Team:Aberdeen Scotland/WetLab/betagalactosidaseassay
From 2009.igem.org
Katriona87 (Talk | contribs) (→Introduction) |
Katriona87 (Talk | contribs) (→Methods) |
||
| Line 10: | Line 10: | ||
<i>E.coli strains:</i><br> | <i>E.coli strains:</i><br> | ||
| - | The XL1-Blue strain, usually used for | + | The XL1-Blue strain, usually used for blue-white screening during cloning, expresses the ω portion of LacZ, referred to as XL1-Blue[ωLacZ]. The Lac minus E.coli strain was transformed with the plasmid pRS425, which carries the gene for the α portion of β-Galactosidase. This is referred to as Lac-[αLacZ]. As positive control the plasmid pRS425 was also transformed into the XL1-Blue strain, therefore this strain now encodes both portions of β-Galactosidase and the two portions are reconstituted within cell environment (XL1-Blue[α/ωLacZ]). <br><br> |
<i>Sonication:</i><br> | <i>Sonication:</i><br> | ||
| - | A sonicator was used to lyse the | + | A sonicator was used to lyse the cells; cells were exposed to sound waves for approximately one minute until the cell suspension changed colour.<br><br> |
<i>Protein Assay:</i><br> | <i>Protein Assay:</i><br> | ||
| - | A standard curve of protein per millilitre versus absorbance was produced using | + | A standard curve of protein per millilitre versus absorbance was produced using BSA. Following that, the concentration of protein in the lysates was determined using a 10µl sample. The concentrations of proteins in the Lac-[αLacZ], XL1-Blue[ωLacZ] and XL1-Blue lysates were normalised, so that all samples had the same concentration of protein, and activity per minute per milligram protein could then be calculated.<br><br> |
<i>Beta-Galactosidase assay:</i><br> | <i>Beta-Galactosidase assay:</i><br> | ||
| - | Overnight cultures used in this assay (as described in “E. coli strains”) were grown in fresh | + | Overnight cultures used in this assay (as described in “E. coli strains”) were grown in fresh LB medium to an OD600 of about 0.6. Ampicillin was added to the strains transformed with the pRS425 plasmid. The 100 ml cultures were harvested by centrifugation, washed and resuspended in 1 ml of Z-Buffer. The cells were stored on ice for 20 min, and then sonicated before the main part of the assay was carried out. Lysate mixtures were then prepared, containing the alpha and omega portions from two different strains (Lac-[αLacZ] and XL1-Blue[ωLacZ]), in differing volumetric ratios. These ratios were 1:1, 4:1, and 1:4 (alpha to omega lysate). The lysate mixtures were incubated for differing lengths of time, to allow recombination of functional beta-galactosidase activity from the component alpha and omega portions, and beta-galactosidase activity was subsequently assayed. For all beta galactosidase reactions, 0.7mL of Z-Buffer, 0.2mL of ortho-nitrophenyl-β-galactoside and 0.1mL of lysates were used. After significant accumulation of the reaction product, the reaction was stopped by adding 0.5mL of sodium carbonate (Na2CO3).<br><br> |
| - | + | ||
| - | + | ||
| - | <br><br> | + | |
==Results== | ==Results== | ||
Revision as of 17:15, 13 October 2009
University of Aberdeen - Pico Plumber
Contents |
Introduction
The idea of our iGEM project involves the production of a biological two - component glue; when the bacteria, Eschericha coli, detects a breach of pipe integrity, expression of two genes, the 2 components of the glue, is triggered. The first component is the adhesive tropoelastin, which will be expressed within the cell plasma. The second component is an enzyme called lysyl oxidase, which functions as an adhesive hardener and will be expressed on the cell surface, by translational fusion with the outer membrane protein OmpX. Once the E.coli reaches the breach, it will lyse, releasing the two glue components. Subsequent reconstitution allows the proteins to start functioning as a glue, thereby healing the pipe.
The following experiments were performed in order to prove that the concept of mixing two components in vitro results in a functional enzyme / protein. This was shown by using the example of alpha and omega complementation of beta-galactosidase activity. The gene encoding beta-galactosidase can be split into two parts, termed alpha and omega portions, and each expressed separately. When both portions are expressed in the same cell, enzyme activity is reconstituted and its activity can be detected with a beta-galactosidase assay. In these experiments, the alpha and omega portions were expressed in separate cells, the two transformant types were lysed, and the lysates combined to test whether functional protein interactions can occur in crude bacterial lysates.
Methods
E.coli strains:
The XL1-Blue strain, usually used for blue-white screening during cloning, expresses the ω portion of LacZ, referred to as XL1-Blue[ωLacZ]. The Lac minus E.coli strain was transformed with the plasmid pRS425, which carries the gene for the α portion of β-Galactosidase. This is referred to as Lac-[αLacZ]. As positive control the plasmid pRS425 was also transformed into the XL1-Blue strain, therefore this strain now encodes both portions of β-Galactosidase and the two portions are reconstituted within cell environment (XL1-Blue[α/ωLacZ]).
Sonication:
A sonicator was used to lyse the cells; cells were exposed to sound waves for approximately one minute until the cell suspension changed colour.
Protein Assay:
A standard curve of protein per millilitre versus absorbance was produced using BSA. Following that, the concentration of protein in the lysates was determined using a 10µl sample. The concentrations of proteins in the Lac-[αLacZ], XL1-Blue[ωLacZ] and XL1-Blue lysates were normalised, so that all samples had the same concentration of protein, and activity per minute per milligram protein could then be calculated.
Beta-Galactosidase assay:
Overnight cultures used in this assay (as described in “E. coli strains”) were grown in fresh LB medium to an OD600 of about 0.6. Ampicillin was added to the strains transformed with the pRS425 plasmid. The 100 ml cultures were harvested by centrifugation, washed and resuspended in 1 ml of Z-Buffer. The cells were stored on ice for 20 min, and then sonicated before the main part of the assay was carried out. Lysate mixtures were then prepared, containing the alpha and omega portions from two different strains (Lac-[αLacZ] and XL1-Blue[ωLacZ]), in differing volumetric ratios. These ratios were 1:1, 4:1, and 1:4 (alpha to omega lysate). The lysate mixtures were incubated for differing lengths of time, to allow recombination of functional beta-galactosidase activity from the component alpha and omega portions, and beta-galactosidase activity was subsequently assayed. For all beta galactosidase reactions, 0.7mL of Z-Buffer, 0.2mL of ortho-nitrophenyl-β-galactoside and 0.1mL of lysates were used. After significant accumulation of the reaction product, the reaction was stopped by adding 0.5mL of sodium carbonate (Na2CO3).
Results
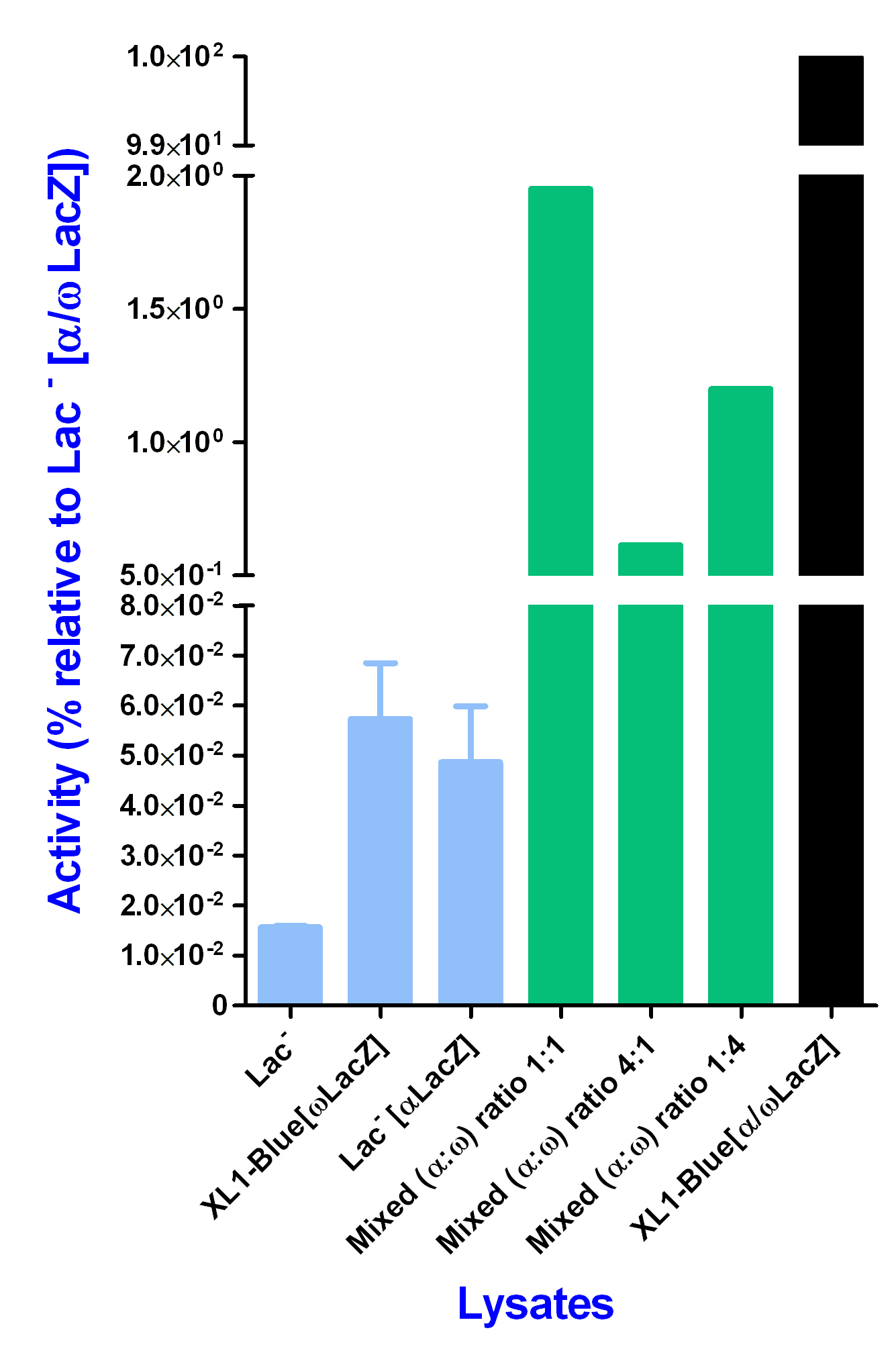
The activity gained by in vivo reconstitution of the alpha and omega subunit inside the XL1-Blue[α/ωLacZ] strain was said to be the maximum achievable and therefroe said to be 100% activity. All the other activities when given in percentage are compared to this maximum of activity.
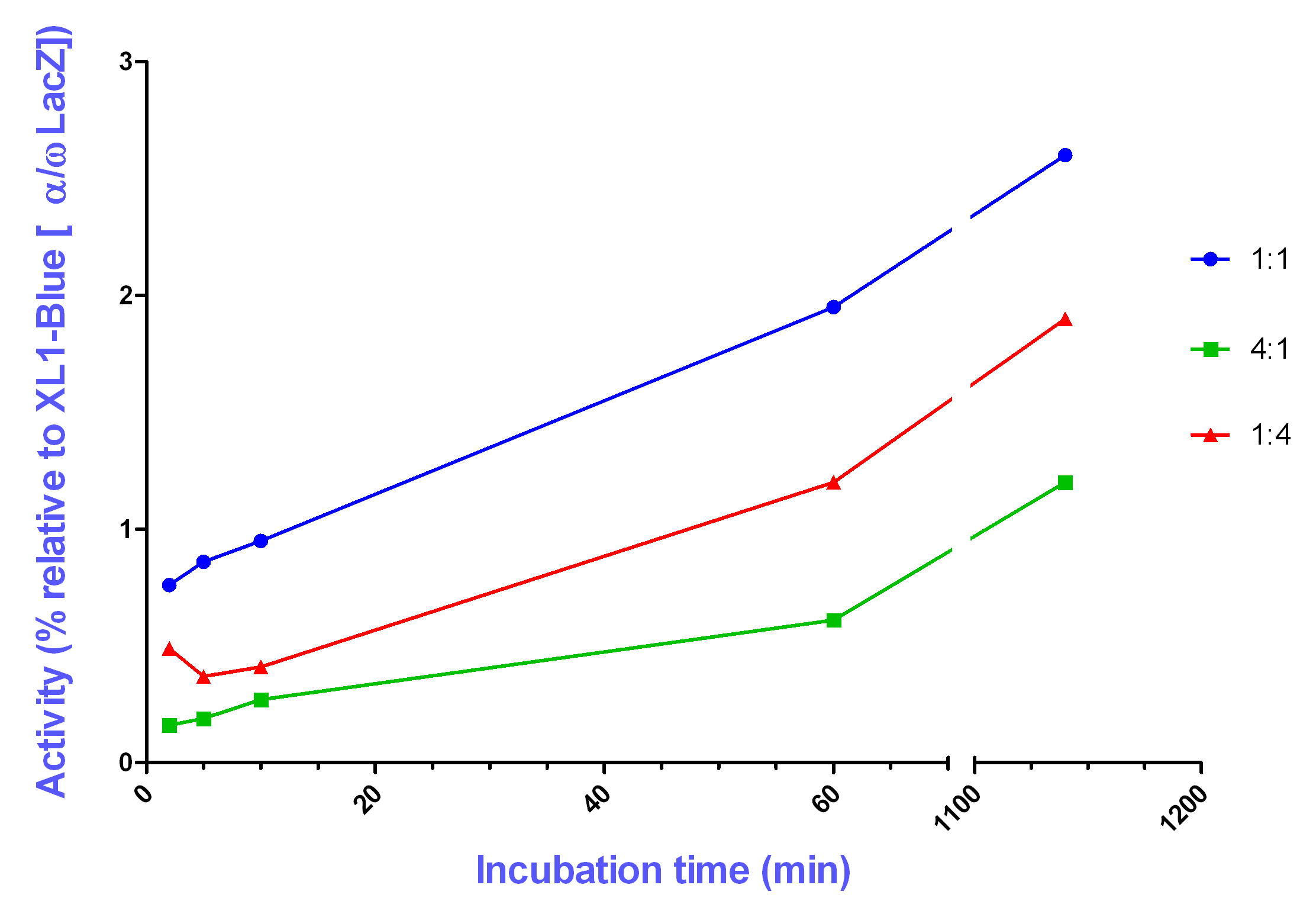
Generally, an increased enzymatic activity is achieved with increasing incubation time which allows the two components, alpha and omega, to be built together. Measurements were taken at an incubation time of 2 minutes, 5 minutes, 10 minutes, 60 minutes, and 19 hours. However, the activity is higher for a mixing ration of 1:1 (alpha to omega) than for the other ratios (4:1 and 1:4).
Never the less, at an incubation time of 60 minutes all three different ratios gave a significant higher activity in comparison with the negative ontrols, but much less activity than the in vivo reconstitution of the alpha and omega subunit.
Conclusion
It was demonstrated that in vitro reconstitution of a protein, from two component polypeptides, results in a functional protein; the alpha and omega subunits of the enzyme beta-galactosidase were combined, outside the cellular environment, and enzyme activity was successfully reconstituted.
The results suggest that the most effective reconstitution is achieved by mixing the alpha and omega portion in a 1:1 ratio. The achieved in vitro reconstituted enzymatic activity, in comparison to the in vivo reconstitution, however, was very low. It is possible that alternative buffer conditions could increase the in vitro reconstituted activity.
For most pre-incubation times tested, the enzyme activity was found to increase during the pre-incubation time. These results suggest that a longer pre-incubation time, permits the two components, alpha and omega, greater opportunity to associate and reconstitute beta-galactosidase activity. It is expected that the decreased activity observed, for 19 hours pre-incubation, was due to protein degradation.
This experiment has proved the concept of the 2-component glue; it successfully demonstrated that the mixing of two subunits of an enzyme, outside the cellular environment, can result in reconstitution of enzyme function.
 Back to Quorum Sensing Results Back to Quorum Sensing Results
|
Continue to Wet Lab Procedures 
|
 "
"