Team:British Columbia/Project
From 2009.igem.org
m (Team:British Columbia/Tools moved to Team:British Columbia/Project over redirect) |
(→'Overview of the Traffic Light Biosensor: A flexible, modular, and transparent system for multi-level assessment of variable inputs.') |
||
| Line 4: | Line 4: | ||
== 'Overview of the Traffic Light Biosensor: <br> A ''flexible'', ''modular'', and ''transparent'' system for multi-level assessment of variable inputs.' == | == 'Overview of the Traffic Light Biosensor: <br> A ''flexible'', ''modular'', and ''transparent'' system for multi-level assessment of variable inputs.' == | ||
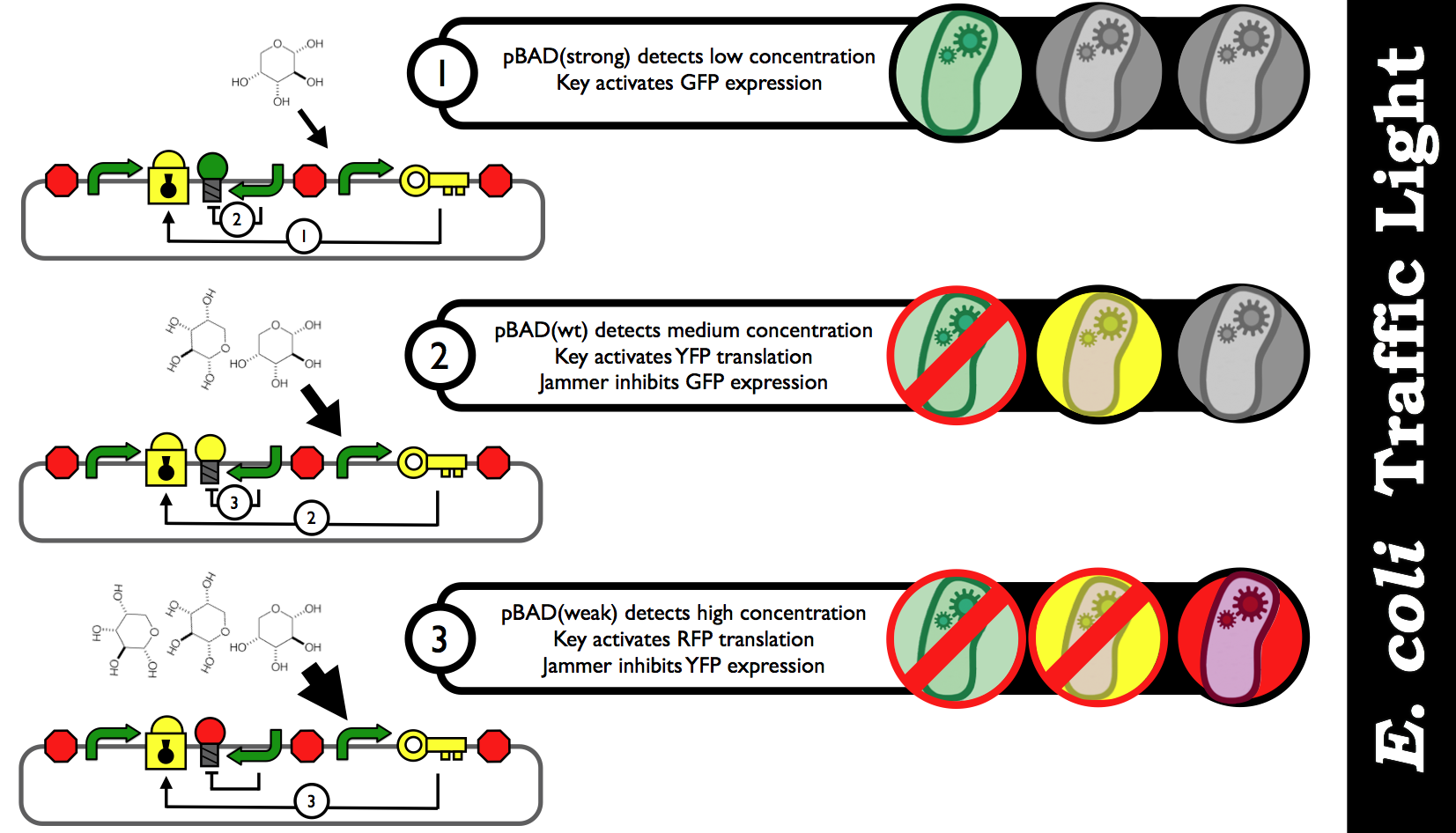
| - | + | Depending on the concentration of a particular substrate in the medium, E. coli will respond accordingly by producing different coloured fluorescence proteins. A diagram would look like this: | |
[[Image:E_coli_Traffic_Light_Subprojects.png|center|thumb||400px|The ''E. coli'' Traffic Light Biosensor is composed of three major subparts: variable arabinose-inducible promoters, RNA lock and key system, and reverse antisense promoters for input detection, color activation and traffic light switching respectively.]] | [[Image:E_coli_Traffic_Light_Subprojects.png|center|thumb||400px|The ''E. coli'' Traffic Light Biosensor is composed of three major subparts: variable arabinose-inducible promoters, RNA lock and key system, and reverse antisense promoters for input detection, color activation and traffic light switching respectively.]] | ||
| - | + | Here is the design for how we can build our "traffic light". | |
| + | [[Image:E_coli_Traffic_Light_Step_by_Step.png|thumb|center|700px|Schematic black-box representation of the E. coli Biosensor that detects various concentration inputs and color outputs. The idea is discrete analog outputs based on a user-specified threshold for each range of concentration.]] | ||
| - | + | Please go to the sub pages for more information on the subparts:<br> | |
1. [https://2009.igem.org/Team:British_Columbia/pBAD A variable sensitivity biosensor]<br> | 1. [https://2009.igem.org/Team:British_Columbia/pBAD A variable sensitivity biosensor]<br> | ||
2. [https://2009.igem.org/Team:British_Columbia/LockandKey A lock-and-key logic gate system]]<br> | 2. [https://2009.igem.org/Team:British_Columbia/LockandKey A lock-and-key logic gate system]]<br> | ||
Revision as of 02:45, 22 October 2009
'Overview of the Traffic Light Biosensor:
A flexible, modular, and transparent system for multi-level assessment of variable inputs.'
Depending on the concentration of a particular substrate in the medium, E. coli will respond accordingly by producing different coloured fluorescence proteins. A diagram would look like this:
Here is the design for how we can build our "traffic light".
Please go to the sub pages for more information on the subparts:
1. A variable sensitivity biosensor
2. A lock-and-key logic gate system]
3. An antisense "off" switch
Miscellaneous Data
We also produced a couple tools to help out the project:
- Biobricks.zip - Fasta file containing every biobrick from Here
- http://www.pkts.ca/bb - Biobrick digestion engine - enter the name of a biobrick plasmid and biobrick insert, and this will show you the product of an EcoRI and PstI digestion/ligation as a FASTA file (suitable for viewing in your favorite program).
- http://www.pkts.ca/brickedit/ - Biobrick picture maker - enter a sequence of letters corresponding to the icons, and the program will produce a concatenated file of the Biobrick.
Links
http://rna.tbi.univie.ac.at/ - a package of prediction tools for RNA structures; we used RNAfold to annotate the key and lock structures
http://mobyle.pasteur.fr/cgi-bin/portal.py - a set of web-accessible bioinformatics tools including Mfold, which determines 2D RNA structure and draws it
http://frodo.wi.mit.edu/ - Primer3, a primer design program
 "
"