Team:TorontoMaRSDiscovery/Bioinformatics
From 2009.igem.org
(→Overview:) |
|||
| Line 17: | Line 17: | ||
===Overview:=== | ===Overview:=== | ||
| - | Spatial segregation is widely believed to be a defining organizational feature of eukaryotic cells: proteins, nucleic acids and small molecules are contained within and often actively transported between the many membrane-bound, subcellular organelles such as mitochondria, lysosomes, Golgi apparatus, etc. However, it has recently been found that some bacteria conditionally express proteinaceous microcompartments. These polyhedral organelles are usually 100-150 nm in cross section [1] and consist of proteinaceous outer shells, reminiscent of viral capsids, surrounding a core of enzymes[2]. It is thought that microcompartments allow bacteria to sequester specific metabolic enzymes and their substrates to enhance enzymatic efficiency and protect cells from the toxic effects of certain intermediates. While several examples of these compartments have been reported (as reviewed below) the diversity of this phenomenon as well as the variety of enzymes involved have not been explored. | + | Spatial segregation is widely believed to be a defining organizational feature of eukaryotic cells: proteins, nucleic acids and small molecules are contained within and often actively transported between the many membrane-bound, subcellular organelles such as mitochondria, lysosomes, Golgi apparatus, etc. However, it has recently been found that some bacteria conditionally express proteinaceous microcompartments. These polyhedral organelles are usually 100-150 nm in cross section [1] and consist of proteinaceous outer shells, reminiscent of viral capsids, surrounding a core of enzymes[2]. It is thought that microcompartments allow bacteria to sequester specific metabolic enzymes and their substrates to enhance enzymatic efficiency and protect cells from the toxic effects of certain intermediates. While several examples of these compartments have been reported (as reviewed below) the diversity of this phenomenon as well as the variety of enzymes involved have not been fully explored. We used publicly available, sequence-based resources to extend what is known about the diversity of microcompartments and their associated enzymes. An understanding of the natural biology of microcompartments will allow us to explore a variety of alternative approaches for our engineered system as well as help us to identify potential downstream applications. |
| - | + | ||
===Classifications:=== | ===Classifications:=== | ||
Revision as of 18:44, 19 October 2009
| Home | The Team | The Project | Parts Submitted to the Registry | Modeling | Bioinformatics | Notebook |
|---|
Contents |
Background information
Microcompartments:
Overview:
Spatial segregation is widely believed to be a defining organizational feature of eukaryotic cells: proteins, nucleic acids and small molecules are contained within and often actively transported between the many membrane-bound, subcellular organelles such as mitochondria, lysosomes, Golgi apparatus, etc. However, it has recently been found that some bacteria conditionally express proteinaceous microcompartments. These polyhedral organelles are usually 100-150 nm in cross section [1] and consist of proteinaceous outer shells, reminiscent of viral capsids, surrounding a core of enzymes[2]. It is thought that microcompartments allow bacteria to sequester specific metabolic enzymes and their substrates to enhance enzymatic efficiency and protect cells from the toxic effects of certain intermediates. While several examples of these compartments have been reported (as reviewed below) the diversity of this phenomenon as well as the variety of enzymes involved have not been fully explored. We used publicly available, sequence-based resources to extend what is known about the diversity of microcompartments and their associated enzymes. An understanding of the natural biology of microcompartments will allow us to explore a variety of alternative approaches for our engineered system as well as help us to identify potential downstream applications.
Classifications:
So far, most of our knowledge on bacterial microcompartments has been derived from the three well-studied microcompartment systems in nature.
- Carboxysomes
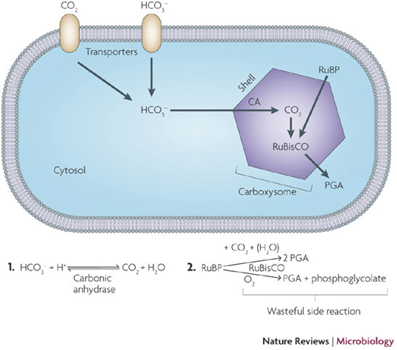
- First reported in 1956, carboxysomes were the first bacteria microcompartments to be discovered. They are often present in cyanobacteria and other chemoautotrophic bacteria [3]. They are known to play a key role in enhancing autotrophic carbon fixation in the Calvin cycle. The shell of the carboxysome encodes the enzymes carbonic anhydrase (CA) and ribulose bis-phosphate carboxylase monooxygenase (RuBisCO). CA converts bicarbonate ions into carbon dioxide, which is then converted into 3-phosphoglycerate (3-PGA) by RuBisCO. Carboxysome not only allows for co-localization of CA and RuBisCO, but also acts as a diffusion barrier to retain carbon dioxide in the immediate vicinity of RuBisCO and thus catalyzes the conversion [2].
- Pdu microcompartment
- In 1994, homologues of carboxysome shell proteins were reported in S. enterica. They are amongst a cluster of genes that are involved in coenzyme B12-dependent metabolism of 1,2 propanediol [4]. The gene cluster was later termed the pdu operon and the microcompartment formed was later termed propanediol utilization microcompartment. The proposed fuction of the pdu microcompartment is to encapsulate the enzymes that are necessary for cell to degrade propanediol and most importantly, to protect the cell from the toxic effects of propionaldehyde, an intermediate formed during the process [5].
- Eut microcompartment
- Later, similar structures were also found in E.coli and S. enterica when they were grown using ethanolamine as energy source. These structures were named ethanolamine utilization microcompartment (encoded by eut operon) and they often display very high genetic similarity with pdu microcompartment. Eut microcompartments contain enzymes involved in the degradation of ethanolamine and protect cell from acetaldehyde [6-8].
- Generally speaking, both pdu and eut microcompartments are less uniform in size and more irregular in shape than carboxysomes[2].
- Other putative microcompartment structures
- In addition to the three major systems mentioned above, several other putative microcompartments have been inferred from gene clustering analyses:
- A microcompartment was suggested to be involved in the oxidation of ethanol by Clostridium kluyveri. This was explored as an alternative platform to our current construction of encapsulin nanocomparment [9].
- A microcompartment associated with a puryvate-formate lyase homolog is proposed to be involved in the production of ethanol from pyruvate [10].
- A putative microcompartment in Rhodopirellula baltica is proposed to be associated with a lactate dehydrogenase homologue [1].
- A putative microcompartment in Carboxydothermus hydrogenoformans is associated with an isochorismatase-family protein [1].
- A putative microcompartment Solibacter usitatus can be associated with a dihydrdipicolinate synthase homologue [1].
Identification of conserved protein domain
Identification of new microcompartment is often achieved through bioinformatics analyses. (ie The pdu operon was first discovered as homologous to carboxysome genes.) The observation that diverse microcompartment structures are composed of proteins with homologous sequences led to the identification of a protein domain in the shell of all polyhedral bacterial microcompartments. This conserved protein domain, Pfam00936, is known as the Bacterial Microcompartment (BMC) domain. It is approximately 84 amino acids long and can be either found as a part of a large protein or in tandem copies within the same operon. This domain is found to be present in 189 bacterial species to date. [2]
Applications in nature
- Microcompartments are used to substantially catalyze a particular metabolic pathway by sequestering or co-localizing multiple metabolically related enzymes. This is true for all microcompartment structures discussed above because the product of one enzymes reaction will be in extreme vicinity to the next enzyme and will be delivered at high concentration as a substrate for the next enzyme. This prevents any potential diffusion loss of substrate and greatly enhances the enzymatic efficiency.
- Microcompartments are used to occlude toxic intermediates that cannot be degraded by normal bacterial machinery. This was shown to be true in pdu and eut microcompartments. The degradation pathway of 1,2 propanediol and ethanolamine both proceed through aldehyde intermediates (propionaldehyde and acetaldehyde respectively). The toxicity of these aldehyde intermediates was later shown in growth assays in which cells accumulating large amounts of propionaldehyde underwent growth arrest due to propionaldehyde toxicity [5, 11]. Therefore, it is reasonable to propose that bacteria are using microcompartment to mitigate the toxicity of aldehyde by encapsulating these poisonous intermediates.
Future Directions
Our ultimate goal is to exploit an understanding of bacterial microcompartments to design new microcompartments with modified properties or novel enzymatic activities, which could result in potentially useful applications in biotechnology. With those two objectives in mind, we attempted to address the following two questions using various bioinformatics tools: (1) What enzymes will benefit the most from enzymatic channeling? and (2) Is there any other alternative microcompartment chassis that could be explored for enzymatic channeling engineering?
1. What enzymes will benefit the most from enzymatic channeling?
We’d like to identify several enzyme pairs that display observable benefit from enzymatic channeling. Our approach to address this question is proposed based on the following assumptions:
- Enzymes pair with specific thermodynamic energetic properties will likely to benefit from enzymatic channeling. For two sequential reactions, the second one will be thermodynamically more favorable than the first reaction. Consequently, the products from the first reaction be consumed at a faster rate than being generated, hence driving the reaction to completion.
- Enzymes involved in reactions with toxic intermediates will likely to benefit from enzymatic channeling.
- Enzymes that were identified as gene fusion products in other organisms will likely to benefit from enzymatic channeling.
Based on these three premises, the following enzyme tables were constructed:
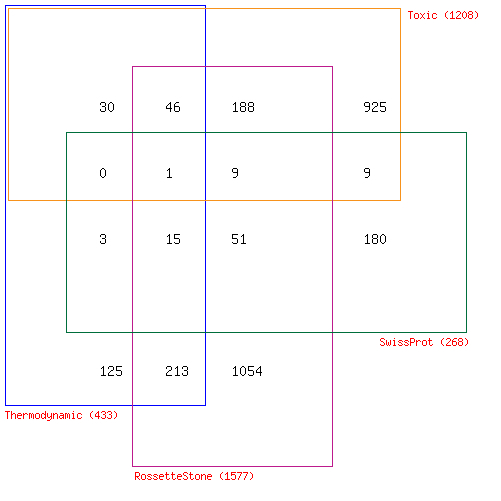
- Thermodynamic properties:
- Scored.csv: This is the list of all predictions based on free energy values.
- Toxic intermediates:
- ToxicPairs.xls: Enzyme pairs that possess a common metabolite that is known to be toxic.
- Gene fusion or multicomplex proteins:
- RosettaEnzymePairs.xls: The enzyme pairs predicted to form multifunctional enzymes by the Prolinks Rosetta Stone analysis.
- SwissProtPairs.xls: The enzyme pairs annotated to form multifunctional enzymes in SwissProt database.
By looking up their EC numbers in BRENDA database, the enzymes that are not immediate adjacent to each other in the pathway are eliminated. The enzymes with molecular weight well beyond microcompartment range (100 kDa) are also eliminated from the final table. The remaining enzyme pairs are considered as primary candidates for enzymatic channeling (Fig.2).
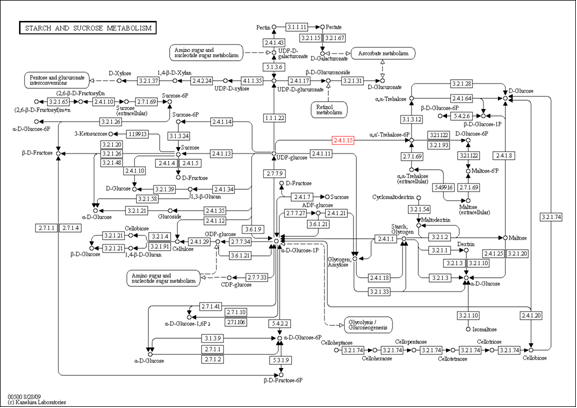
For example:
| EC 1 | Molecular Weight 1 | EC 2 | Molecular Weight 2 | Sources | Pathway Involved | Final Product |
| 2.4.1.15 | 45-630 | 3.1.3.12 | 25-973 | Rosette | Starch and sucrose metabolism | a,a trehalose |
2. What alternative microcompartment chassis could be explored?
Upon further examination, most of our known microcompartments tend to consist of a large number of protein subunits with very complex structures. Because of their complicated regulatory patterns and stoichiometry, these structures are not ideal to be used in enzymatic channeling engineering.
- Carboxysome: 80-150 nm in cross section. Composed of several thousands polypeptide of 10-15 different types. Contain as many as 250 RuBisCO per carboxysome molecule[12, 13].
- Pdu Microcomparment: 100-150 nm in cross section. Composed of about 18000 individual polypeptides of about 14-18 different types [1].
- Eut Microcomparment: Similar to pdu microcompartment in terms of both size and protein composition.
One of the putative microcompartment systems in Clostridium kluyveri associated with the oxidation of ethanol, however, shows great promises as an alternative microcompartment chassis because:
- The gene cluster involved contains only seven genes: including two genes for two almost identical acetaldehyde dehydrogenases, three genes for highly similar ethanol dehydrogenases, and two genes for microcompartment proteins, which are orthologs of ethanolamine using genes (eutML) of Salmonella typhimurium [9].
- From cell extracts of C. kluyveri a macromolecular complex of ethanol dehydrogenase and acetaldehyde dehydrogenase can be purified by differential manganese sulfate precipitation, indicating the presence of a functional and intact microcompartment enclosing these two enzymes[9].
Microcompartment protein gene sequences retrieved from NCBI database:
| CKL_1072 |
| 153953697|ref|YP_001394462.1| microcompartment protein [Clostridium kluyveri DSM 555] |
| MGQEALGMIETKGLVGAIEAADSMVKAANVALIGYEKIGSGLVTVMVRGDVGAVKAATDAGAASAKRVGE
VISVHVIPRPHTDVEKILPNIG |
| CKL_1073 |
| 153953698|ref|YP_001394463.1| microcompartment protein [Clostridium kluyveri DSM 555] |
| MNNELIEKVLGEVRKSLDLKNFDQEKLNKVVESTTEKLSDSKKEEAIKEAKPDVKVAEESKQAVVEQKAN
DVKTAPTMTEFVGTAGGDTVGLVIANVDSLLHKHLGLDNTCRSIGIISARVGAPAQMMAADEAVKGTNTE VATIELPRDTKGGAGHGIFIVLKAADVSDARRAVEIALKQTDKYLGNVYLCDAGHLEVQYTARASLIFEK AFGAPSGQAFGIMHAAPAGVGMIVADTALKTADVKLITYGSPTNGVLSYTNEILITISGDSGAVLQSLTA ARKAGLSILRSMGQDPVSMSKPTF |
Position Specific Iterative (PSI) BLAST was conducted on these two sequences respectively. PSI-BLAST constructs and modifies the Position Specific Scoring Matrix constantly and uses it during the next round of search. This iterative searching strategy results in increased sensitivity. The top 50 results were saved after 5 iterations. (please add linkout pages to the following documents)
CKL_1072 Hit Table CKL_1072 Taxonomy Report
CKL_1073 Hit Table CKL_1073 Taxonomy Report
Later, the top 50 hit genes and their adjacent genetic environment (10 genes 5’ upstream and 3’ downstream of the hit gene) were aligned using MUSCLE alignment. This is mainly used to identify any potential conserved sequences as targeting sequences to translocate enzymes into the microcompartment.
References
1.Cheng, S., et al., Bacterial microcompartments: their properties and paradoxes. Bioessays, 2008. 30(11-12): p. 1084-95.
2.Yeates, T.O., et al., Protein-based organelles in bacteria: carboxysomes and related microcompartments. Nat Rev Microbiol, 2008. 6(9): p. 681-91.
3.Shively, J.M., Inclusion bodies of prokaryotes. Annu Rev Microbiol, 1974. 28(0): p. 167-87.
4.Chen, P., D.I. Andersson, and J.R. Roth, The control region of the pdu/cob regulon in Salmonella typhimurium. J Bacteriol, 1994. 176(17): p. 5474-82.
5.Havemann, G.D., E.M. Sampson, and T.A. Bobik, PduA is a shell protein of polyhedral organelles involved in coenzyme B(12)-dependent degradation of 1,2-propanediol in Salmonella enterica serovar typhimurium LT2. J Bacteriol, 2002. 184(5): p. 1253-61.
6.Brinsmade, S.R., T. Paldon, and J.C. Escalante-Semerena, Minimal functions and physiological conditions required for growth of salmonella enterica on ethanolamine in the absence of the metabolosome. J Bacteriol, 2005. 187(23): p. 8039-46.
7.Penrod, J.T. and J.R. Roth, Conserving a volatile metabolite: a role for carboxysome-like organelles in Salmonella enterica. J Bacteriol, 2006. 188(8): p. 2865-74.
8.Stojiljkovic, I., A.J. Baumler, and F. Heffron, Ethanolamine utilization in Salmonella typhimurium: nucleotide sequence, protein expression, and mutational analysis of the cchA cchB eutE eutJ eutG eutH gene cluster. J Bacteriol, 1995. 177(5): p. 1357-66.
9.Seedorf, H., et al., The genome of Clostridium kluyveri, a strict anaerobe with unique metabolic features. Proc Natl Acad Sci U S A, 2008. 105(6): p. 2128-33.
10.Wackett, L.P., et al., Genomic and biochemical studies demonstrating the absence of an alkane-producing phenotype in Vibrio furnissii M1. Appl Environ Microbiol, 2007. 73(22): p. 7192-8.
11.Sampson, E.M. and T.A. Bobik, Microcompartments for B12-dependent 1,2-propanediol degradation provide protection from DNA and cellular damage by a reactive metabolic intermediate. J Bacteriol, 2008. 190(8): p. 2966-71.
12.Yeates, T.O., et al., Self-assembly in the carboxysome: a viral capsid-like protein shell in bacterial cells. Biochem Soc Trans, 2007. 35(Pt 3): p. 508-11.
13.Tanaka, S., et al., Atomic-level models of the bacterial carboxysome shell. Science, 2008. 319(5866): p. 1083-6.
 "
"