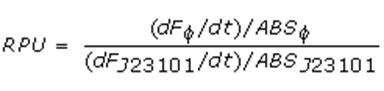Team:UNIPV-Pavia/Parts Characterization
From 2009.igem.org
(→Growth conditions) |
|||
| Line 124: | Line 124: | ||
*strain, plasmid copy number, antibiotic, growth medium, growth conditions, protein generator assembled downstream of the promoter must be the same in the promoter of interest and in J23101 reference standard. | *strain, plasmid copy number, antibiotic, growth medium, growth conditions, protein generator assembled downstream of the promoter must be the same in the promoter of interest and in J23101 reference standard. | ||
*steady state must be validated, so the considered RPU values are in exponential growth phase, when dF/dt/ABS (proportional to the GFP synthesis rate per cell) does not vary. | *steady state must be validated, so the considered RPU values are in exponential growth phase, when dF/dt/ABS (proportional to the GFP synthesis rate per cell) does not vary. | ||
| + | |||
| + | |||
| + | = Materials = | ||
| + | *Long term glycerol stocks were stored at -80°C with a final glycerol concentration of 20% | ||
| + | *Antibiotics were: Ampicillin (Amp) 100 ug/ml, Kanamycin (Kan) 50 ug/ml and Chloramphenicol (Cm) 12.5 ug/ml. All of them were stored at -20°C in 1000x stocks. Amp and Kan were dissolved in water, while Cm was dissolved in ethanol 100%. | ||
| + | *LB medium was prepare with: 1% NaCl, 1% bactotryptone, 0.5% yeast extract. The medium was not buffered with NaOH. | ||
| + | *M9 supplemented medium was prepared according to: [http://openwetware.org/wiki/Knight:M9_supplemented_media]. | ||
| + | *3OC6-HSL (Sigma) was dissolved in water and stored at -20°C in a 2mM stock. | ||
| + | *aTc (Clontech) was dissolved in ethanol 50% and stored at -20°C in a 100 ug/ml stock. All the following dilutions were performed in water. | ||
| + | *Ready made IPTG (Sigma) was stored at -20°C in a 200mM stock. | ||
</td> | </td> | ||
</tr> | </tr> | ||
</table> | </table> | ||
Revision as of 08:59, 19 October 2009

|
|
|
Parts Characterization |
|
|
Here we describe the characterization results of 4 parts of our own design, 2 existing parts re-built because they were inconsistent and 7 existing parts taken from the Registry. When not reported differently, all the experiments have been performed according to Growth conditions and Data analysis sections.
Re-built existing parts (BBa_our part code/BBa_existing part code): Existing parts from the Registry:
Our new partsBBa_K173003 - ethanol producing deviceBBa_K173007 - aTc inducible device with J23100 promoterBBa_K173011 - aTc inducible device with J23118 promoterBBa_K173010 - lactose/IPTG inducible device with J23118 promoterRe-built existing partsBBa_K173004/BBa_I732019 - beta-galactosidase protein generatorBBa_K173005/BBa_Q04400 - tetR QPIExisting parts from the RegistryBBa_J23100, BBa_J23101, BBa_J23118 - constitutive promoter family membersBBa_F2620 - 3OC6HSL receiver deviceBBa_K116001 - nhaA promoterBBa_K116001 from iGEM 2008 NYMU-Taipei We received this BioBrick from iGEM in September but the bacterial strain that contained the plasmid wasn't declared. So we decided to sequence it as check and transform it into E.coli TOP10. We wanted to perform some experiments to better understand how it works and if can be successfully used. Experiment #1MotivationIn our opinion the working principle of the antiporter Na+/H+ channel described in [1-2-3-4] makes the nhaA promoter a Na+ sensor and only under certain conditions (presence of Na+) a pH sensor. Methods
BBa_K112808 - Enterobacteria phage T4 Lysis DeviceBBa_R0011 - Plac hybrid promoterGrowth conditionsFor every culture we tested:
Data analysisGrowth curvesThe presented growth curves have all been processed as OD600_culture-OD600_broth for each time sample. OD600_broth is the medium in the same conditions as in the culture (e.g. induced with the same inducer concentration as in the culture). Relative Promoter Units (RPUs)The RPUs are standard units proposed by Kelly J. et al., 2008, in which the transcriptional strength of a promoter can be measured using a reference standard, just like the ground in electric circuits. RPUs have been computed as: in which:
RPU measurement has the following advantages:
The hypotheses on which RPU theory is based can be found in Kelly J. et al., 2008, as well as all the mathematical steps. From our point of view, the main hypotheses to satisfy are the following:
Materials
|
 "
"