Team:Calgary/13 August 2009
From 2009.igem.org
(Difference between revisions)
Babydimples (Talk | contribs) |
Jamie.feng (Talk | contribs) |
||
| (18 intermediate revisions not shown) | |||
| Line 194: | Line 194: | ||
<br> | <br> | ||
<div class="heading"> | <div class="heading"> | ||
| - | + | TO THE RADIO STATION | |
</div> | </div> | ||
<br> | <br> | ||
<div class="desc"> | <div class="desc"> | ||
</html> | </html> | ||
| - | + | ||
| + | At 1:00 Me , Emily , Fadh and Mandy went to MacHall , the CJWS studio for an interview . It went very well and was an interesting experience. | ||
| + | |||
| + | After the interview we began preparing for the Fort Mcmurray presentation Friday . Building slide order , research detail etc. | ||
<html> | <html> | ||
| Line 221: | Line 224: | ||
<br> | <br> | ||
<div class="heading"> | <div class="heading"> | ||
| - | + | Glycerol Stocks, Plasmid Switch, Transformation and radio interview | |
</div> | </div> | ||
<br> | <br> | ||
<div class="desc"> | <div class="desc"> | ||
</html> | </html> | ||
| - | + | ||
| + | |||
| + | *This morning I made glycerol stocks of my completed and sequenced Mutant Circuit! | ||
| + | |||
| + | *Today me and Vicki started a Plasmid switch of her Mutant Circuit from psB1AK to psB1AC3. In order to test the reporter circuit, we need to transform the mutant circuits into Kevin's reporter circuit which has qrr4 and GFP. Because this circuit is in a vecotr with K resistance however, we can't have either of the mutants in AK as this would not be good selection pressure. To deal with this, we shall move Vicki's circuit into the psB1AC3 vector; the vector that my circuit is currently in. To do this we set up an overnight restriction digest of J13002-LuxOD47A-B0015 and the psB1AC3 vector with XbaI and PstI retriction enzymes at 37 C. Tomorrow we will do antarctic phosphotase and ligation of this followed by verification. | ||
| + | *I also did two transformations today: one of my mutant circuit (J13002-LuxOD47E-B0015) into KT1144 cells, and one of my mutant circuit into TOP10 cells with the qrr4 promoter and GFP. The KT1144 celles were plated on C and K plates and we made KC plates and plated the mutants in TOP10 on these plates. We left these for overnight growth and will check them for glow tomottow morning. | ||
| + | |||
| + | *Prima, Vcicki, Chinee and I talked about and wrote out a plan for the Fort Mac presentation tomorrow. We planned out the slides and divided up who is going to do which ones tonight. We'll compile them and practice the presentation tomrrow morning. | ||
| + | |||
| + | *Today Fahd, Chinee, Mandy and I also went for an interview with Joe Burima from CJSW. We talked about the project in general and then briefly went over the sub projects, focusing on lab, outreach and Second Life. We also talked a bit about aGEM and iGEM and Joe invited us to come back in November after iGEM, so that we can talk about it. | ||
| + | |||
<html> | <html> | ||
| Line 247: | Line 260: | ||
<br> | <br> | ||
<div class="heading"> | <div class="heading"> | ||
| - | + | CJSW Interview and Marketing Blog | |
</div> | </div> | ||
<br> | <br> | ||
<div class="desc"> | <div class="desc"> | ||
</html> | </html> | ||
| - | + | Today I started my day by writing up a blog for the marketing team. The blog summarized are marketing team activity for the past two weeks. I included events such as Bake sale and the Oil Sands tour and our new sponsors (VWR Scientific and Corning Life Sciences). The blog has been posted on www.igemcalgary.blogspot.com . | |
| + | <br> | ||
| + | <br> | ||
| + | I also prepared for our radio interview with our University of Calgary Community Radio Station (C JSW 90.9). Some of the things we discussed with Joe Burima (the radio jockey) included such as details about iGEM, Synthetic Biology and our project. I want to sincerely thank CJSW and Joe for their support and time. | ||
| + | |||
<html> | <html> | ||
| Line 273: | Line 290: | ||
<br> | <br> | ||
<div class="heading"> | <div class="heading"> | ||
| - | + | Membrane Computing Visualization in Progress | |
</div> | </div> | ||
<br> | <br> | ||
<div class="desc"> | <div class="desc"> | ||
</html> | </html> | ||
| - | + | Making a movie/animation for complete cascade when AI2 is present/not present. We are also working on our paper to be handed out tomorrow. | |
| - | + | ||
<html> | <html> | ||
</div> | </div> | ||
| Line 299: | Line 315: | ||
<br> | <br> | ||
<div class="heading"> | <div class="heading"> | ||
| - | + | Preparing for AI-2 activity assay | |
</div> | </div> | ||
<br> | <br> | ||
<div class="desc"> | <div class="desc"> | ||
| - | </ | + | Made LM media and autoinducer bioassay media for culturing <i>Vibrio harveyi</i> and <i>Salmonella typhimurium</i>. Also set up overnight cultures for AI-2 extraction and reporter strains. |
| - | + | ||
| - | + | ||
| - | < | + | |
</div> | </div> | ||
</td> | </td> | ||
| Line 325: | Line 338: | ||
<br> | <br> | ||
<div class="heading"> | <div class="heading"> | ||
| - | + | PCR to verify presence of LuxPQ-LuxOU construct in pCS26 in the correct direction | |
</div> | </div> | ||
<br> | <br> | ||
<div class="desc"> | <div class="desc"> | ||
</html> | </html> | ||
| - | + | The pCS26-S-F primer was designed to anneal on the 5’ end of the multiple cloning site of the pCS26 vector, and its use is to help verify (in conjunction with the LuxPQ-Reverse primer) that LuxPQ-LuxOU was cloned in the correct direction. This primer was synthesized then diluted into working stock concentrations. A PCR was set up with plasmid from eight colonies of the PQ-OU in pCS26, with the following two pairs of primers: pCS26-S-F and LuxPQ-Reverse; pCS26-S-F and LuxOU-Reverse. The conditions for this PCR were as follows: 94ºC for 3 minutes; 36X (94ºC for 30 seconds; 55ºC for 45 seconds; 72ºC for 4 min(pCS26-S-F and LuxPQ-R) or for 6 min 20 sec (pCS26-S-F and LuxOU-R)); 72ºC for 10 minutes; held at 4ºC overnight. For results, see August 14. | |
<html> | <html> | ||
| Line 351: | Line 364: | ||
<br> | <br> | ||
<div class="heading"> | <div class="heading"> | ||
| - | + | Modifying Replication and Updating Notebook | |
</div> | </div> | ||
<br> | <br> | ||
<div class="desc"> | <div class="desc"> | ||
</html> | </html> | ||
| - | + | Primase for DNA replication now rezzes a primer on the top strand as well and in the proper orientation and the DNA polymerase now rezzes the nucleotides in the proper orientation as well along the leading strand although since it is in a testing stage the nucleotides will remain temporary for now. I have also started writing the script that will lock and unlock activities until it is time for the actions to be performed. | |
| + | <br> | ||
| + | <br> | ||
| + | Things left to do for replication: | ||
| + | <br> | ||
| + | <br> | ||
| + | * Get it user friendly - I will now have to make it easier to interact with the exhibit, so avatars may specify that they have to add primers before the DNA polymerase may add nucleotides etc. The user can be asked questions and if the user is correct something ends up happening and for example, if the polymerase is not ready then give a hint. "You still need to make some primers so that the polymerase knows where to begin replication. Do you know what enzyme can do this?" | ||
| + | * Make a second polymerase for the lagging strand. If polymerase two says it is ready the position should be set to this and then rez nucleotides every 0.3 meters, | ||
| + | * Label the strands including what end it is (5’ or 3’) | ||
| + | |||
| + | I also updated May 4<sup>th</sup> and 11<sup>th</sup> in the wiki notebook and looked into constructivist theory in an attempt to find out more about the reasons as to why education in Second Life has the possibility to be a successful educational platform. | ||
<html> | <html> | ||
| Line 377: | Line 400: | ||
<br> | <br> | ||
<div class="heading"> | <div class="heading"> | ||
| - | + | Competent cells | |
</div> | </div> | ||
<br> | <br> | ||
<div class="desc"> | <div class="desc"> | ||
</html> | </html> | ||
| - | + | The TOP10 cells with <i>Pqrr4</i>+I13500 plasmid have been made competent again in order to test the mutants. | |
| - | + | <html> | |
| + | </div> | ||
| + | <br> | ||
| + | <div class="heading"> | ||
| + | Worked on symbiology models | ||
| + | </div> | ||
| + | <br> | ||
| + | <div class="desc"> | ||
| + | </html> | ||
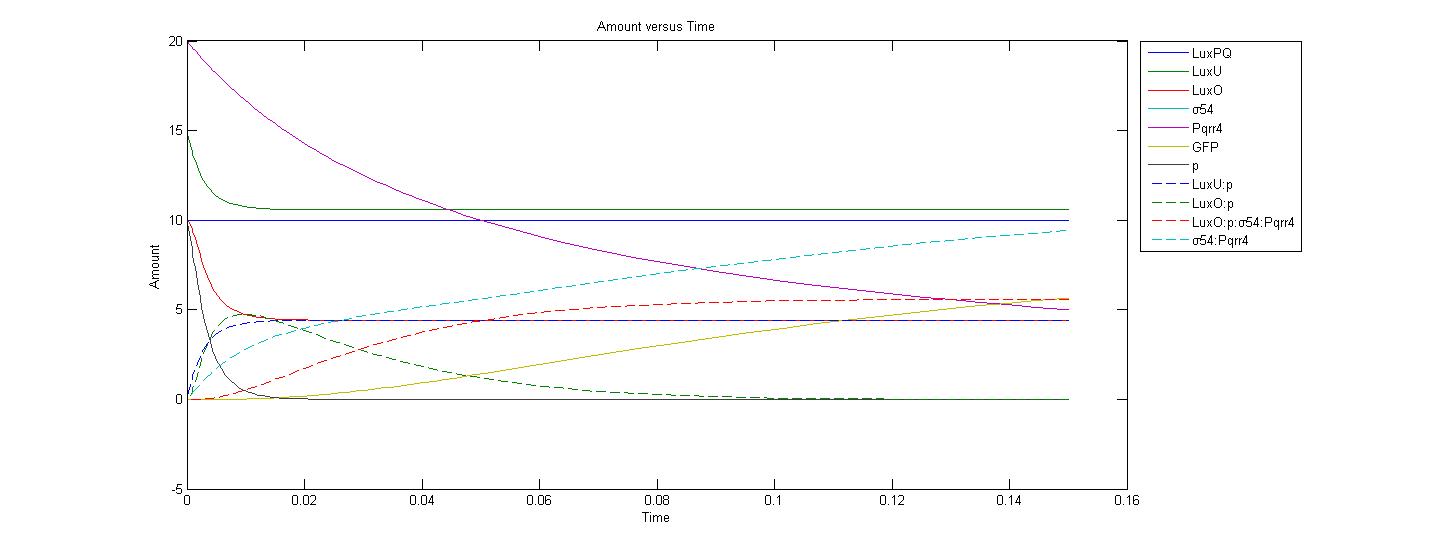
| + | Our previous AI-2 model has been improved, and now it is able to produce a visual representation of our model. The following graph represents the system without AI-2, and when there are 20 Pqrr4 promoters in the cell: | ||
| + | <br> | ||
| + | [[Image:Calgary_WithoutAI2graph3.jpg|700px]] | ||
| + | <br> | ||
| + | <br> | ||
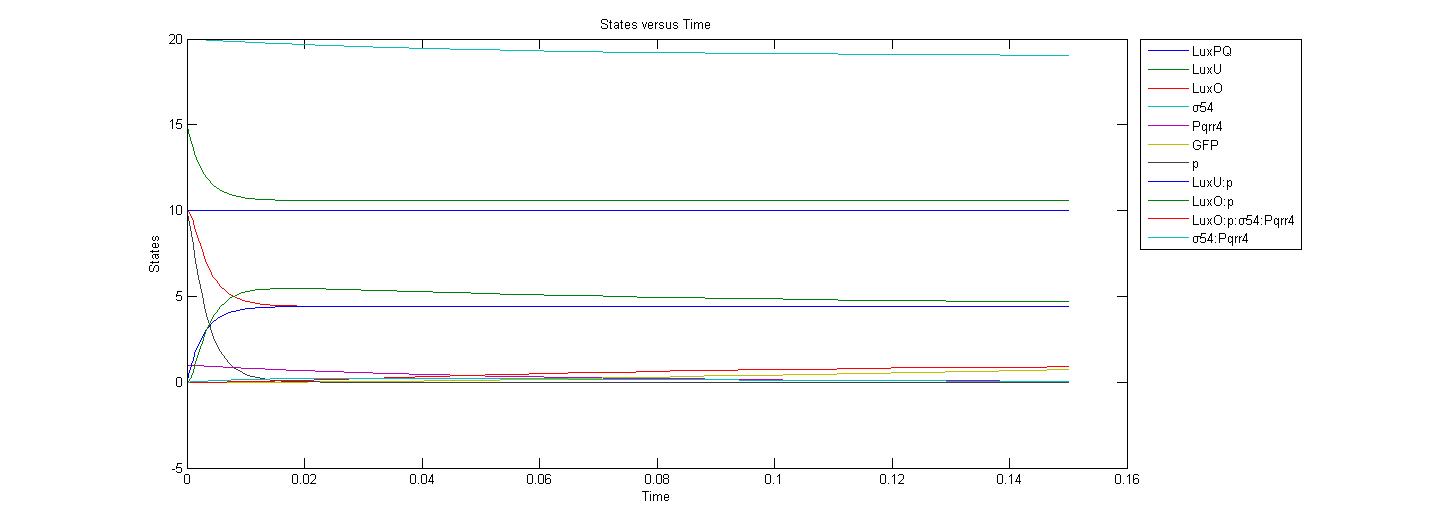
| + | The next graph represents the system without AI-2, and when there is only 1 Pqrr4 promoter in the cell: | ||
| + | <br> | ||
| + | [[Image:Calgary_WithoutAI2graph4.jpg|700px]] | ||
| + | <br> | ||
| + | <br> | ||
| + | Although most of the initial values and reaction rates of the parameters are not accurate, one can see from above graphs that the production of GFP drastically decreases when the amount of <i>Pqrr4</i> promoter decreases from 20 to 1. | ||
<html> | <html> | ||
</div> | </div> | ||
| Line 403: | Line 445: | ||
<br> | <br> | ||
<div class="heading"> | <div class="heading"> | ||
| - | + | Biobricker Design Continued and Misc. | |
</div> | </div> | ||
<br> | <br> | ||
<div class="desc"> | <div class="desc"> | ||
</html> | </html> | ||
| - | + | I completed all of the proteins, promoters, and other coding sequence buttons for the biobricker, continuing from yesterday. In addition, I also made menu buttons for multiple functions required for the biobricker, including: | |
| - | + | *Help | |
| + | *Unbind | ||
| + | *Delete | ||
| + | <br> | ||
| + | In addition, as mentioned above, we had our CJSW interview today, in order to let more people on campus (and in Calgary) know about our project. | ||
| + | <br> | ||
| + | Did a little bit of reading on the Constructivist Theory, to aid in justification of our activities in Second Life. | ||
<html> | <html> | ||
| + | <br> | ||
</div> | </div> | ||
</td> | </td> | ||
| Line 462: | Line 511: | ||
<br> | <br> | ||
<div class="heading"> | <div class="heading"> | ||
| - | + | Continuation of Restriction Digest of aiiA-B0015 into psB1AK3 | |
</div> | </div> | ||
<br> | <br> | ||
<div class="desc"> | <div class="desc"> | ||
</html> | </html> | ||
| - | + | This morning, I took out the insert and vector tubes from the water bath and phosphatased the vector (psB1AK3), ligated and transformed my construct (aiiA-B0015) into Top 10 cells. I plated the cells on Kan plates since the construct is in the psB1AK3 vector. I left the plates in the incubator to grow overnight at 37 degrees. I also picked up the gene-specific aiiA primers from the synthesis lab. | |
| - | + | I spoke with Emily and Fahd about how to prepare and present the Fort-Mac presentation for tomorrow. Then Emily and I worked to make the slides and add pictures and animations. | |
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| + | I called a few new companies but unfortunately, I couldn't get a hold of anyone. I left messages and my contact information and I'll follow up with them later this week. | ||
<html> | <html> | ||
</div> | </div> | ||
| Line 514: | Line 540: | ||
<br> | <br> | ||
<div class="heading"> | <div class="heading"> | ||
| - | + | Parameter optimisation, presentation preparation and plasmid transformation | |
</div> | </div> | ||
<br> | <br> | ||
<div class="desc"> | <div class="desc"> | ||
</html> | </html> | ||
| - | + | Today involved a miscellany of activities across the board: | |
| + | * I experimented with the SimBiology platform and have found a function that estimates model parameters to fit a set of data. I managed to get the example to work and am working on implementing that for our signalling pathway so that we can optimise our parameters as soon as we have data to work with. To accomplish this, I made up a set of data, with the idea that if I can get this function to spit out a set of parameters that suit the fabricated data, it will also be useful in spitting out a set of parameters that suit real data as soon as it comes. | ||
| + | * I also explored the sensitivity analysis. I'm not through with it yet, but the gist of it is that it will allow us to assess how sensitive the output it to changes in the different parameters. In layman's terms, this tells us which parameters can be adjusted and result in large changes in the output, versus the parameters that, when tweaked, won't give us any change in output worth mentioning. | ||
| + | * I investigated the possibility of setting up different compartments to represent different cells. I didn't make it too far as I began working on the parameter estimation process instead, but my goal is to figure out a representation at the command line level so that we can set up 10^8 or more compartments (to represent 10^8 cells present in culture) without having to insert all of them manually. | ||
| + | * We also explored areas in which synthetic biology could be useful on the oilpatch. My main area of interest was in using enzymes for cracking purposes on the bitumen, as an alternate approach to lowering its viscosity and reducing the temperature/pressure required to allow it to pass through a pipe. This was met with little success, as metallic catalysts seem to be the norm in industry and there aren't any research papers or patents on enzymatic cracking processes. Enzymes are used in other parts of the oilpatch, such as Greenzyme (TM), which releases hydrocarbons from carbon surfaces and is used in maximising oil collection. I am currently generating other ideas for useful synthetic biology projects in this area. | ||
| + | * We began preparations for mutant circuit testing. As my mutant (LuxO D47A) was on an AK plasmid and our reporter circuit is on a K plasmid, I prepared a restriction digest with <i>Xba1</i> and <i>Pst1</i> for eventual switch into an AC plasmid. | ||
<html> | <html> | ||
Latest revision as of 03:38, 22 October 2009
UNIVERSITY OF CALGARY

 "
"