|
|
| (12 intermediate revisions not shown) |
| Line 336: |
Line 336: |
| | Using a 96 well microplate, AI-2 isolated from <i>Salmonella typhimurium</i> 14028 supernatant will be tested. <i>V. harveyi</i> reporter strain MM32 (luxS- and luxN-) will luminesce in the presence of AI-2. Controls used in the experiment include <i>S. typhimurium</i> SS007, <i>V. harveyi</i> BB120 and <i> E. coli</i> DH5a. Overnight OD600 and luminescence (CPS) readings will be taken by a VICTOR (thank you Surette lab, especially Margot!). <br /><br /> | | Using a 96 well microplate, AI-2 isolated from <i>Salmonella typhimurium</i> 14028 supernatant will be tested. <i>V. harveyi</i> reporter strain MM32 (luxS- and luxN-) will luminesce in the presence of AI-2. Controls used in the experiment include <i>S. typhimurium</i> SS007, <i>V. harveyi</i> BB120 and <i> E. coli</i> DH5a. Overnight OD600 and luminescence (CPS) readings will be taken by a VICTOR (thank you Surette lab, especially Margot!). <br /><br /> |
| | | | |
| - | Preliminary testing was done on the bacterial transformation and genomic DNA isolation activities in the virtual lab. Additionally, I now have transparent wings which glow. Cool beans. | + | Preliminary testing was done on the bacterial transformation and genomic DNA isolation activities in the virtual lab. Additionally, I now have transparent wings which glow. |
| | </div> | | </div> |
| | </td> | | </td> |
| Line 381: |
Line 381: |
| | <br> | | <br> |
| | <div class="heading"> | | <div class="heading"> |
| - | Descriptive Title of What You're Doing
| + | Fundamental Displays |
| | </div> | | </div> |
| | <br> | | <br> |
| | <div class="desc"> | | <div class="desc"> |
| | </html> | | </html> |
| - | WIKI CODING HERE
| + | I recently remembered that I will have to start making cleaning scripts for a lot of the equipment in the lab to prevent clutter for easy maintenance. I started to make these and added them to bacterial transformation, the sequencing station and DNA extraction. It is not imperative that these be completed immediately since this type of abuse is not expected for testing. |
| | + | <br> |
| | + | <br> |
| | + | Added rho termination, but I am not satisfied with it yet and will continue to change where objects travel too although the pushing part looks satisfactory I need a better place for the mRNA strand. I have no idea how I am going to do the hairpin termination at this time but it will probably mean I have to fiddle with rotation again. There is now a complementary strand of DNA that will change its bases when an avatar makes changes to the template strand (the bottom one strand in this animation) so that the two strands will remain complementary to one another. |
| | + | <br> |
| | + | <br> |
| | + | Also did some more wiki updates. |
| | | | |
| | <html> | | <html> |
| Line 412: |
Line 418: |
| | <div class="desc"> | | <div class="desc"> |
| | </html> | | </html> |
| - | Will be updated soon
| + | Yesterday, varieties of bacterial cultures were grown in order to test the expression of GFP's. The fluorescence of GFP are measured by the plate reader, and following are the results that Emily and I were able to get today: |
| - | | + | *Fluorescent levels of different cultures (Stored in the plate reader for 5 hours after dilution of R0040+GFP) |
| | + | <br> |
| | + | [[Image:Calgary_GFP_reading_5hrs_Plate_reader.png|700px]] |
| | + | <br> |
| | + | *Fluorescent levels of different cultures (Stored in the shaker at 37˚C for 5 hours after dilution of R0040+GFP) |
| | + | <br> |
| | + | [[Image:Calgary_GFP_reading_5hrs_shaker.png|700px]] |
| | + | <br> |
| | + | We were forced to wait for 5 hours after the dilution of R0040+GFP culture as we were asked to leave the lab that had the plate reader by another organization. We were able to measure once quickly right after the dilution; however, with no accurate settings, meaning we cannot make any comparisons. |
| | + | As for the above results, it seems convincing enough for me to say those cells that have the reporter and mutant (LuxOD47E) circuits in them are doing something special that make their readings higher. Our KT1144 cells have a cosmid that contains <i>Qrr4</i> promoter driving an expression of GFP; thus, as shown by the data, express GFP in larger quantities when it has the LuxOD47E mutant than when it doesn't. Our TOP10 cells also have a plasmid containing <i>Pqrr4</i> and GFP, which behaves the same way as our KT1144 cells. |
| | + | Both data show that the fluorescence reading is about 2 times higher in cells that have both mutants and reporters than the cells that doesn't have mutant circuits. As we can see in well B7, the cells that have R0040, a constitutive promoter, driving GFP have a much higher reading than the rest, which is what was expected. Also as expected, diluting the R0040+GFP cells to 1/10, 1/50, and 1/100 ratios resulted in much lower readings of fluorescence. Two interesting things to note about the above collected data are: |
| | + | <br> |
| | + | 1. Both with and without mutant readings of TOP10 cells with reporter are than the readings of KT1144 cells. I do not know exactly why it is as such, but my guess is that different type of cells absorb different amount of wavelengths. |
| | + | <br> |
| | + | 2. The fluorescent levels of the cells that were grown for an extra 5 hours in the shaker were lower than the cells that grew in the plate reader, except for the cells that were diluted with fresh LB media. As above, I do not know the exact reason. Perhaps it is because the cells have already reached their 16~20 hours of growing overnight, growing for an extra 5 more hours at 37˚C killed the bacteria, decreasing the production of GFP, while the growth of the cells that were stored in the plate reader for an extra 5 hours slowed down due to a relatively colder temperature, thus being able to produce GFP more stably. |
| | + | As for the diluted cells with R0040+GFP, because they were transferred to a fresh media before the additional growth of 5 hours, storing them in the shaker at 37˚C led to faster replication, which sped up the production of GFP, while storing them in the colder plate reader slowed down the replication, which slowed the production of GFP. |
| | <html> | | <html> |
| | </div> | | </div> |
| | <br> | | <br> |
| | <div class="heading"> | | <div class="heading"> |
| - | 2. Sequencing | + | 2. Sequencing results |
| | </div> | | </div> |
| | <br> | | <br> |
| | <div class="desc"> | | <div class="desc"> |
| | </html> | | </html> |
| - | Will be updated soon
| + | The sequencing results of <i>Pqrr4</i>+B0034+K082003 came back today. Unfortunately, B0034 was still not there despite the change of <i>Pqrr4</i>+B0034 colonies from C9 to C1. Surprisingly, the sequence for the previous construction that used <i>Pqrr4</i>+B0034 C9 actually match the sequence that came back today, which used <i>Pqrr4</i>+B0034 C1. (99% match) The following is the alignment of the two sequences: |
| | | | |
| | + | >lcl|28877 |
| | + | Length=801 |
| | + | |
| | + | Score = 1472 bits (797), Expect = 0.0 |
| | + | Identities = 800/801 (99%), Gaps = 1/801 (0%) |
| | + | Strand=Plus/Plus |
| | + | |
| | + | Query 9 GGTGACACCTTGCCCTTTTTTGCCGGACTGCAGCGGCCGCTACTAGTATTAAGCTACTAA 68 |
| | + | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||| |
| | + | Sbjct 1 GGTGACACCTTGCCCTTTTTTGCCGGACTGCAGCGGCCGCTACTAGTATTAAGCTACTAA 60 |
| | + | |
| | + | Query 69 AGCGTAGTTTTCGTCGTTTGCAGCAGGCCTTTTGTATAGTTCATCCATGCCATGTGTAAT 128 |
| | + | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||| |
| | + | Sbjct 61 AGCGTAGTTTTCGTCGTTTGCAGCAGGCCTTTTGTATAGTTCATCCATGCCATGTGTAAT 120 |
| | + | |
| | + | Query 129 CCCAGCAGCTGTTACAAACTCAAGAAGGACCATGTGGTCTCTCTTTTCGTTGGGATCTTT 188 |
| | + | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||| |
| | + | Sbjct 121 CCCAGCAGCTGTTACAAACTCAAGAAGGACCATGTGGTCTCTCTTTTCGTTGGGATCTTT 180 |
| | + | |
| | + | Query 189 CGAAAGGGCAGATTGTGTGGACAGGTAATGGTTGTCTGGTAAAAGGACAGGGCCATCGCC 248 |
| | + | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||| |
| | + | Sbjct 181 CGAAAGGGCAGATTGTGTGGACAGGTAATGGTTGTCTGGTAAAAGGACAGGGCCATCGCC 240 |
| | + | |
| | + | Query 249 AATTGGAGTATTTTGTTGATAATGGTCTGCTAGTTGAACGCTTCCATCTTCAATGTTGTG 308 |
| | + | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||| |
| | + | Sbjct 241 AATTGGAGTATTTTGTTGATAATGGTCTGCTAGTTGAACGCTTCCATCTTCAATGTTGTG 300 |
| | + | |
| | + | Query 309 TCTAATTTTGAAGTTAACTTTGATTCCATTCTTTTGTTTGTCTGCCATGATGTATACATT 368 |
| | + | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||| |
| | + | Sbjct 301 TCTAATTTTGAAGTTAACTTTGATTCCATTCTTTTGTTTGTCTGCCATGATGTATACATT 360 |
| | + | |
| | + | Query 369 GTGTGAGTTATAGTTGTATTCCAATTTGTGTCCAAGAATGTTTCCATCTTCTTTAAAATC 428 |
| | + | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||| |
| | + | Sbjct 361 GTGTGAGTTATAGTTGTATTCCAATTTGTGTCCAAGAATGTTTCCATCTTCTTTAAAATC 420 |
| | + | |
| | + | Query 429 AATACCTTTTAACTCGATTCTATTAACAAGGGTATCACCTTCAAACTTGACTTCAGCACG 488 |
| | + | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||| |
| | + | Sbjct 421 AATACCTTTTAACTCGATTCTATTAACAAGGGTATCACCTTCAAACTTGACTTCAGCACG 480 |
| | + | |
| | + | Query 489 TGTCTTGTAGTTCCCGTCATCTTTGAAAAATATAGTTCTTTCCTGTACATAACCTTCGGG 548 |
| | + | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||| |
| | + | Sbjct 481 TGTCTTGTAGTTCCCGTCATCTTTGAAAAATATAGTTCTTTCCTGTACATAACCTTCGGG 540 |
| | + | |
| | + | Query 549 CATGGCACTCTTGAAAAAGTCATGCTGTTTCATATGATCTGGGTATCTCGCAAAGCATTG 608 |
| | + | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||| |
| | + | Sbjct 541 CATGGCACTCTTGAAAAAGTCATGCTGTTTCATATGATCTGGGTATCTCGCAAAGCATTG 600 |
| | + | |
| | + | Query 609 AACACCATAACCGAAAGTAGTGACAAGTGTTGGCCATGGAACAGGTAGTTTTCCAGTAGT 668 |
| | + | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||| |
| | + | Sbjct 601 AACACCATAACCGAAAGTAGTGACAAGTGTTGGCCATGGAACAGGTAGTTTTCCAGTAGT 660 |
| | + | |
| | + | Query 669 GCAAATAAATTTAAGGGTAAGTTTTCCGTATGTTGCATCACCTTCACCCTCTCCACTGAC 728 |
| | + | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||| |
| | + | Sbjct 661 GCAAATAAATTTAAGGGTAAGTTTTCCGTATGTTGCATCACCTTCACCCTCTCCACTGAC 720 |
| | + | |
| | + | Query 729 AGAAAA-TTTGTGCCCATTAACATCACCATCTAATTCAACAAGAATTGGGACAACTCCAG 787 |
| | + | |||||| ||||||||||||||||||||||||||||||||||||||||||||||||||||| |
| | + | Sbjct 721 AGAAAAATTTGTGCCCATTAACATCACCATCTAATTCAACAAGAATTGGGACAACTCCAG 780 |
| | + | |
| | + | Query 788 TGAAAAGTTCTTCTCCTTTAC 808 |
| | + | ||||||||||||||||||||| |
| | + | Sbjct 781 TGAAAAGTTCTTCTCCTTTAC 801 |
| | + | |
| | + | And the following is the previous sequencing result that I have gotten for <i>Pqrr4</i>+B0034 that was used for construction: |
| | + | |
| | + | <b>Sequence of Pqrr4+B0034 (RBS) C1</b> |
| | + | <Br> |
| | + | Legend |
| | + | <Br> |
| | + | <font color="red">Vector contamination</font> |
| | + | <br> |
| | + | <font color="green">BBK Restriction sites</font> |
| | + | <br> |
| | + | <font color="yellow"><i>Pqrr4</i> Sequence</font> |
| | + | <br> |
| | + | <font color="cyan">B0034 sequence</font> |
| | + | <br> |
| | + | |
| | + | <font color="red"> |
| | + | GGCGTATCACGAGGCAGAATTTCAGATAAAAAAAATCCTTAGCTTTCGCTAAGGATGATTTCTG</font> |
| | + | <font color="green"> |
| | + | GAATTCGCGGCCGCTTCTAGA</font><font color="yellow">GTATCAGCAAAAACACTACGGTGGATAATCAGTAAAACCATGAAACTAGAGCCCCGCA |
| | + | CACTTGCGGGGCTTTTTAATTTTGAATTTCTTTCTTATTAAAACGCCATTTTTCTGATAAATGTATTAGTAGCAA |
| | + | TGCGCATGGTGGCATATTTGCATCATTTTGCATTTTGCAAATGCGATTTGCAAAATGCGTGCTCAATAAAGCACC |
| | + | AATATGCATCAGGATCGAAGAAAAAAGGCGTTTTTAAAAGTTGGCACGCATCGTGCTTTATACAGAT</font><font color="purple">ACTAGAG</font><font color="cyan">AAAGAGGAGAAA</font> |
| | + | <font color="green">TACTAGTAGCGGCCGCTGCAG</font> |
| | + | |
| | + | <font color="red">TCCGGCAAAAAAGGGCAAGGTGTCACCACCCTGCCCTTTTTCTTTAAAACCGAAAAGATTACTT |
| | + | CGCGTTATGCAGGCTTCCTCGCTCACTGACTCGCTGCGCTCGGTCGTTCGGCTGCGGCGAGCGGTATCAGCTCACTCAAAGG |
| | + | CGGTAATACGGTTATCCACAGAATCAGGGGATAACGCAGGAAAGAACATGTGAGCAAAAGGCCAGCAAAAGGCCAGGAACCG |
| | + | TAAAAAGGCCGCGTTGCTGGCGTTTTTCCACAGGCTCCGCCCCCCTGACGAGCATCACAAAAATCGACGCTCAAGTCAGAGG |
| | + | TGGCGAAACCCGACAGGACTATAAAGATACCAGGCGTTTCCCCCTGGAAGCTCCCTCGTGCGCTCTCCTGTTCCGACCCTGC |
| | + | CGCTTACCGGATACCTGTCCGCCTTTCTCCCTTCGGGAAGCGTGGCGCTTTCTCATAGCTCACGCTGTAGGTATCTCAGTTC |
| | + | GGTGTAGGTCGTTCGCTCCAAGCTGGGCTGTGTGCACGAACCCCCCGTTCAGCCCGACCGCTGCGCCTTATCCGGTAACTAT |
| | + | CGTCTTGAGTCCAACCCGGTAGACACGACTTATCGCCACTG</font> |
| | + | |
| | + | <br> |
| | + | When I carefully analyzed today's <i>Pqrr4</i>+B0034(C1)+K082003, I could find part of the B0034 embeded between <i>Pqrr4</i> and K082003. The following explains this further: |
| | + | <br> |
| | + | <br> |
| | + | <b>Legend</b> |
| | + | <br> |
| | + | <Font color="yellow">end of <i>Pqrr4</i> sequence</font> |
| | + | <br> |
| | + | <Font color="green">Biobrick restriction sites</font> |
| | + | <br> |
| | + | <Font color="cyan">Part of B0034 (Complete sequence is AAAGAGGAGAAA</font> |
| | + | <br> |
| | + | <Font color="white">Part of scar? (Complete sequence is ACTAGAG)</font> |
| | + | <br> |
| | + | <Font color="pink">Beginning of K082003</font> |
| | + | <br> |
| | + | |
| | + | <Font color="yellow">...AGAT</font><font color="green">ACTAGAG</font><font color="cyan">AAAGAGG</font><font color="white">CTAG</font><font color="pink">ATGCGTA...</font> |
| | + | |
| | + | |
| | + | <br> |
| | + | <br> |
| | + | A mistake of sending two wrong colonies of <i>Pqrr4</i>+B0034 for sequencing is highly unlikely, and also a mistake of using a wrong colony for construction two times is highly unlikely, especially because I was not willing to mess up the construction the second time. I will have to investigate this further. |
| | <html> | | <html> |
| | </div> | | </div> |
| Line 534: |
Line 674: |
| | <div class="desc"> | | <div class="desc"> |
| | </html> | | </html> |
| | + | I ran the gene-specifc PCR with aiiA-F/R primers and ran it with a 1% gel at 100V. |
| | + | <br> |
| | + | <center> |
| | + | [[Image:Calgary_2009.08.19.gene.specific.aiiapcr.png|500px]] |
| | + | </center> |
| | + | <br> |
| | + | I had contaminated my negative control (which was just the mastermix) with my aiiA gene. Since all three bands are around 756 bp, we can say that the gene exists. |
| | + | |
| | I followed up with couple pharmaceutical companies. Some of our potential sponsors are currently deciding on sponsoring iGEM Calgary. Again, I couldn't get hold of other companies by phone but I left a message with a call back number. I also emailed those companies with a follow- up email. The good news is that Beckman Coulter has generously donated $200.00 to our iGEM Team this year. Unfortuantely, Alda Phamaceuticals Corporation is unable to sponsor our team this year. | | I followed up with couple pharmaceutical companies. Some of our potential sponsors are currently deciding on sponsoring iGEM Calgary. Again, I couldn't get hold of other companies by phone but I left a message with a call back number. I also emailed those companies with a follow- up email. The good news is that Beckman Coulter has generously donated $200.00 to our iGEM Team this year. Unfortuantely, Alda Phamaceuticals Corporation is unable to sponsor our team this year. |
| | | | |
| Line 559: |
Line 707: |
| | <br> | | <br> |
| | <div class="heading"> | | <div class="heading"> |
| - | Descriptive Title of What You're Doing
| + | Testing...Testing |
| | </div> | | </div> |
| | <br> | | <br> |
| | <div class="desc"> | | <div class="desc"> |
| | </html> | | </html> |
| - | WIKI CODING HERE
| + | Today, a generic avatar was created (aptly named Generic Lennie) that was in a separate group that had access to our island. He was used to test each area on our sim. We don't want visitors to be able to move, objects, change script and make new permanent objects. For the synthetic Kingdom everything was in order: |
| | + | <br> |
| | + | <br> |
| | + | *All objects were not modifiable |
| | + | *Generic Lennie was able to move the vitamins, Bad Guy/Champion cells, quorum sensing bacteria and oil clean-up bacteria, which is ok because they have a self-destruct script inside |
| | + | *GFP station drag and drop worked! |
| | + | <br> |
| | + | This means that the Synthetic Kingdom is unofficially done! |
| | | | |
| | <html> | | <html> |
| Line 585: |
Line 740: |
| | <br> | | <br> |
| | <div class="heading"> | | <div class="heading"> |
| - | Descriptive Title of What You're Doing
| + | ...And some more work on the modelling front |
| | </div> | | </div> |
| | <br> | | <br> |
| | <div class="desc"> | | <div class="desc"> |
| | </html> | | </html> |
| - | WIKI CODING HERE
| + | * Chinee and I met with Afshin and Iman to align our models with each other. The SimBiology and Membrane Computing reactions were subsequently updated, so that we are now modelling the same thing where applicable. The only major differences lie in that the membrane computing model accounts for LuxS (which the SimBiology model omits) and the membrane computing model accounts for AI-2 molecules leaving the cell and going to another one (while ours just includes a reverse reaction for AI-2 binding to LuxPQ, as we are only modelling performance in one cell). |
| | + | * I helped Chinee download Latex and will provide a lesson on how to use that platform |
| | + | * I looked up degradation rates for GFP:LVA mRNA |
| | + | * Our sensitivity analysis crashes with the addition of more reactions to our model. I worked on fixing that, to little avail. I will try again tomorrow. |
| | | | |
| | <html> | | <html> |
|
|
CAROL
Descriptive Title of What You’re Doing
WIKI CODING HERE
|
|
|
CHINMOYEE
Wiki Modelling page planning and Modelling meeting
MatLab paper formatting
Abstract
Introduction – talk about our system , have a system diagram
Methodology —algorithms , lab procedures etc
Results and discussions —graphs , pics of glow bac
Conclusions
(Do in LaTeX)
Wiki Formatting
Main frame
(put pics beside overview )
Left frame
Matlab modeling --- The tab contains definitions and overview of methodology like
Robustness
transfer function
response time
Parameter sensitivity
Optimization
Chemical kinetic equation / Law of mass action
more...
Membrane computing--- this tab contain pics and relavant information for overview of concepts
Paper 1 Matlab Modelling---
Paper 2 Membrane computing--- pdf file , made with LaTeX
The Meeting
Meeting with Afshin and Iman in main campus . Compared reactions present in both of our systems .
Need to inquire into mRNA degradation .
|
|
|
EMILY
Descriptive Title of What You're Doing
|
|
|
FAHD
Human Practices for August 20th 2009
Today, I continued to work on the ethics paper. I started reading literature on the environmental, legal and societal issues surrounding Quorum Sensing and I found a lot of interesting information that I can add on to the ethics paper. I will continue to work on the ethics paper over the next couple of days and try to improve it further.
I also contacted some companies that were on my marketing list. Most of these companies were Oil and gas development companies. I will do a follow-up with these companies next week.
|
|
|
IMAN
Descriptive Title of What You're Doing
|
|
|
JAMIE
Setting up the V. harveyi MM32 AI-2 Assay and Testing Experiments in Second Life's Virtual Lab
Using a 96 well microplate, AI-2 isolated from Salmonella typhimurium 14028 supernatant will be tested. V. harveyi reporter strain MM32 (luxS- and luxN-) will luminesce in the presence of AI-2. Controls used in the experiment include S. typhimurium SS007, V. harveyi BB120 and E. coli DH5a. Overnight OD600 and luminescence (CPS) readings will be taken by a VICTOR (thank you Surette lab, especially Margot!).
Preliminary testing was done on the bacterial transformation and genomic DNA isolation activities in the virtual lab. Additionally, I now have transparent wings which glow.
|
|
|
JEREMY
Descriptive Title of What You're Doing
|
|
|
KATIE
Fundamental Displays
I recently remembered that I will have to start making cleaning scripts for a lot of the equipment in the lab to prevent clutter for easy maintenance. I started to make these and added them to bacterial transformation, the sequencing station and DNA extraction. It is not imperative that these be completed immediately since this type of abuse is not expected for testing.
Added rho termination, but I am not satisfied with it yet and will continue to change where objects travel too although the pushing part looks satisfactory I need a better place for the mRNA strand. I have no idea how I am going to do the hairpin termination at this time but it will probably mean I have to fiddle with rotation again. There is now a complementary strand of DNA that will change its bases when an avatar makes changes to the template strand (the bottom one strand in this animation) so that the two strands will remain complementary to one another.
Also did some more wiki updates.
|
|
|
KEVIN
1. Measuring fluorescence of GFP
Yesterday, varieties of bacterial cultures were grown in order to test the expression of GFP's. The fluorescence of GFP are measured by the plate reader, and following are the results that Emily and I were able to get today:
- Fluorescent levels of different cultures (Stored in the plate reader for 5 hours after dilution of R0040+GFP)

- Fluorescent levels of different cultures (Stored in the shaker at 37˚C for 5 hours after dilution of R0040+GFP)
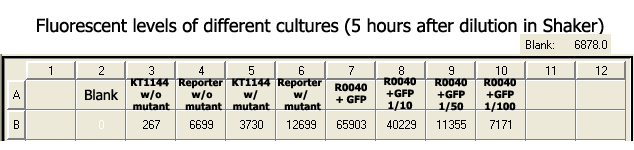
We were forced to wait for 5 hours after the dilution of R0040+GFP culture as we were asked to leave the lab that had the plate reader by another organization. We were able to measure once quickly right after the dilution; however, with no accurate settings, meaning we cannot make any comparisons.
As for the above results, it seems convincing enough for me to say those cells that have the reporter and mutant (LuxOD47E) circuits in them are doing something special that make their readings higher. Our KT1144 cells have a cosmid that contains Qrr4 promoter driving an expression of GFP; thus, as shown by the data, express GFP in larger quantities when it has the LuxOD47E mutant than when it doesn't. Our TOP10 cells also have a plasmid containing Pqrr4 and GFP, which behaves the same way as our KT1144 cells.
Both data show that the fluorescence reading is about 2 times higher in cells that have both mutants and reporters than the cells that doesn't have mutant circuits. As we can see in well B7, the cells that have R0040, a constitutive promoter, driving GFP have a much higher reading than the rest, which is what was expected. Also as expected, diluting the R0040+GFP cells to 1/10, 1/50, and 1/100 ratios resulted in much lower readings of fluorescence. Two interesting things to note about the above collected data are:
1. Both with and without mutant readings of TOP10 cells with reporter are than the readings of KT1144 cells. I do not know exactly why it is as such, but my guess is that different type of cells absorb different amount of wavelengths.
2. The fluorescent levels of the cells that were grown for an extra 5 hours in the shaker were lower than the cells that grew in the plate reader, except for the cells that were diluted with fresh LB media. As above, I do not know the exact reason. Perhaps it is because the cells have already reached their 16~20 hours of growing overnight, growing for an extra 5 more hours at 37˚C killed the bacteria, decreasing the production of GFP, while the growth of the cells that were stored in the plate reader for an extra 5 hours slowed down due to a relatively colder temperature, thus being able to produce GFP more stably.
As for the diluted cells with R0040+GFP, because they were transferred to a fresh media before the additional growth of 5 hours, storing them in the shaker at 37˚C led to faster replication, which sped up the production of GFP, while storing them in the colder plate reader slowed down the replication, which slowed the production of GFP.
2. Sequencing results
The sequencing results of Pqrr4+B0034+K082003 came back today. Unfortunately, B0034 was still not there despite the change of Pqrr4+B0034 colonies from C9 to C1. Surprisingly, the sequence for the previous construction that used Pqrr4+B0034 C9 actually match the sequence that came back today, which used Pqrr4+B0034 C1. (99% match) The following is the alignment of the two sequences:
>lcl|28877
Length=801
Score = 1472 bits (797), Expect = 0.0
Identities = 800/801 (99%), Gaps = 1/801 (0%)
Strand=Plus/Plus
Query 9 GGTGACACCTTGCCCTTTTTTGCCGGACTGCAGCGGCCGCTACTAGTATTAAGCTACTAA 68
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
Sbjct 1 GGTGACACCTTGCCCTTTTTTGCCGGACTGCAGCGGCCGCTACTAGTATTAAGCTACTAA 60
Query 69 AGCGTAGTTTTCGTCGTTTGCAGCAGGCCTTTTGTATAGTTCATCCATGCCATGTGTAAT 128
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
Sbjct 61 AGCGTAGTTTTCGTCGTTTGCAGCAGGCCTTTTGTATAGTTCATCCATGCCATGTGTAAT 120
Query 129 CCCAGCAGCTGTTACAAACTCAAGAAGGACCATGTGGTCTCTCTTTTCGTTGGGATCTTT 188
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
Sbjct 121 CCCAGCAGCTGTTACAAACTCAAGAAGGACCATGTGGTCTCTCTTTTCGTTGGGATCTTT 180
Query 189 CGAAAGGGCAGATTGTGTGGACAGGTAATGGTTGTCTGGTAAAAGGACAGGGCCATCGCC 248
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
Sbjct 181 CGAAAGGGCAGATTGTGTGGACAGGTAATGGTTGTCTGGTAAAAGGACAGGGCCATCGCC 240
Query 249 AATTGGAGTATTTTGTTGATAATGGTCTGCTAGTTGAACGCTTCCATCTTCAATGTTGTG 308
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
Sbjct 241 AATTGGAGTATTTTGTTGATAATGGTCTGCTAGTTGAACGCTTCCATCTTCAATGTTGTG 300
Query 309 TCTAATTTTGAAGTTAACTTTGATTCCATTCTTTTGTTTGTCTGCCATGATGTATACATT 368
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
Sbjct 301 TCTAATTTTGAAGTTAACTTTGATTCCATTCTTTTGTTTGTCTGCCATGATGTATACATT 360
Query 369 GTGTGAGTTATAGTTGTATTCCAATTTGTGTCCAAGAATGTTTCCATCTTCTTTAAAATC 428
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
Sbjct 361 GTGTGAGTTATAGTTGTATTCCAATTTGTGTCCAAGAATGTTTCCATCTTCTTTAAAATC 420
Query 429 AATACCTTTTAACTCGATTCTATTAACAAGGGTATCACCTTCAAACTTGACTTCAGCACG 488
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
Sbjct 421 AATACCTTTTAACTCGATTCTATTAACAAGGGTATCACCTTCAAACTTGACTTCAGCACG 480
Query 489 TGTCTTGTAGTTCCCGTCATCTTTGAAAAATATAGTTCTTTCCTGTACATAACCTTCGGG 548
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
Sbjct 481 TGTCTTGTAGTTCCCGTCATCTTTGAAAAATATAGTTCTTTCCTGTACATAACCTTCGGG 540
Query 549 CATGGCACTCTTGAAAAAGTCATGCTGTTTCATATGATCTGGGTATCTCGCAAAGCATTG 608
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
Sbjct 541 CATGGCACTCTTGAAAAAGTCATGCTGTTTCATATGATCTGGGTATCTCGCAAAGCATTG 600
Query 609 AACACCATAACCGAAAGTAGTGACAAGTGTTGGCCATGGAACAGGTAGTTTTCCAGTAGT 668
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
Sbjct 601 AACACCATAACCGAAAGTAGTGACAAGTGTTGGCCATGGAACAGGTAGTTTTCCAGTAGT 660
Query 669 GCAAATAAATTTAAGGGTAAGTTTTCCGTATGTTGCATCACCTTCACCCTCTCCACTGAC 728
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
Sbjct 661 GCAAATAAATTTAAGGGTAAGTTTTCCGTATGTTGCATCACCTTCACCCTCTCCACTGAC 720
Query 729 AGAAAA-TTTGTGCCCATTAACATCACCATCTAATTCAACAAGAATTGGGACAACTCCAG 787
|||||| |||||||||||||||||||||||||||||||||||||||||||||||||||||
Sbjct 721 AGAAAAATTTGTGCCCATTAACATCACCATCTAATTCAACAAGAATTGGGACAACTCCAG 780
Query 788 TGAAAAGTTCTTCTCCTTTAC 808
|||||||||||||||||||||
Sbjct 781 TGAAAAGTTCTTCTCCTTTAC 801
And the following is the previous sequencing result that I have gotten for Pqrr4+B0034 that was used for construction:
Sequence of Pqrr4+B0034 (RBS) C1
Legend
Vector contamination
BBK Restriction sites
Pqrr4 Sequence
B0034 sequence
GGCGTATCACGAGGCAGAATTTCAGATAAAAAAAATCCTTAGCTTTCGCTAAGGATGATTTCTG
GAATTCGCGGCCGCTTCTAGAGTATCAGCAAAAACACTACGGTGGATAATCAGTAAAACCATGAAACTAGAGCCCCGCA
CACTTGCGGGGCTTTTTAATTTTGAATTTCTTTCTTATTAAAACGCCATTTTTCTGATAAATGTATTAGTAGCAA
TGCGCATGGTGGCATATTTGCATCATTTTGCATTTTGCAAATGCGATTTGCAAAATGCGTGCTCAATAAAGCACC
AATATGCATCAGGATCGAAGAAAAAAGGCGTTTTTAAAAGTTGGCACGCATCGTGCTTTATACAGATACTAGAGAAAGAGGAGAAA
TACTAGTAGCGGCCGCTGCAG
TCCGGCAAAAAAGGGCAAGGTGTCACCACCCTGCCCTTTTTCTTTAAAACCGAAAAGATTACTT
CGCGTTATGCAGGCTTCCTCGCTCACTGACTCGCTGCGCTCGGTCGTTCGGCTGCGGCGAGCGGTATCAGCTCACTCAAAGG
CGGTAATACGGTTATCCACAGAATCAGGGGATAACGCAGGAAAGAACATGTGAGCAAAAGGCCAGCAAAAGGCCAGGAACCG
TAAAAAGGCCGCGTTGCTGGCGTTTTTCCACAGGCTCCGCCCCCCTGACGAGCATCACAAAAATCGACGCTCAAGTCAGAGG
TGGCGAAACCCGACAGGACTATAAAGATACCAGGCGTTTCCCCCTGGAAGCTCCCTCGTGCGCTCTCCTGTTCCGACCCTGC
CGCTTACCGGATACCTGTCCGCCTTTCTCCCTTCGGGAAGCGTGGCGCTTTCTCATAGCTCACGCTGTAGGTATCTCAGTTC
GGTGTAGGTCGTTCGCTCCAAGCTGGGCTGTGTGCACGAACCCCCCGTTCAGCCCGACCGCTGCGCCTTATCCGGTAACTAT
CGTCTTGAGTCCAACCCGGTAGACACGACTTATCGCCACTG
When I carefully analyzed today's Pqrr4+B0034(C1)+K082003, I could find part of the B0034 embeded between Pqrr4 and K082003. The following explains this further:
Legend
end of Pqrr4 sequence
Biobrick restriction sites
Part of B0034 (Complete sequence is AAAGAGGAGAAA
Part of scar? (Complete sequence is ACTAGAG)
Beginning of K082003
...AGATACTAGAGAAAGAGGCTAGATGCGTA...
A mistake of sending two wrong colonies of Pqrr4+B0034 for sequencing is highly unlikely, and also a mistake of using a wrong colony for construction two times is highly unlikely, especially because I was not willing to mess up the construction the second time. I will have to investigate this further.
|
|
|
MANDY
Descriptive Title of What You're Doing
|
|
|
PATRICK
The Next Big Thing: Levels
As the iGEM season starts winding up, it is becoming very urgent to make the second life project publicly presentable! I spent today getting organized and nailing down the design for the last big feature that will live in the Biobrick Simulator heads up display object: levels.
The simulator is not a game (though it will be fun), but the 'playing through levels' concept is widely known amongst young people these days, and it's much more appealing than something like 'concepts', or 'chapters', even though those terms are better descriptions of what a 'level' will be. Each level is a self contained set of parts, with the goal of introducing one or two new features of the simulator and concepts in molecular biology to the user. The preliminary list of levels is a good illustration of the progression I'm looking for:
- First Transcription
- Activators (a simple activatable device, and an activator)
- Repressors (a simple repressable device)
- External Activator (the activator/repressor is produced by a device, instead of given)
- External Repressor
- Negative Autoregulation (Control)
- Positive Autoregulation (Memory)
- Or Gates
- Inverters (Not Gate)
- And Gates (two inverters w. dual repressed promoter)
- Bistable Toggle Switch
- Tristable Toggle Switch (See [http://openwetware.org/wiki/IGEM:Brown/2007/Tri-Stable_Switch Brown's 2007 project.])
- Repressilator
Unfortunately, there are also a good number of systems I had wanted to make into levels, but whose features aren't yet supported by the simulator's engine! Including, but not limited to:
- Lac Operon
- Tet Operon
- And Gates (additional varieties)
- Lux Operon / Quorum Sensing
- Feedforward systems (various types)
- XOR Gates
- Ara Operon?
In addition, a number of the levels I will be building would benefit from features these other systems require, especially inducers for LacI and TetR. Inducers make a huge amount of sense for systems like the toggle switches, for getting from one state to another; and also for positive autoregulation, for turning on/off cellular memory. That's to say nothing of incorporating environmental signals like temperature, osmolarity, and light, each of which have sensor parts within the registry, leading to more systems and more levels... but I digress! In a perfect world I'd have a chance to implement all of these things before putting together a selection of levels. But since I would like a version of the simulator with most of its *features* in place before the end of summer, choices have to be made about content. And who knows, there might still be time.
Fortunately, none of the code that remains to be written for the levels is very complicated. The Level Selection screen makes the levels and their parts available, the Home screen gives some basic guidance for each level, and the Parts screen makes supplementary objects, and replacement objects in case you happen to delete the ones you're working with (or wish to combine multiple levels' parts together. That will work just fine!) All three screens are necessary for the levels to work, and none have new or complicated features, it's mostly a question of adapting code I've already written for other purposes.
And all of that isn't even the last major portion of the island that needs to be developed! To support all of this biobrickery, users will first visit a zone describing the basics of molecular biology and biochemistry (in Second Life's interactive style). Katie is already working on a display of DNA replication for this zone. Also fortunately, each of the exhibits I have in mind are more or less stand-alone displays, as compared to the biobrick simulator where every part interacts with half a dozen others, so I don't anticipate it taking as long.
But, with development as with biology, a good safe bet is to multiply all your time estimates by three...
|
|
|
PRIMA
Marketing and tieing up loose ends
I ran the gene-specifc PCR with aiiA-F/R primers and ran it with a 1% gel at 100V.
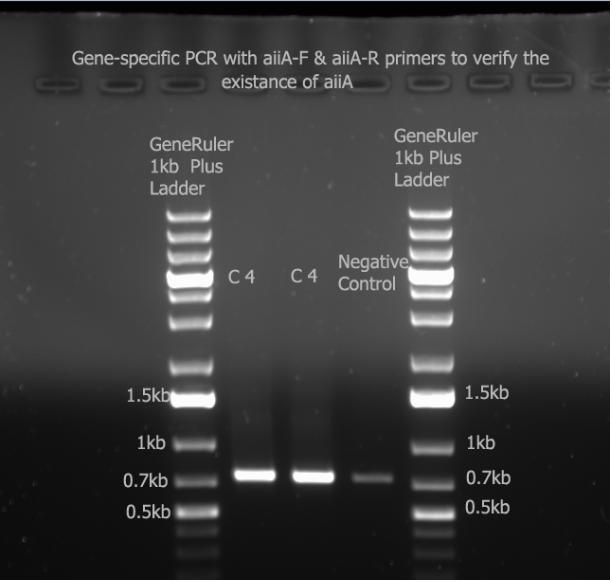
I had contaminated my negative control (which was just the mastermix) with my aiiA gene. Since all three bands are around 756 bp, we can say that the gene exists.
I followed up with couple pharmaceutical companies. Some of our potential sponsors are currently deciding on sponsoring iGEM Calgary. Again, I couldn't get hold of other companies by phone but I left a message with a call back number. I also emailed those companies with a follow- up email. The good news is that Beckman Coulter has generously donated $200.00 to our iGEM Team this year. Unfortuantely, Alda Phamaceuticals Corporation is unable to sponsor our team this year.
I updated some of the days on the wiki notebook. I also worked on my lab notebook to make sure all the gels were labeled and all pages are numbered.
Finally, I began working on the iGEM articles for media purposes and for our meeting with Leanne next week. I should have that completed by this weekend. Tomorrow, I'll discuss my gel with Thane and Sonja and decide where to go from here in the lab. For marketing, I'll continue to contact companies (mostly via email) for sponsorship for this year and next year.
|
|
|
STEFAN
Testing...Testing
Today, a generic avatar was created (aptly named Generic Lennie) that was in a separate group that had access to our island. He was used to test each area on our sim. We don't want visitors to be able to move, objects, change script and make new permanent objects. For the synthetic Kingdom everything was in order:
- All objects were not modifiable
- Generic Lennie was able to move the vitamins, Bad Guy/Champion cells, quorum sensing bacteria and oil clean-up bacteria, which is ok because they have a self-destruct script inside
- GFP station drag and drop worked!
This means that the Synthetic Kingdom is unofficially done!
|
|
|
VICKI
...And some more work on the modelling front
- Chinee and I met with Afshin and Iman to align our models with each other. The SimBiology and Membrane Computing reactions were subsequently updated, so that we are now modelling the same thing where applicable. The only major differences lie in that the membrane computing model accounts for LuxS (which the SimBiology model omits) and the membrane computing model accounts for AI-2 molecules leaving the cell and going to another one (while ours just includes a reverse reaction for AI-2 binding to LuxPQ, as we are only modelling performance in one cell).
- I helped Chinee download Latex and will provide a lesson on how to use that platform
- I looked up degradation rates for GFP:LVA mRNA
- Our sensitivity analysis crashes with the addition of more reactions to our model. I worked on fixing that, to little avail. I will try again tomorrow.
|

 "
"













