Team:Heidelberg/HEARTBEAT network
From 2009.igem.org
HEARTBEAT Fuzzy ModelingContents
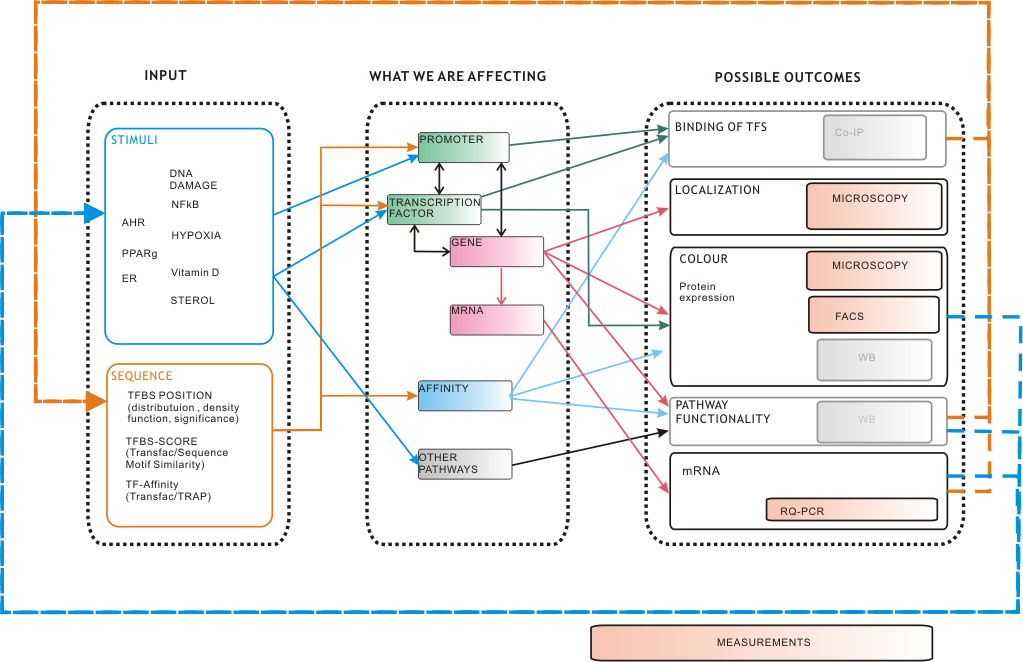
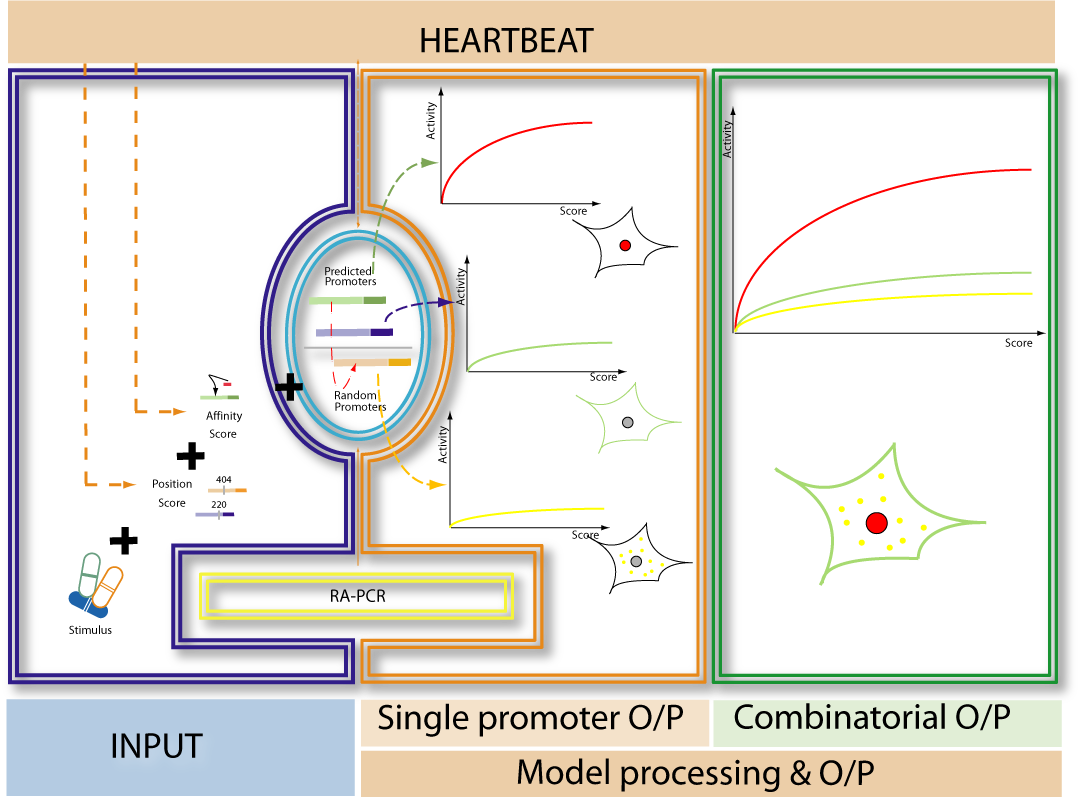
IntroductionThe research field of regulation of gene expression in eukaryotes is a field of biological research growing rapidly [1,2]. Hereby the interaction of DNA with certain proteins known as transcription factors (TFs) plays an essential role for the complex mechanism of transcriptional activation [3,4]. One strong focus of synthetic biology aims at the reconstruction of such gene regulatory networks [5-8]. To act within the scope of synthetic biology’s duties, the iGEM Team Heidelberg 09 claims that any synthetic promoter can be constructed by using our two methods for the construction of synthetic promoters. However, the only efficient way to construct systems of high complexity (such as computers or airplanes) is simulating these systems on the computer prior to construction [9]. In our case, this strongly emphasizes the necessity of HEARTBEAT (Heidelberg Artificial Transcription Factor Binding Site Engineering and Assembly Tool) which comprises data analysis (HEARTBEAT_database), a graphical user interface (HEARTBEAT GUI) and network modeling (HEARTBEAT fuzzy network (FN) modeling). Contributing to the HEARTBEAT project, HEARTBEAT FN focuses on simulating the promoter activity by integrating a variety of signals and sequence characteristics as well as on predicting distinct pathway functionalities. This, especially in eukaryotic systems, is a tough challenge since transcriptional activity of a gene is not directly correlated to protein expression [10]. For this purpose we propose fuzzy logic (FL) modeling as an approach to logic-based modeling which is capable of incorporating qualitative data but producing quantitative predictions. New insights will be provided about the operation of gene regulatory networks and relationships between promoter sequence composition and TF-DNA interaction will be unraveled that is understood only marginally so far [11-12]. [TOP] Background / MotivationWe present two different approaches for promoter design resulting in three different types of synthetic promoters: randomly assembled constitutive and inducible promoters as well as rationally designed promoters. As an additional type of promoters those occurring in nature can be integrated into vector systems. These heterogeneous cocktail of promoters can be combined for a precise regulation of pathways. This represents the power of our entire HEARTBEAT project. Synthesized promoters can be then used e.g. as a combinatorial gene therapy, i.e. several promoters that are of different types and/or have different strength will be applied as treatment agents. Therefore, a model that not only simulates single promoter activity and following gene expression but also accurately predicts gene expression from combined promoter sequences is indispensable. We constructed a Fuzzy Logic model to provide a formal mathematical framework for prediction of combined activity of multiple promoters upon several stimuli and to gain insight into the mechanisms that generate diverse expression levels. A Short Introduction into Fuzzy Logic ModelingFuzzy Logic is a rule-based approximate artificial reasoning method developed by [http://en.wikipedia.org/wiki/Lofti_Zadeh| Lotfi Zadeh] in 1965. Its motivation is the observation that humans often think and communicate in a vague way, and yet can make precise decisions [13]. It has been widely used in engineering and Artificial Intelligence approaches such as Fuzzy Controllers and Fuzzy Expert Systems. Fuzzy Logic has also been used for the modeling of biological pathways [14] and very recently to analyze gene regulatory networks [15]. Key advantages of Fuzzy logic-based approaches are (i) the ability to construct models based on prior knowledge of the system and experimental data and (ii) encode intermediate states for inputs and outputs, thus improving other logic-approaches that can only deal with ON/OFF states such as Boolean models [16] and (iii) simulations can be derived from both qualitative and quantitative data, both of which can be cast into the form of IF-THEN rules. Thus, FL constitutes a powerful approach for the understanding of heterogeneous datasets. A Model combining in silico prediction with experimental dataIn our project, the complete set of rules will capture the behavior of each promoter in a Multiple-Input Single-Output (MISO) Fuzzy Logic model. Combining the MISO models in a network of all promoters will constitute the final Multiple-Input Multiple-Output (MIMO) model allowing for the simulation and prediction of combined activation of pahways regulated by our promoters. A key advantage of this methodology towards understanding the exclusive pathway activation of our promoters of interest is the possibility to study not only the individual activity of each promoter but also the combined activity, as the signal progresses from one MISO to another. A general idea of our modeling network which is already adapted to our whole project is shown in Fig. 1.
AchievementsModel descriptionOur HEARTBEAT fuzzy network modelThe HEARTBEAT fuzzy network model we have developed has four layers with different functionalities (see Fig.1). Its global aim is of a particular importance! The Heartbeat tells you what sequence allows for regulation of the expression of your gene of interest, but how do you know how to combine more than one promoter, i.e. what combined activity to expect? What if these promoters crosstalk (see Fig.1)? What if you want to use several exclusive promoters to regulate several pathways? Our model predicts the final activity of all promoters by using their Affinity and Position Scores and regulatory events such as inducible/constitutive activity, fixed stimulus, or increasing stimuli. A key advantage of the fuzzy logic core of the modeling approach is the use of both quantitative and qualitative experimental data as well as literature-prior knowledge. Two Measures for scoring promoter sequencesTo capture the quality of the promoter sequence of interest we introduced two measures: the affinity of the designated TF to the sequence as well as the impact of the position of the binding site in the sequence. The affinity of a TF to the sequence of interest was calculated with TRAP (TRanscription factor Affinity Prediction) which predicts a relative binding affinity to a DNA sequence using a physical binding model [17]. The position of the binding site of a TF was assessed using its probability density function which was derived from the HEARTBEAT database as described previously. A sliding window of 20bp was shifted over the binding distribution of the transcription factor to determine its preferential binding region. The area under the curve (AUC) at the determined position was then scaled according to the number of total hits of the particular transcription factor. In order to further evaluate the significance of a certain TF binding site, we first calculated the difference between the AUC of interest and the mean over all values (ΔAUC(x)). We then calculated the difference between the maximal AUC value and the mean over all values (ΔAUC(max)) and finally estimated the quotient (Eq.1). By this, we were able to characterize a promoter of interest from two different aspects, biophysical affinity and spatial distance to the TSS, respectively. Model assumptionsWhile constructing the model, we made several assumptions which our model is based on. First of all, we choose JeT as a established methodological reference for our data analysis prior to model development. We furthermore assume that a high TF affinity corresponds to a high promoter activity. At the same time, we assume that a high TFBS position score also correlates with a high promoter activity. A system describing an inducible promoter cannot be invoked if either affinity or position score is low. Upon stimulation, promoter having such characteristics should not gain a high activity. Obviously their basal activity is considered not to be high as well. We also consider that the optimal transcriptional activation of a promoter of interest cannot be reached by a single high score. For this purpose, both scores have to be at a high range. Regarding the scoring of the promoter sequences, the main problem which had to be overcome was the integration of single scores to one overall promoter score. TRAP calculates one score which describes the affinity of one TF to a given sequence. The position score, moreover, is calculated for each single TFBS of each TF on a given sequence. We decided to choose the maximum of the scores ("pos.max.score" or "af.max.score") because:
At this point it is indispensable to mention that we assume that a better binding of a TF to a promoter sequence leads to a enhanced transcrptional activity. Since the promoter activity was captured by fluorescence, we assume that the increase in transcription of target genes (which in our case were coding for the fluorescent proteins) should lead to a higher expression of the respective protein as well. For the last part of our modeling, the combination of distinct pathways, the different systems are integrated by either applying a fuzzy inference system or simply calculating cumulative activity.
Defining Fuzzy Membership Functions & Fuzzy Rules: capturing the behavior of our experimental dataAs a first step to create a model using fuzzy logic, experimental data had to be studied in order to define several so-called membership functions. Based on data analysis techniques we defined 13 classes in which our data could be clustered due to differentail activity for a total of 5 variables. In Fuzzy Logic, each class requires a membership function that accurately defines its behavior. Therefore we established as many membership functions (MF) as classes for all inputs. These classes were characterized as follows:
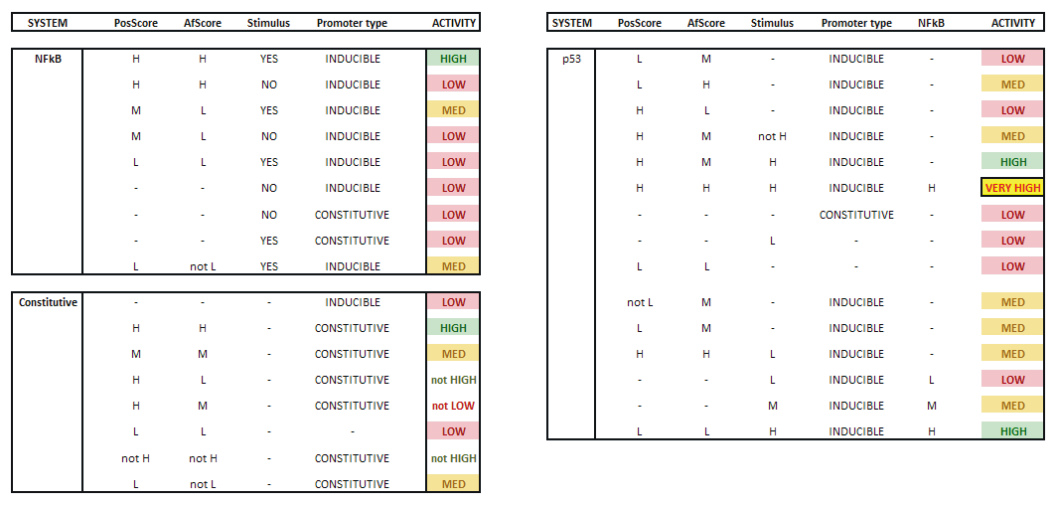
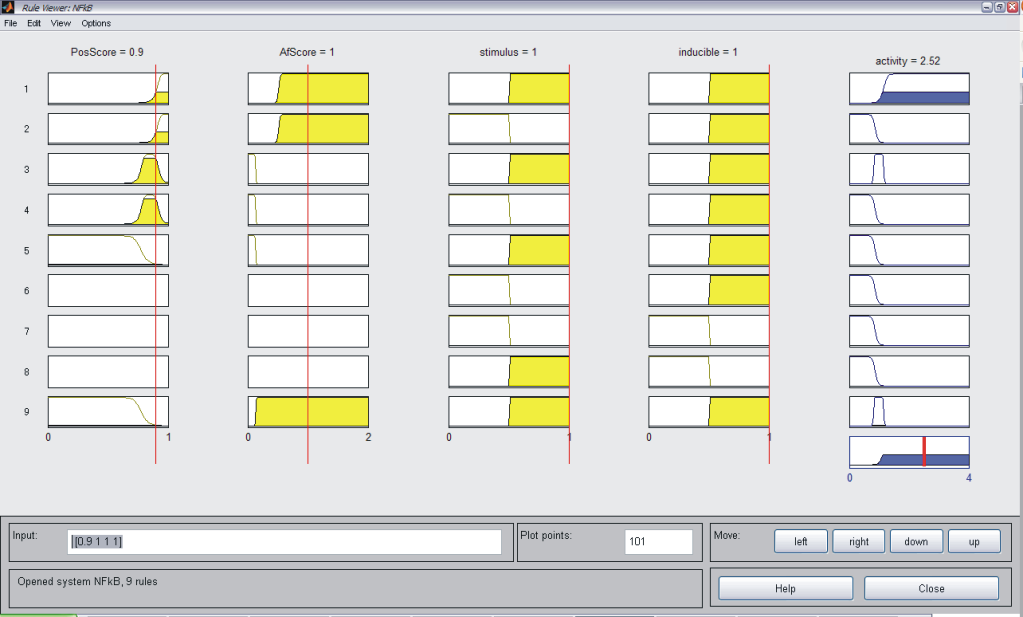
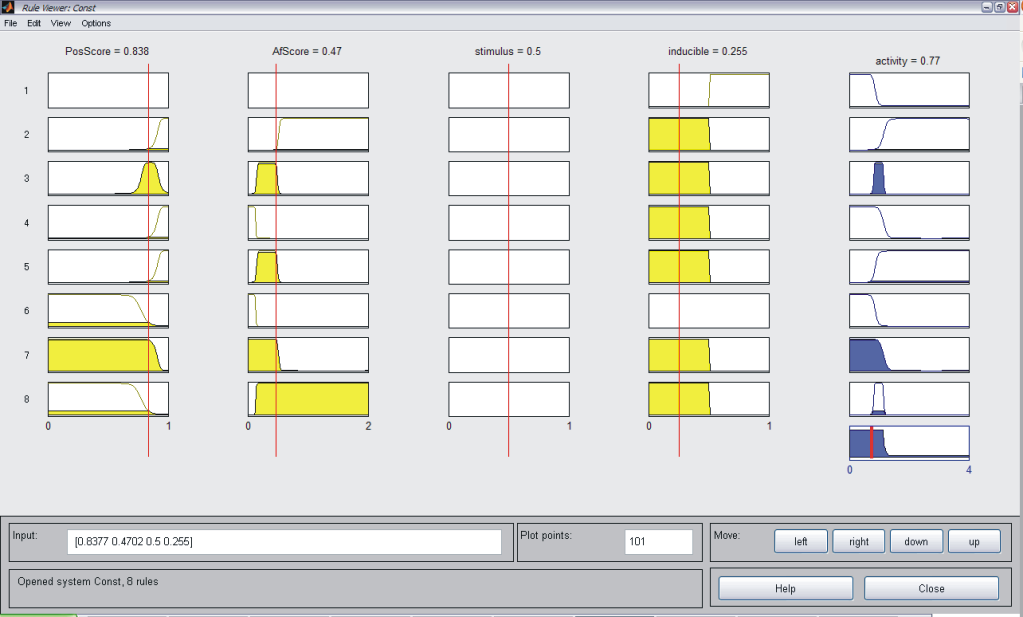
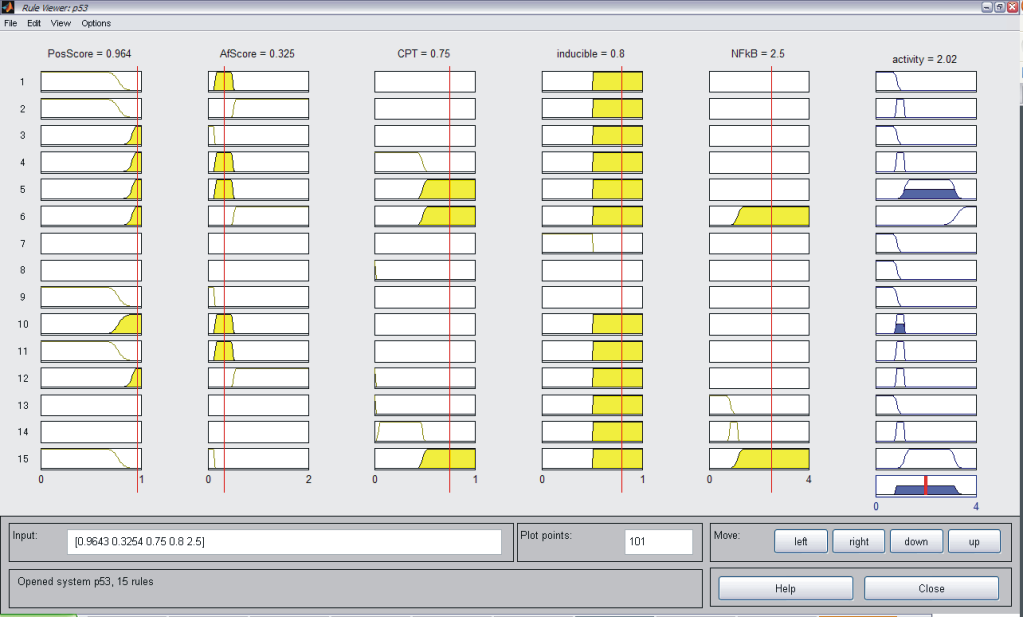
As described above, we considered JeT as a well established medium active promoter (taken as basal activity here). Based on this assumption, our system should be able to predict decreased level of activity when compared to CMV:JeT as well as an increased activity when compared to CMV (see Measurement). Our model should be able to reflect the behaviour of at least two inducible NFκB-responsive sequences (I L 12, II L 10). Having different position an affinity scores, these can be considered as "differential" promoters since they are of the same promoter type and induced by the same stimulus (TNFα), thus corresponding to the same pathway. Our model should also be able to describe the behaviour of at least six constitutive (NFκB responsive) promoters (L1, L4, L5, S4, S5, S10). These can be also taken as "differential" because the only differ in their sequence scores. Also, the system should be capable of reproducing the activity characteristics of eight p53 responsive promoters (S6, S8, S9, S12, S24, L8, L19, L22). They do not only differ in their physical sequence scores like in the two case above, but also differ in the concentration of the stimulating agent. Taking all these assumptions into account, we defined our rules which is described in Tab. 1. As an example for each single system the MATLAB fuzzy logic toolbox (FLT) rule viewer is displayed (Figs. A-C). The viewer shows the combination of the rules which are defined as an array which has the structure as follows: [position score; affinity score; stimulation; inducibilty; NFkB-activity (only for p53)]. All rules are combined with an AND gate. Combining Exclusive PathwaysTo see how we combine several exclusive pathways see results. [TOP] ResultsBased on our experimental measurements, subsequent data analysis as well as on our assumption we were able to design a network model which is based on fuzzy logic. This mathematical model reflected the characteristics of all investigated promoter sequences. This model was able to describe the behaviour of three standard promoters (JeT, CMV, CMV:JeT), six constitutively active promoters as well as at least ten inducible promoters (two NFκB- and eight p53-responsive) (data not shown) and it was also able to characterize the joined effect of distinct promoter combinations. This section briefly summarizes our results. Please click here to read more about our results! [TOP] Discussion[TOP] References[1] Harbison C. T., Gordon D. B., Lee T. I., Rinaldi N. J., Macisaac K. D., Danford T. W., Hannett N. M., Tagne J. B., Reynolds D. B., Yoo J., Jennings E. G., Zeitlinger J., Pokholok D. K., Kellis M., Rolfe P. A., Takusagawa K. T., Lander E. S., Gifford D. K., Fraenkel E., Young R. A. Transcriptional regulatory code of a eukaryotic genome. Nature 431: 99-104 (2004). [2] Hu Z., Killion P. J., Iyer, V. R. Genetic reconstruction of a functional transcriptional regulatory network. Nature Genet. 39: 683-687 (2007). [3] Gertz J., Siggia E. D., Cohen B. A. Analysis of combinatorial cis-regulation in synthetic and genomic promoters. Nature 457: 215-218 (2009) [4] Roider H. G., Kanhere A., Manke T., Vingron M. Predicting transcription factor affinities to DNA from a biophysical model. Bioinformatics 23: 134-141 (2006) [5] Carrera J., Rodrigo G., Jaramillo A. Towards the automated engineering of a synthetic genome. Mol. Biosyst. 5: 733-43 (2009). [6] Agapakis C. M., Silver P. A. Synthetic biology: exploring and exploiting genetic modularity through the design of novel biological networks. Mol. Biosyst. 5: 704-13 (2009) [7] Purnick P. E., Weiss R. The second wave of synthetic biology: from modules to systems. Nat Rev Mol Cell Biol. 10: 410-22 (2009). [8] Bhalerao K. D. Synthetic gene networks: the next wave in biotechnology? Trends Biotechnol. 27: 368-74 (2009). [9] Andrianantoandro E., Basu S., Karig D. K., Weiss R. Synthetic biology: new engineering rules for an emerging discipline. Mol Sys Biol (2006) [10] Alberts B., Johnson A., Walter P., Lewis J. Molecular Biology of the Cell, 5th edition, 2008. Garland Science, Chapter 6 [11] Vardhanabhuti S., Wang J., Hannenhalli S. Position and distance specificity are important determinants of cis-regulatory motifs in addition to evolutionary conservation. Nucl Acid Res 35: 3203-3213 (2007). [12] Yokoyama K. D., Ohler U., Wray G. A. Measuring spatial preferences at fine-scale resolution identifies known and novel cis-regulatory element candidates and functional motif-pair relationships. Nucl Acid Res 37: e92 (2009) [13] Nelles O. Nonlinear System Identification Springer Verlag GmbH & Co., Berlin, 2000. [14] Bosl W. J. Systems biology by the rules: hybrid intelligent systems for pathway modeling and discovery.
BMC Systems Biology1:13 (2007). [15] Laschov D., Margaliot M. Mathematical modeling of the lambda switch:a fuzzy logic approach. J Theor Biol. 21:475-89 (2009) [16] Aldridge B. B., Saez-Rodriguez J., Muhlich J. L., Sorger P. K., Lauffenburger D. A. Fuzzy logic analysis of kinase pathway crosstalk in TNF/EGF/insulin-induced signaling PLoS Comput Biol.5:e1000340 (2009).
[17] Roider H. G., Kanhere A., Manke T., Vingron M. Predicting transcription factor affinities to DNA from a biophysical model. Bioinformatics 23: 134-141 (2007). |
 "
"