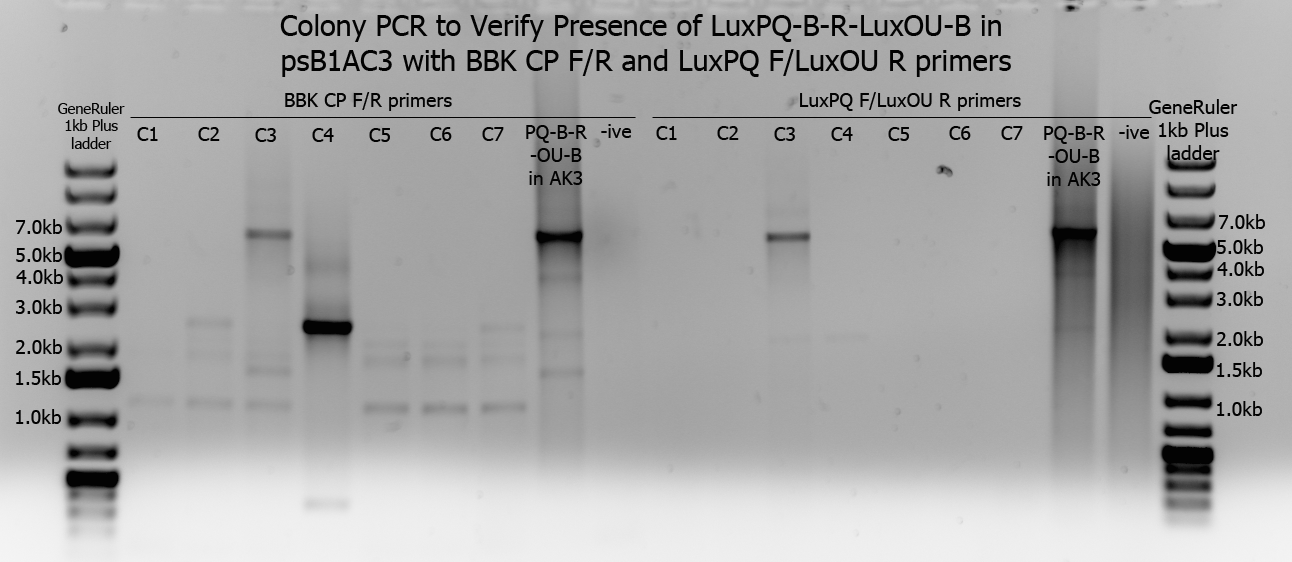Team:Calgary/30 July 2009
From 2009.igem.org
(Difference between revisions)
Prima.moinul (Talk | contribs) |
|||
| (37 intermediate revisions not shown) | |||
| Line 157: | Line 157: | ||
<br> | <br> | ||
<div class="heading"> | <div class="heading"> | ||
| - | + | Gradient PCR of R0040 with BamHI and XhoI sites | |
</div> | </div> | ||
<br> | <br> | ||
<div class="desc"> | <div class="desc"> | ||
</html> | </html> | ||
| - | + | * Received primers that will amplify R0040 with BamHI and XhoI sites on the ends from the University of Calgary synthesis lab. This will allow us to clone in the promoter for vector, pCS26, that can give us an observable luciferase expression. | |
| + | * Diluted primers and ran a gradient PCR (45 to 55 degrees Celsius) to amplify R0040+enzyme sites. | ||
| + | * Ran 4% gel at 60V to see if gradient PCR was successful or not. | ||
| + | <br> | ||
| + | <html> | ||
| + | <center> | ||
| + | <table> | ||
| + | <tr> | ||
| + | <td bgcolor="#ffffff" width="640px"> | ||
| + | </html> | ||
| + | [[Image:Calgary_BamHI+XhoI+R0040_Gradient_PCR.png|400px]] | ||
| + | <html> | ||
| + | </td> | ||
| + | </tr> | ||
| + | </table> | ||
| + | </center> | ||
| + | </html> | ||
| + | <br> | ||
| + | The gel shows that there is a faint band at ~75bp. This is approximately the size of R0040+BamHI+XhoI, so the promoter was PCR purified, then digested overnight at 37<sup>o</sup>C with XhoI and BamHI. | ||
| + | <br> | ||
| + | * The modelling team had a meeting with Dr. Nygren. We discussed about what we have done for modelling and we other parameters we would like to characterize. | ||
| + | |||
<html> | <html> | ||
</div> | </div> | ||
| Line 209: | Line 230: | ||
<br> | <br> | ||
<div class="heading"> | <div class="heading"> | ||
| - | + | Verification Digest of J13002-LuxOD47E-B0015 | |
</div> | </div> | ||
<br> | <br> | ||
<div class="desc"> | <div class="desc"> | ||
</html> | </html> | ||
| - | + | ||
| + | *Objective: Verify once more that B0015 has been cloned into the J13002-LuxOD47E construct before we send a colony down for sequencing. The previous gel looked a little strange and we saw some unexpected bands in the uncut lanes, so to be safe, we shall run the restriction digest again before sending anything down for sequencing. | ||
| + | <Br> | ||
| + | *Protocol: Cut with XbaI and PstI according to contruction technique protocol | ||
| + | <Br> | ||
| + | Visualized on a 1% agarose gel with uncut plasmid of the same colonies as a control. Also ran with the B0015-J13002-LuxOD47E construct as a size control. Bands of approximately 1.6 kb and 3 kb were expected represneting the construct and the psB1AC3 vector respectively. | ||
| + | <Br> | ||
| + | [[Image:Calgary_2009.07.30.J13002_LUXOD47E_B0015.RD_2.png|400px]] | ||
| + | <Br> | ||
| + | *Results: Lane 1, 3 and 5 are J13002-LuxOD7E-B0015 colonies 1, 2 and 3 digested with XbaI and PstI. | ||
| + | Lanes 2, 4 and 6 are colonies 1, 3 and 5 uncut. Lane 7 is B0015-J13002-LuxOD47E. The bands are the expected sizes which is good. We will send colony 3 down for sequecing tomorrow as the band for the construct looks the higest, so it has the greatest chance of having the terminator in it. | ||
| + | <Br> | ||
| + | |||
| + | Outreach | ||
| + | <Br> | ||
| + | <Br> | ||
| + | Contacted Rob McConeghy via E-Mail, about collaborating on an outreach initiative with the team. | ||
| + | <Br> | ||
| + | Attended a seminar on interdisciplinary learning in Second Life to get more familiar with Second Life conferences and get some ideas for Ethics. | ||
| + | |||
<html> | <html> | ||
| Line 235: | Line 275: | ||
<br> | <br> | ||
<div class="heading"> | <div class="heading"> | ||
| - | + | Outreach, Ethics and Marketing | |
</div> | </div> | ||
<br> | <br> | ||
<div class="desc"> | <div class="desc"> | ||
</html> | </html> | ||
| - | + | I started of my day by exploring and getting familiar with the online virtual world called Second Life (TM). I attended a conference on the Science Centre Island. I browsed different ways for organizing the Ethics conference in Second Life. | |
| + | <br> | ||
| + | <br> | ||
| + | For marketing, I did research on some more Oil & Gas companies starting off with Enbridge Pipelines and finishing off with Pajak Engineering. I e-mailed out newsletters to companies, updated our iGEM marketing list, filled out a grant application and printed posters for our Third University of Calgary iGEM Bake Sale. | ||
| + | <br> | ||
| + | <br> | ||
| + | I also looked into potential organizations that would help us in our 2009 iGEM Education and Outreach Campaign. I have decided to contact the respective individuals as early as next week. | ||
| + | |||
<html> | <html> | ||
| Line 287: | Line 334: | ||
<br> | <br> | ||
<div class="heading"> | <div class="heading"> | ||
| - | + | Paper writing and T-shirt brainstorming | |
</div> | </div> | ||
<br> | <br> | ||
<div class="desc"> | <div class="desc"> | ||
</html> | </html> | ||
| - | + | First draft of the paper is complete! | |
| + | We also came up with some ideas for the Tshirt. | ||
<html> | <html> | ||
| Line 313: | Line 361: | ||
<br> | <br> | ||
<div class="heading"> | <div class="heading"> | ||
| - | + | Verification of LuxPQ-LuxOU construct and cl lambda inverter | |
</div> | </div> | ||
<br> | <br> | ||
<div class="desc"> | <div class="desc"> | ||
</html> | </html> | ||
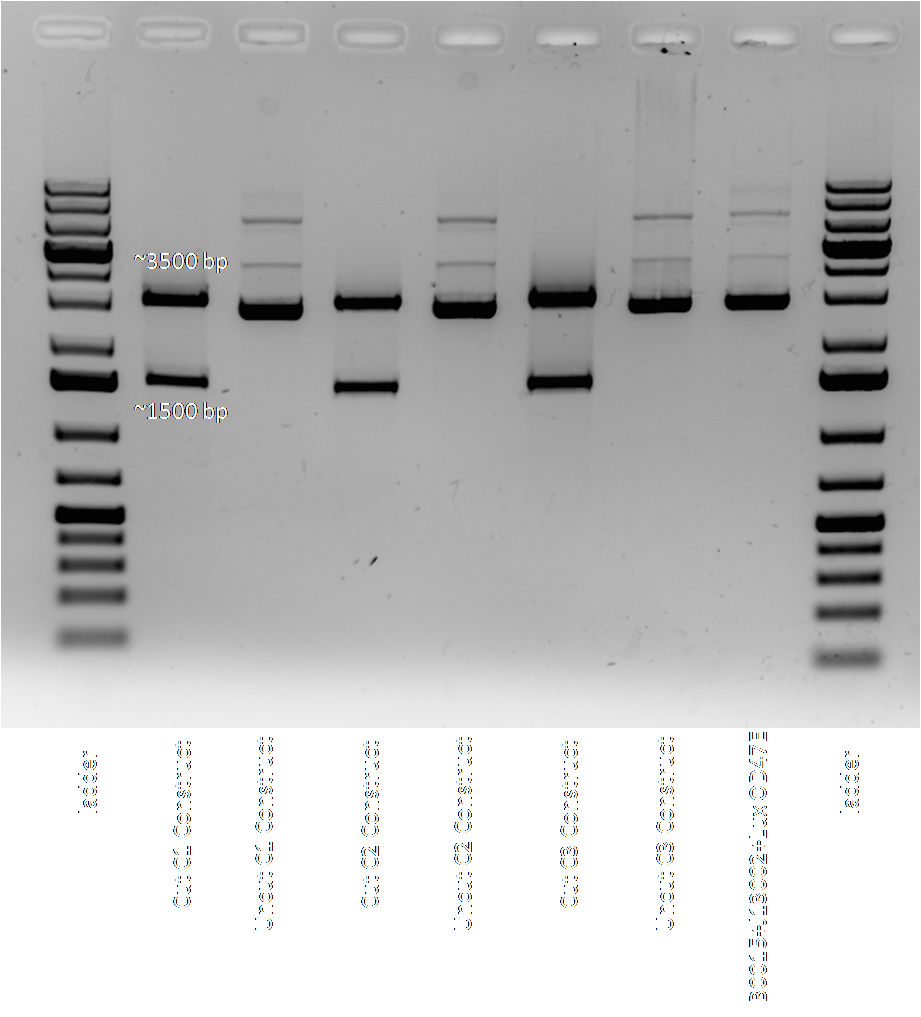
| - | + | A 0.7% agarose gel was run on the colony PCR of LuxPQ-B0015-R0040-LuxOU-B0015 in psB1AC3 set up on July 29. The following figure shows the results. This construct is ~6.0kb, and we see the right sized band for colony 3 for both sets of primers (BBK CP and gene specific.) Note that the positive control used is the construct in the AK3 plasmid. | |
| + | <br> | ||
| + | [[Image:2009.07.30.PQ_B_R_OU_B_AC3_construction.png|700px]] | ||
| + | <br> | ||
| + | Plasmid from colony 3 was then isolated using the QIAprep Spin MiniPrep Kit. Simultaneously, plasmid from cl lambda was also isolated. | ||
| + | <br> | ||
| + | <br> | ||
| + | A 2 hour restriction digest at 37ºC was set up for these isolated plasmids with ~500ng of DNA, XbaI and PstI in React 2 buffer (Invitrogen). This was then run on a 0.7% agarose gel and the PQ-OU construct appeared to be there, whereas the plasmid from cl lambda gave a faint band with no band at 1kb (the size of the gene). With this in mind, overnight liquid cultures were made with the intention of isolated plasmid again from cl lamda and then pooling the tubes together and vacufuging to increase the concentration of this piece. | ||
| + | |||
<html> | <html> | ||
| Line 344: | Line 400: | ||
<div class="desc"> | <div class="desc"> | ||
</html> | </html> | ||
| - | + | Phosphatase treatment is now being combined with the restriction digest activity so I moved some of the script that was within the recipient tube for restriction digest, into the phosphatase treatment script and had to add new sections to the recipient tube so that the product you receive has to be taken to phosphatase treatment manually, before an avatar may receive the proper product to begin construction. This required the new method of searching for inventory, which is much more effective. | |
| + | <br> | ||
| + | <br> | ||
| + | I completed instructions for the bacterial transformation activity and it was discovered that some of the items our avatars were using for PCR were named incorrectly, which was causing some major issues with testing out the equipment so those items have been renamed and the machine is in the process of being test once again. | ||
| + | <br> | ||
| + | <br> | ||
| + | <html> | ||
| + | </div> | ||
| + | <div class="heading"> | ||
| + | Communication between Objects | ||
| + | </div> | ||
| + | <br> | ||
| + | <div class="desc"> | ||
| + | </html> | ||
| + | I was also able to test the DNA extraction activity and it functions correctly as long as you follow the instructions provided throughout the activity. To improve upon it, we will have to get the objects to communicate with each other, which I have now started and the constant messaging would have to be reset for all steps within the product tube. | ||
| + | <br> | ||
| + | <br> | ||
| + | Since I find that the restriction digest is not unique compared to the other activity, tomorrow I believe that instead of moving the tube form water bath to heating block automatically, I will: | ||
| + | <br> | ||
| + | <br> | ||
| + | * Add to the listen event in restriction digest so that it may wait for prompts from the water bath and the heating block | ||
| + | * When the avatar is asked what the tube needs to be sent to they may touch the equipment and it will send a message of conformation to the tube, which was inspired by the touch capabilities of the DNA extraction activity have lead me to consider | ||
| + | * It will move to the equipment that was touched only if it was the correct place for the tube to be sent | ||
| + | * The listen will have to be removed after to prevent tube from being sent to equipment that is touched at random out of curiosity | ||
<html> | <html> | ||
| Line 365: | Line 444: | ||
<br> | <br> | ||
<div class="heading"> | <div class="heading"> | ||
| - | + | Verification of the transformed reporter circuit | |
</div> | </div> | ||
<br> | <br> | ||
<div class="desc"> | <div class="desc"> | ||
</html> | </html> | ||
| - | + | Now that hopefully the whole reporter circuit is transformed into TOP10 cells, I have performed a colony PCR of those colonies with Biobrick CP F/R primers and with the following conditions: | |
| + | 94°C for 3 min; 36x (94°C for 30s; 53°C for 45s; 72°C for 1 min and 20s;) 72°C for 10 min; held at 4°C. | ||
| + | <br><br> | ||
| + | Using Biobrick CP primers was a mistake, however, because I should have used <i>Pqrr4</i> F primer and Biobrick CP reverse primer. This would have allowed me to verify whether or not <i>Pqrr4</i> was not cut out during the construction. Another mistake was noticed, as I did not load any positive size control. Without any positive size controls to compare against, it is hard for me to tell whether or not it contains every piece of the circuit. Because of today's mistakes, I am planning to run another cPCR of the reporter circuit with 2 positive size controls, <i>Pqrr4</i> by itself and <i>Pqrr4</i> with B0034, and use <i>Pqrr4</i> Forward and biobrick CP Reverse primers. | ||
| + | <br><br> | ||
| + | The cPCR products were ran on 2% agarose gel, at 90 volts, and the following image is the picture of the gel. | ||
| + | <BR><BR> | ||
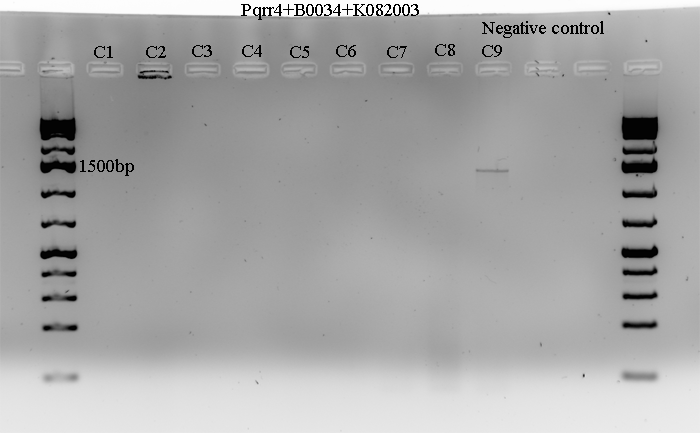
| + | Figure 1. cPCR of <i>Pqrr4</i>+B0034(RBS)+K082003(GFP+LVA)<br> | ||
| + | [[Image:Calgary_2009.07.30.Reporter_circuit.png]] | ||
| + | <br> | ||
| + | <br> | ||
| + | Colonies 1 to 8 did not get amplified at all, and Colony 9 seemed to have been amplified, but not of the right size. The expected size is about 1043bp with some additional length added due to the primers annealing outside, and the band in Colony 9 lane is a bit higher than the expected size. | ||
| + | <html> | ||
| + | <div class="heading"> | ||
| + | Modelling meeting | ||
| + | </div> | ||
| + | <br> | ||
| + | <div class="desc"> | ||
| + | </html> | ||
| + | Anders, one of our facilitators, came and gave us an advise on modelling. We discussed about limitations, advantages of our characterization methods and what we should focus on. | ||
| + | <br> | ||
| + | <Br> | ||
| + | <html> | ||
| + | <div class="heading"> | ||
| + | Updated Wiki notebooks from May 26 to June 11 | ||
| + | </div> | ||
| + | <br> | ||
| + | <div class="desc"> | ||
| + | </html> | ||
| + | Because our wiki notebook page has been created recently by Mandy, we were not able to post our previous updates, so old updates are now being done along with the recent updates. | ||
<html> | <html> | ||
| Line 391: | Line 499: | ||
<br> | <br> | ||
<div class="heading"> | <div class="heading"> | ||
| - | + | Second Life: Exploring Potentials of Seminars for Ethics | |
</div> | </div> | ||
<br> | <br> | ||
<div class="desc"> | <div class="desc"> | ||
</html> | </html> | ||
| - | + | Today, I took Emily and Fahd to an in-world seminar being held by Science Enthusiasts regarding the value of interdisciplinary learning, and its contrasts from 'multidisciplinary learning'. While the topic of the seminar was related to our project (as we participate in exactly that), the main purpose of our exploration was to monitor good (and unfortunately, bad) methods of running talks and seminars in Second Life. Thus, we could form a better idea of how we want to organize our proposed ethics seminar. | |
| - | + | <br> | |
| + | <br> | ||
| + | During the seminar, I was also able to meet an educator by the name of 'Graham Mill', who is exploring the uses of Second Life in teaching Microbiology. He has heard of iGEM and expressed interest in our project. Although he is occupied for the next few weeks, I will be meeting him again to tell him more about what we are doing and see what he has done to teach microbiology in SL. | ||
| + | <Br> | ||
| + | <br> | ||
<html> | <html> | ||
</div> | </div> | ||
| + | <div class="heading"> | ||
| + | Revision of Notecard Prompts for Second Life Lab Missions | ||
| + | </div> | ||
| + | <div class="desc"> | ||
| + | </html> | ||
| + | The three 'lab missions' that guide users through the lab section of our island are still in development. To make experiments more comparable in length (and to reinforce the concepts specifically related to bacterial transformation, an important procedure), I have added steps to two of the lab missions. | ||
| + | <br> | ||
| + | <br> | ||
| + | I have also been editting the notecards describing laboratory procedures, the lab missions ,and Katie's basics of microbiology (which she is awesome at). With the help of Max Chatnoir, the creator of Genome Island, and Thane, I have begun thinking of the best (and simplest) way to define a gene and its function (likely through an analogy - such as computers). Thane is really good at analogies, let me tell you. | ||
| + | <br> | ||
| + | <br> | ||
| + | Most of the time spent in second life were in the seminar and editting notecards. I've turned by DNA extraction activity over to Katie for fixing (as she is a master scripter and has figured out a way to solve my problem in which people can jump in at the end of the activity without doing anything at all. | ||
| + | <br> | ||
| + | <br> | ||
| + | <html> | ||
| + | </div> | ||
| + | <div class="heading">T-Shirt Design</div> | ||
| + | <div class="desc"> | ||
| + | </html> | ||
| + | Brainstormed some ideas for t-shirts with Jamie. These ideas involve a t-rex battling a giant squid. This will be a metaphorical representation of our team overcoming struggles with the quorum sensing system. Also, we are using that image because it is so awesome. | ||
| + | <html> | ||
| + | </div> | ||
| + | <br> | ||
</td> | </td> | ||
</tr> | </tr> | ||
| Line 443: | Line 578: | ||
<br> | <br> | ||
<div class="heading"> | <div class="heading"> | ||
| - | + | Colony PCR of LuxPQ-B-R-OU-B and marketing continued | |
</div> | </div> | ||
<br> | <br> | ||
<div class="desc"> | <div class="desc"> | ||
</html> | </html> | ||
| - | + | Today: | |
| + | '''Lab:''' | ||
| + | (shadowed Jeremy) I helped Jeremy in the lab again. We did a colony PCR to verify which colonies contained the LuxPQ-B-R-OU-B. The positive control was the psPAK3 plasmid. When we ran it on the gel, we had Colonies 1 to 7 set up on wells 2 to 8 with ladders on each end. These were cut plasmids. Then we had the positive control and then the same colonies assembled in the same order but these were uncut. However, only colony 3 seemed to have worked out with the correct band size since it aligned with the positive control. | ||
| + | |||
| + | Then we ran a restriction digest of Cl lambda to verify the size of Cl lambda. The order in the gel were: ladder in the first well, the cut Cl lambda (new plasmid that we ordered recently) on the second well then the uncut Cl lambda (new one)in the third, the ladder again and finally the uncut Cl lambda (old -that we used before). After running it on the 0.7% agarose gel, only the old uncut Cl lambda bands showed up. Aside from the ladders we couldn’t’ see any thing else. But before the restriction digest, when he did the nano drop, he said that there was a decent concentration of plasmid but we’re not sure why the bands didn’t show up. | ||
| + | |||
| + | '''For marketing:''' | ||
| + | President of Beckman Coulter called me back and said that he spoke to (someone in the BHSc O’brian Centre – I forgot the name, I’ll give it to you tomorrow) about iGEM because he was in Calgary for a conference for a few hours. He’s interested in helping us but he wants me to meet up with his representative who’s coming down from Edmonton next week. He said he’ll email him on my behalf since he didn’t have his number. So I sent the president another email with the sponsorship pkg and a new email about our phone convo so that he could forward that to the representative. The president told me to follow up with him in a few more days. | ||
| + | |||
| + | Cayman Chemicals I have to follow up on Monday – more time needed for consideration. | ||
| + | |||
| + | Company 1 -emailed back and said they can’t help due to economic probs. | ||
| + | The Wordans T-shirt guy Sales Manager emailed me back. I asked him for an estimate of costs for about 35-40 shirts (pink and black and sponsors on the back with black and white = total 3 colors) by October and a student discount/ group quote. | ||
| + | |||
| + | I wrote up the marketing blog today and posted it. I also started filling out some of the dates on the wiki notebook. I also mailed out a few more july newsletters. I checked over a few emails for Fahd and edited the outreach email. I also finished the email I plan to send out to Ms. Ellechuk (my hs bio teacher) for a presentation proposition and I had fahd and Emily double check it. I’ll ask Jeremy and Jamie to check it too tomorrow. Fahd and I printed off the bake sale posters. If he didn’t post all of them up today, I’ll finish posting tomorrow. We made a bake sale event on facebook and invited all our friends to our bake sale event. We were thinking of asking Lisa to send out an email to everyone just as a reminder next week. | ||
| + | |||
<html> | <html> | ||
| Line 469: | Line 619: | ||
<br> | <br> | ||
<div class="heading"> | <div class="heading"> | ||
| - | + | Developing a path | |
</div> | </div> | ||
<br> | <br> | ||
<div class="desc"> | <div class="desc"> | ||
</html> | </html> | ||
| - | + | The Synthetic Kingdom is a large area with plenty of things to see. If it is a person's first time going in, everything might be a bit overwhelming. Thus, I've decided it is essential to have a path of some sort that can lead people through. The different stations along the way will be bacteria "exhibits" that show uses of synthetic biology. The path was done in such a way as to not be obtrusive and lighted platforms when you step on the will indicate a station. Most of it was built but the main exit towards the island is a little more fantastic in scope and still under construction. | |
| + | <br><br> | ||
| + | <center> | ||
| + | [[Image:Calgary Pathway 002.png|700px]] | ||
| + | </center> | ||
<html> | <html> | ||
| Line 495: | Line 649: | ||
<br> | <br> | ||
<div class="heading"> | <div class="heading"> | ||
| - | + | Technical Report writing for LuxO D47A | |
</div> | </div> | ||
| - | |||
<div class="desc"> | <div class="desc"> | ||
</html> | </html> | ||
| - | + | Today's update will not be as exciting as the last few days: it's amazing how much time goes into technical writing and condensing a few months of work into a few short sentences. I made a lot of progress on my discussion section after consulting with Thane on what content is appropriate to include. We have a nicely-referenced paragraph on where LuxO D47A fits into the greater scheme of our signalling and reporter circuits, and how it will be used in conjunction with LuxO D47E to test the latter. | |
| + | |||
| + | <div class="heading"> | ||
| + | Mathematical modelling meeting with Professor Nygren | ||
| + | </div> | ||
| + | <div class="desc"> | ||
| + | Our modelling team met with Thane and Prof Nygren to discuss our recent progress and later direction on the mathematical modelling front. We presented the aspects of the circuit that we plan to characterise, which we will reveal tomorrow in the handout that we have prepared over the last few days. I will spare you the gory details for now, but we're planning to explore static performance (including a colony range analysis and a curve fit to the Hill Equation), dynamic performance (time dependence of our system), response time (based on the dynamic performance) and robustness (in terms of varying the promoter strengths and the surrounding conditions). We will also explore other areas that might be worth testing for robustness, based on the literature and what we can predict from our model. | ||
| + | |||
| + | <div class="heading"> | ||
| + | Preparation of meeting handout | ||
| + | </div> | ||
| + | <div class="desc"> | ||
| + | We had prepared a short document outlining the areas of our circuit that we plan to characterise. I have cleaned it up a bit so that it is more suited for our lab meeting presentation tomorrow. As mentioned earlier, the contents will be revealed in detail tomorrow. | ||
<html> | <html> | ||
Latest revision as of 07:02, 21 August 2009
UNIVERSITY OF CALGARY

 "
"