Team:Cornell/Project/Standardization
From 2009.igem.org
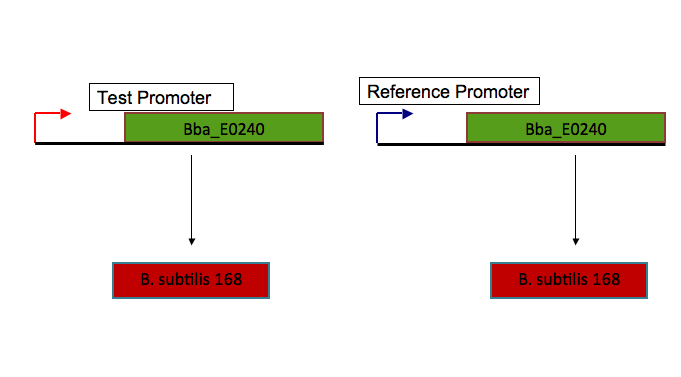
Standardization of partsOver the last six years, iGEM teams have created hundreds of BioBricks and submitted them to the Registry of Parts. These BioBricks adhere to standard assembly methods that have been developed by pioneering iGEM labs and are now widely used by teams everywhere. Although BioBricks are synthesized in the same manner, no standards yet exist that determine how they can be compared to each other. Of the hundreds of parts in the registry, only a few have been “characterized,” or studied in terms of how efficiently they function. Those that have been characterized often cannot be directly compared because their performance has been measured in different units, and they may work optimally in different organisms. Therefore, standardized measurement techniques and units of measurement are needed in order to directly compare similar BioBricks from the same chassis that have been made in different laboratory environments. Measurement standards will improve research on the design, construction, and use of gene parts [1]. A proposed unit of measurement for promoter activity is polymerases per second (PoPS), the rate at which RNA polymerase moves past a particular location in the DNA—a gene’s transcription rate. The PoPS value just downstream of a coding region for a gene should be equal to the rate at which a coding sequence is transcribed from DNA into messenger RNA (mRNA). Also, reporting activity in PoPS allows promoters to be directly compared to other parts that affect polymerase activity, such as transcription terminators [1]. Therefore, PoPs is a measurement of a promoter’s efficiency in transcribing the gene under study.[2] Currently, no methods for directly measuring PoPS in vivo have been documented. However, by inserting a promoter upstream of the coding sequence for green fluorescent protein (GFP), the rate of GFP synthesis, or GFP activity, can be used to measure promoter activity indirectly. This method can be used to synthesize plasmids containing either the standard reference promoter or the test promoter, the GFP reporter, and identical sequences following the transcription initiation site to ensure that both promoters transcribe identical mRNA products. After transforming the plasmids into identical cell strains and measuring their GFP fluorescence activity using the same instrument and technique, the values can be related using the following ratio: Relative activity of test promoter (in RPUs) = [A/R] where A represents the GFP activity of the test promoter, and R represents the GFP activity of the standard reference promoter.[1] The ratio measures the absolute GFP activity of the test promoter relative to that of the reference standard promoter—its relative promoter activity. It was derived from an ordinary differential equation for absolute promoter activity in terms of GFP production at steady state (Kelly). Kelly et. al defined relative promoter activity in terms of a new unit: Relative Promoter Units, or RPUs. Therefore, if the test promoter and the standard reference promoter have the same absolute GFP activity, the test promoter has a relative activity of 1 RPU. [1] Because E. coli promoters are not compatible in Bacillus subtilis, we propose to adapt the above method described by Kelly et. al to create a standardized measurement for characterizing B. subtilis promoters. The mrgA promoter from John Helmann’s HB1266 strain is a constitutive, sigma A promoter that will serve as our in vivo reference standard for promoter activity. [3] It will be used to generate relative promoter units (RPUs) for promoters from different B. subtilis strains to develop an intraspecies promoter characterization standard. GFP (part BBa_E0240 from the Bio Bricks kit) will be used as the reporter. Separate plasmids containing the test promoter and the standard reference promoter ligated to GFP will be transformed via double homologous recombination with integration vectors into Bacillus subtilis strain 168, which is sequenced on the SubtiList genomic database.[4] The mrgA gene is transcriptionally induced in growing cells by hydrogen peroxide and in postexponential phase when Mn(II) and iron levels are low. [5] MrgA protein is made when bacteria enter stationary phase in the presence of low levels of these metals, and its induction is inhibited by micromolar levels of Mn(II) or iron. [3] Additionally, its transcription in the presence of hydrogen peroxide has made it a known member of the B. subtilis peroxide regulon.[5] The mrgA promoter that we recommend for use as an in vivo reference standard for promoter activity was created by one of our faculty advisors, Dr. John Helmann, when he studied the effects of hydrogen peroxide stress on the Bacillus subtilis mrgA gene. This promoter was taken from the mutant strain HB1266, in which the first half of the inverted repeat (extending from bases -12 to -20) upstream of the -35 region has been deleted. This cis-acting mutation led to a thorough derepression of the mrgA gene.[5] The activity of the mrgA promoter from strain HB 1266 ranges from 500 in mid-logarithmic phase to 800 Miller units in stationary phase [6]. In comparison, the activity of the wild type strain ranges to 500 Miller units when introduced to 30uM of Zn(II) and grown to mid-logarithmic phase [6], and the cadA promoter ranges up to 65 Miller units when introduced to 0.5mM Zn(II). Additionally the cadA promoter has a response of ~35 Miller units in 5uM Cd(II).[7] By measuring the GFP activity of Bacillus subtilis promoters relative to the mrgA promoter, we can reduce variation in reported promoter activity due to differences in test conditions and measurement instruments. We will measure the GFP activity of our test promoter, the yvgW promoter, with the ratio mentioned above to characterize it in RPUs. References1. Kelly et al. “Measuring the Activity of BioBrick Promoters Using an In Vivo Reference Standard.” Journal of Biological Engineering. 20 March 2009. 2. “PoPS.” Open WetWare. 2009. http://openwetware.org/wiki/PoPS 3. Helmann, John D. “Metalloregulation in Bacillus subtilis: Isolation and Characterization of Two Genes Differentially Repressed by Metal Ions.” Journal of Bacteriology. September 1993. p. 5428 – 5437 4. Moszer I, Jones LM, Moreira S, Fabry C, Danchin A. “SubtiList: the reference database for the Bacillus subtilis genome.” Nucleic Acids Res. 2002;30:62–65. 5. Helmann, John D. “Coordinate Regulation of Bacillus subtilis peroxide stress genes by hydrogen peroxide and metal ions.” Biochemistry. August 1995. Vol. 92, pp. 8190 – 8194. 6. Helmann, John D. “Genetic and Physiological Responses of Bacillus subtilis to Metal Ion Stress.” Molecular Microbiology. May 9 2005. Vol. 57, issue 1. Pp. 27 – 40. 7. Helmann, John D. “Bacillus subtilis CPx-type ATPases Characterization of Cd An Co and Cu Efflux Systems.” BioMetals. April 2003. Vol 53, pp. 497 - 505 |
 "
"