Team:DTU Denmark/notebookbiobrick
From 2009.igem.org

| Home | The Team | The Project | Parts submitted | Modelling | Notebook |
|
Activities relating to our two sub-projects: - The Redoxilator - The USERTM assembly standard - Biobricks - Protocols |
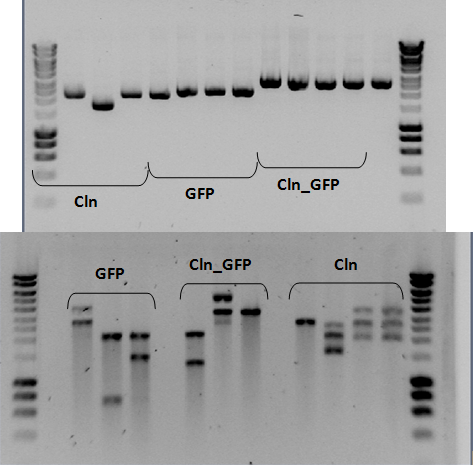
Day-to-day activities Activities relating to the biobricks For more detailed information see results. July 20th Construction of biobricks: The restriction products were evaluated on a 1% agarose gel (see figure 2).
Figure 2: Restriction analysis of the three biobricks. Construction of biobricks: The three plasmids were purified according to the protocol. Following the purification of the plasmids, a restriction analysis was performed using SpeI enzyme (10U) and incubated at 37ºC overnight. July 18th Construction of biobricks: Some colonies were picked from the transformation plates and made overnight cultures in liquid media. July 17th Construction of biobricks: Following purification, all of the PCR product was digested with XbaI (60U) and SpeI (30U). The mixtures were incubated at 37ºC. After 8 hours of incubation, more enzyme was added (XbaI (20 U) and SpeI (10U)) to ensure maximum digestion. Following enzymatic purification a gel was run to estimate the concentration of the vector and the PCR product (see figure 1).
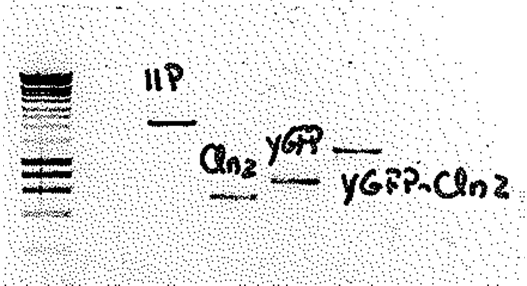
Figure 1: Electrophoresis gel picture of the PCR products of the three biobricks and the vector. The band 11P at 1500 bp corresponds to the vector; the three other bands at 537 bp, 207 bp and 125 bp correspond to the fragments Cln, GFP and Cln_GFP respectively. To meet the 3:1 ratio of fragment to vector 12μl fragment was mixed with 2μl vector, 2μl ligase buffer (10x), 1μl ligase, 3 μl Mq H20. The ligation of the fragment and the vector was carried out on PCR machine using a 3:1 ratio for 1.5 hours using the idea by Lund, A. H., M. Duch, and F. S. Pedersen. 1996. Increased cloning efficiency by temperature-cycle ligation. Nucleic Acids Res. 24:800-801. doi:l50400 [pii]. The final product was finally transformed into competent E coli cells. July 16th Construction of biobricks: The 357bp Cln2 fragment was amplified by PCR using the primers BGHA351/BGHA362 and Phusion polymerase. Primers BGHA368/369 were used to amplify the GFP fragment and finally, primers BGHA368/353 were used to amplify the fragment containing the GFP fused with the Cln. Following amplification the PCR products were run on a 1% agarose gel and purified The primers added tails to the PCR product containing an XbaI site in the forward primer and a SpeI in the reverse primer to meet the “Silver fusion” standard BBF RFC 23. July 15th Construction of biobricks: The plasmid was purified according to the protocol.
Following purification, the plasmid was digested using two enzymes, XbaI (20U) and SpeI (10U). The mixture was incubated at 37ºC. 8 hours after incubation, more enzyme was added to ensure maximum digestion as well as Antartic phosphatase (25U). After one hour the mixture was run on a 1% agarose gel and the 2kb backbone was purified. July 14th Construction of biobricks: Some colonies were picked from the transformation plate and made overnight cultures in liquid media. July 13th Construction of biobricks: We chose pSB1A2 from iGEM spring 2009 DNA collection Kit Plate (well 11P). The crystallized DNA was ressuspended and transformed into competent E. coli cells. |
Work process Our team works parallel in smaller sub-teams. Some of us work hard in the lab, while others are in the process of developing software and the in silico model of the Redoxilator system. However, we constantly keep each other updated, and meet often to exchange ideas and take turns at the different tasks, thus exhausting all of our combined knowledge in every aspect of this project. |
| Comments or questions to the team? Please Email us -- Comments of questions to webmaster? Please Email us |
 "
"