Team:British Columbia/Project
From 2009.igem.org
(Difference between revisions)
(→Traffic Light Overview) |
|||
| (61 intermediate revisions not shown) | |||
| Line 1: | Line 1: | ||
| - | { | + | {{Template:UBCiGEM2009_menu_home}} |
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
<!--- The Mission, Experiments ---> | <!--- The Mission, Experiments ---> | ||
| - | + | ==Traffic Light Overview== | |
| - | + | Depending on the concentration of a particular substrate in the medium, E. coli will respond accordingly by producing different coloured fluorescence proteins. A diagram would look like this: | |
| - | + | ||
| - | + | [[Image:E_coli_Traffic_Light_Subprojects.png|center|thumb||600px|The ''E. coli'' Traffic Light Biosensor is composed of three major subparts: variable arabinose-inducible promoters, RNA lock and key system, and reverse antisense promoters for input detection, color activation and traffic light switching respectively.]] | |
| - | + | Here is what's happening inside our traffic light: | |
| - | + | ||
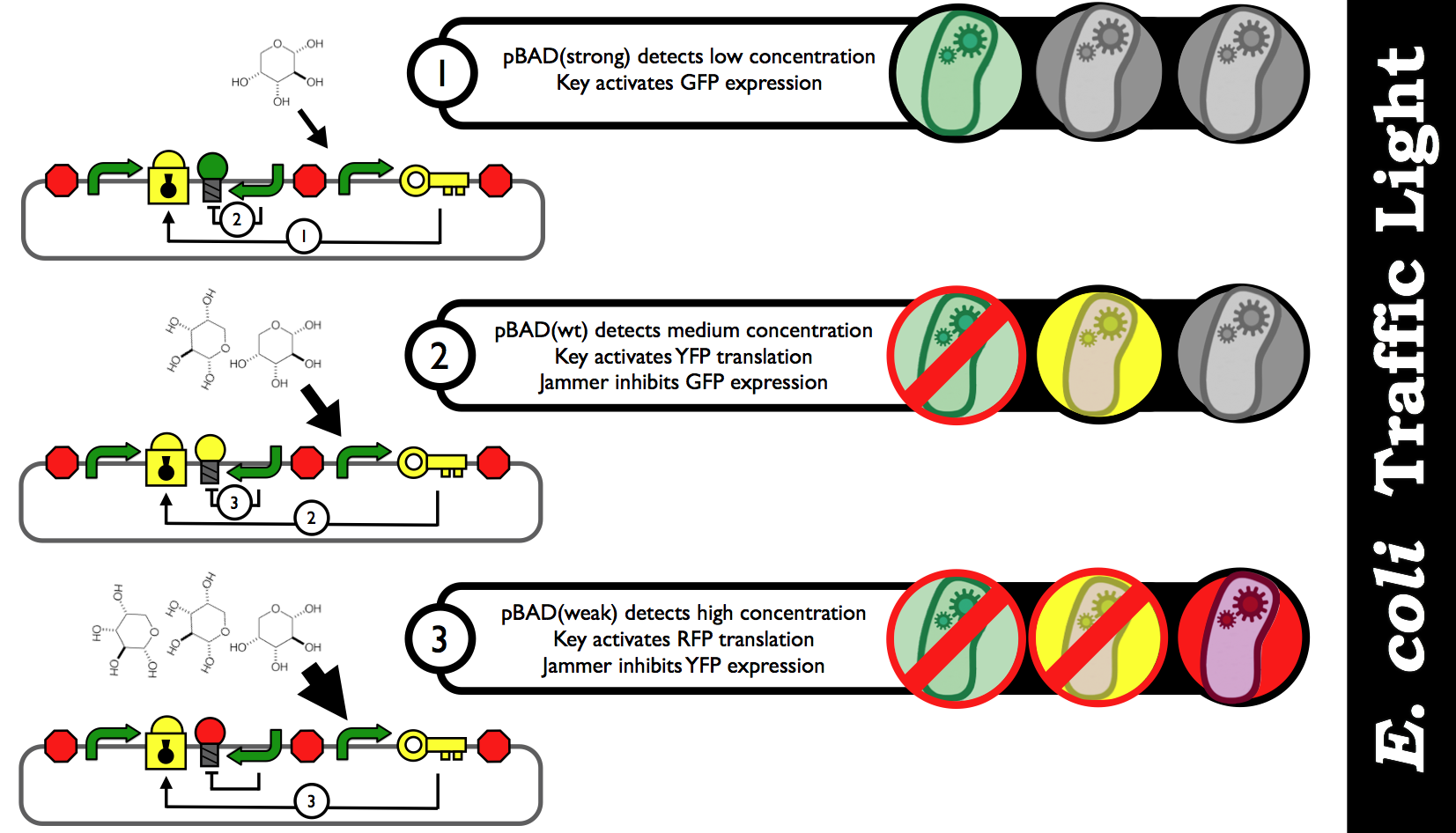
| - | + | [[Image:E_coli_Traffic_Light_Step_by_Step.png|thumb|center|850px|Schematic black-box representation of the E. coli Biosensor that detects various concentration inputs and color outputs. The idea is discrete analog outputs based on a user-specified threshold for each range of concentration.]] | |
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | For our ideas to work, we will need:<br> | |
| + | 1. [https://2009.igem.org/Team:British_Columbia/pBAD A variable sensitivity biosensor]<br> | ||
| + | 2. [https://2009.igem.org/Team:British_Columbia/LockandKey A lock-and-key logic gate system]]<br> | ||
| + | 3. [https://2009.igem.org/Team:British_Columbia/Jammer An antisense "off" switch] | ||
| + | |||
| + | |||
| + | <!-- !!!!!!!make sure the spacer is enough so that the image doesn't run into the next section --> | ||
| + | <br><br><br> | ||
| + | |||
| + | <!-- | ||
== Logic Gates == | == Logic Gates == | ||
| + | In 2004, Isaacs et al. (4) constructed a lock and key system termed the riboregulator. Briefly, the riboregulator consists of a lock sequence immediately 5' of and complementary to the RBS; upon transcription, the lock binds to the RBS and prevents translation of the mRNA. This binding can be "unlocked" by the introduction of a "key" mRNA, which disrupts the binding between the lock and RBS and allows translation to resume. | ||
| + | |||
| + | We intend to convert the lock and key as described by Isaacs et al. into standardized BioBrick parts and integrate them with our library of variable strength promoters in order to create a multi-level logic gate system. We also hope to design an additional component, termed the "jammer", which prevents the key from unlocking the lock. | ||
| + | |||
| + | [[Image:UBC2009-Key-lock_roadmap.jpg|700x700px|center]] | ||
| + | |||
| + | ==Lock and Key== | ||
| + | |||
| + | The key is a short RNA segment that is the reverse-complement of the lock, which binds to the lock, unfolding the hairpin that conceals the ribosome binding site. Ribosomes should then start translating the RNA that was locked. | ||
| + | |||
| + | ==Jammer== | ||
| + | |||
| + | Our analog biosensor requires a component to turn off designed lights (i.e. reporter genes) when the target passes a certain threshold. To do so, we have designed a modular, endogenous RNA knockdown system using antisense RNA hybridization. We have independently conceived of using a reverse promoter, that was first described by O’Connor and Timmis (1987), to generate an antisense transcript that hybridizes to the sense transcript, to reduce gene expression. | ||
| + | |||
| + | This is accomplished at the RNA-level through both transcription and translation interfaces. First, when the reverse promoter recruits RNA polymerases, they transcribe in the antisense direction. This causes the sense and anti-sense oriented polymerases to collide, thus reducing transcription (Ward and Murray, 1979; Crampton et al. 2006). | ||
| + | |||
| + | ==Roadmap== | ||
| + | |||
| + | As of the 2009 Jamboree, not all aspects of our project have been completed. Looking forward, we intend to complete, test, and characterize our lock, key and jammer. Once this is done we intend to do the same for a second set of lock, key and jammer that have different sequences and will therefore not interfere with the first set. This second set will be configured to detect a second molecule, distinct from the first. When this is completed, we will try adding logic gates, allowing us to address nine different combinations: high, medium and low concentrations of arabinose combined with high, medium and low concentrations of whatever we choose as our second molecule, possibly an antibiotic. When plated on media with two perpendicular gradients, we should be able to independently control each square of a 3x3 grid, possibly displaying 9 different colors of fluorescent proteins, or generating other metabolites, such as indigo. | ||
| + | |||
| + | --> | ||
| + | |||
| + | == Tools used and produced == | ||
| + | |||
| + | To assist our project, we produced a Biobrick digestion engine and Biobrick picture maker to help out the project: | ||
| + | |||
| + | :http://www.pkts.ca/bb - Biobrick digestion engine - enter the name of a biobrick plasmid and biobrick insert, and this will show you the product of an EcoRI and PstI digestion/ligation as a FASTA file (suitable for viewing in your favorite program). | ||
| + | :http://www.pkts.ca/brickedit/ - Biobrick picture maker - enter a sequence of letters corresponding to the icons, and the program will produce a concatenated file of the Biobrick. | ||
| + | |||
| + | Also, we generated a handy Fasta file containing every biobrick from [http://partsregistry.org/cgi/partsdb/pgroup.cgi?pgroup=List Here]: | ||
| + | :[[Media:Biobricks.zip|Biobricks.zip]] - Fasta file containing every biobrick | ||
| + | |||
| + | We also found the following tools very helpful: | ||
| + | |||
| + | :http://rna.tbi.univie.ac.at/ - a package of prediction tools for RNA structures; we used RNAfold to annotate the key and lock structures | ||
| + | |||
| + | :http://mobyle.pasteur.fr/cgi-bin/portal.py - a set of web-accessible bioinformatics tools including Mfold, which determines 2D RNA structure and draws it | ||
| + | |||
| + | :http://frodo.wi.mit.edu/ - Primer3, a primer design program | ||
Latest revision as of 03:08, 22 October 2009
[http://www.ubc.ca ]
[http://www.ubc.ca ]
Traffic Light Overview
Depending on the concentration of a particular substrate in the medium, E. coli will respond accordingly by producing different coloured fluorescence proteins. A diagram would look like this:
Here is what's happening inside our traffic light:
For our ideas to work, we will need:
1. A variable sensitivity biosensor
2. A lock-and-key logic gate system]
3. An antisense "off" switch
Tools used and produced
To assist our project, we produced a Biobrick digestion engine and Biobrick picture maker to help out the project:
- http://www.pkts.ca/bb - Biobrick digestion engine - enter the name of a biobrick plasmid and biobrick insert, and this will show you the product of an EcoRI and PstI digestion/ligation as a FASTA file (suitable for viewing in your favorite program).
- http://www.pkts.ca/brickedit/ - Biobrick picture maker - enter a sequence of letters corresponding to the icons, and the program will produce a concatenated file of the Biobrick.
Also, we generated a handy Fasta file containing every biobrick from [http://partsregistry.org/cgi/partsdb/pgroup.cgi?pgroup=List Here]:
- Biobricks.zip - Fasta file containing every biobrick
We also found the following tools very helpful:
- http://rna.tbi.univie.ac.at/ - a package of prediction tools for RNA structures; we used RNAfold to annotate the key and lock structures
- http://mobyle.pasteur.fr/cgi-bin/portal.py - a set of web-accessible bioinformatics tools including Mfold, which determines 2D RNA structure and draws it
- http://frodo.wi.mit.edu/ - Primer3, a primer design program
 "
"