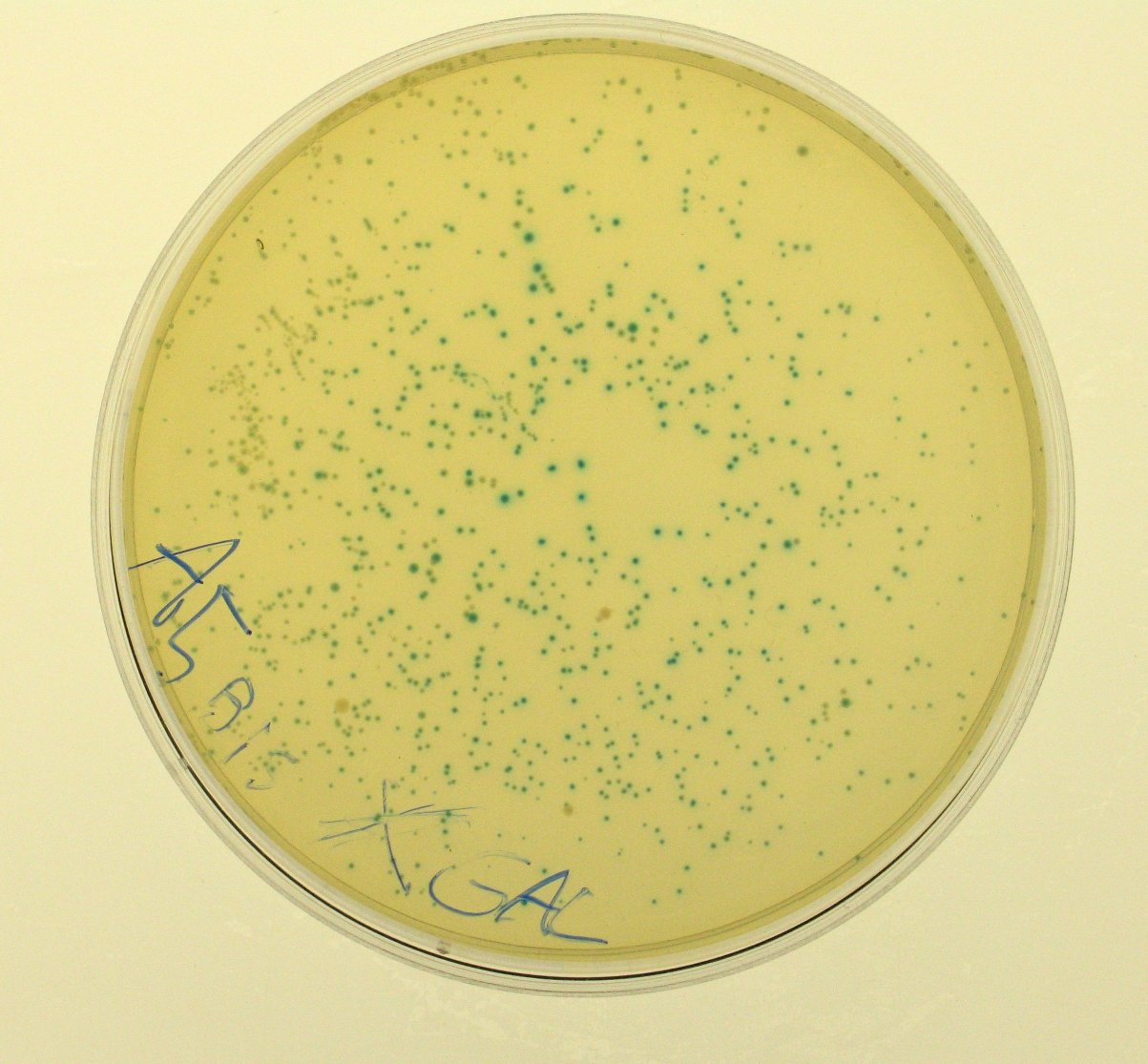
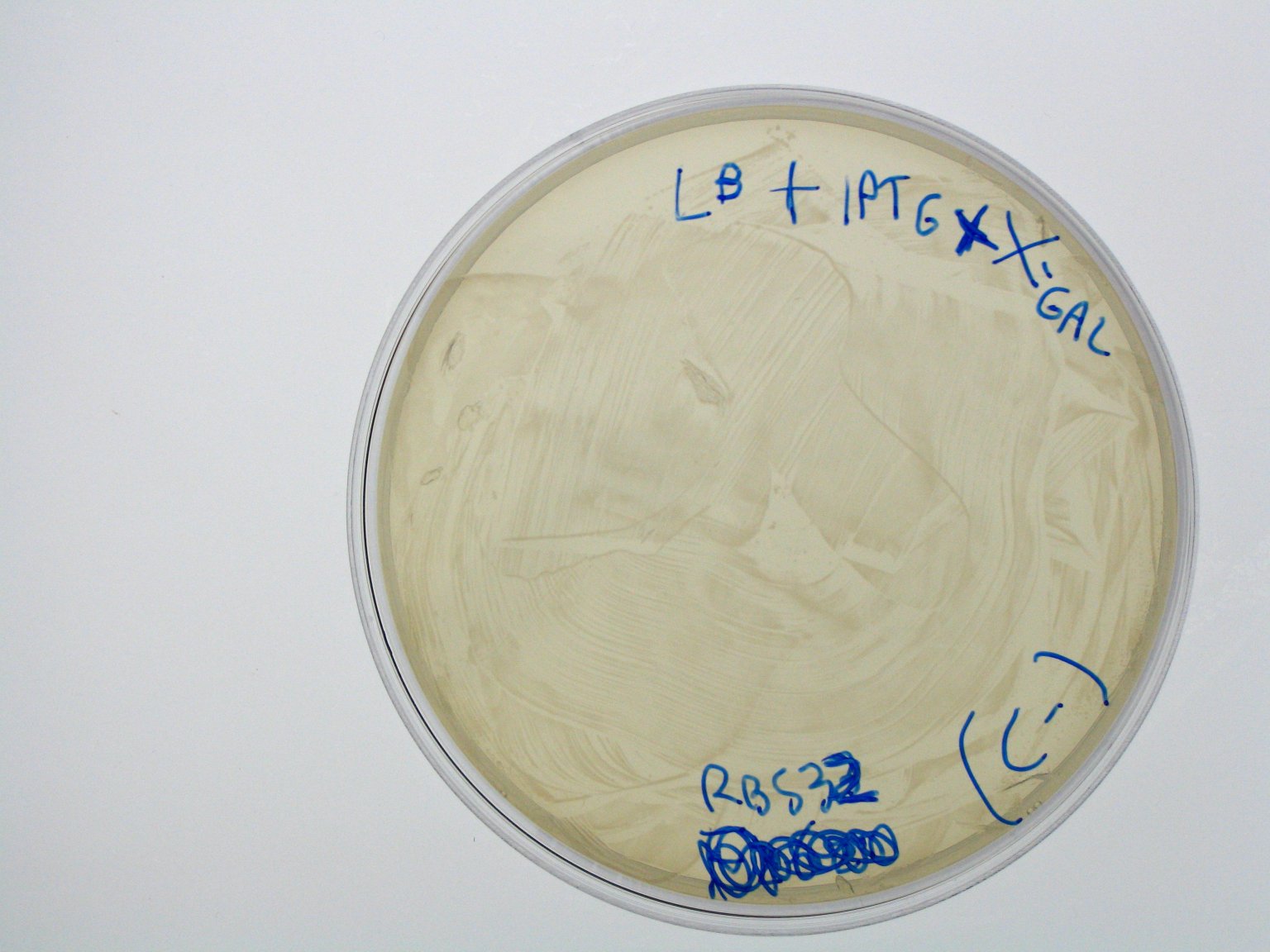
Lactose metabolismA bacterial beta-galactosidase protein generator () was assembled from the parts contained in the Registry, even if it had already been designed and submitted as , whose sequence was consistent according to iGEM QC. The enzyme, encoded by lacZ gene, cleaves lactose in glucose and galactose and can be used, properly assembled downstream of a regulatory module, to speed up the lactose metabolism. We used tetR promoter, contained in , to produce a constitutive expression of lacZ.
This qualitative result shows that:
Anyway, further comparative tests should be done in order to see if lactose cleavage can be performed faster than in wild type E. coli, after the choice of a suitable promoter which controls this protein generator. Promoter characterizationSeveral promoters and devices have been tested by our team during this summer. Our efforts have been focused on standard characterization methodologies, so we decided to try to apply the Relative Promoter Units (Kelly J. et al., 2008) concept to do this. Regulation of gene expression is very important for our project as we want to optimize the expression rate of two actuators, one for lactose conversion to glucose and one for ethanol production from glucose. In our case it essential do have a sort of "genetic regulators user's handbook", in which all the functional characteristics of promoters can be found and in which we can choose the suitable regulator for us. We dedicated a large part of our project in this direction and here we summarize the promoter strength measurements. In the following bar plot we report:
Project conclusions and perspectives |
 "
"