Team:Calgary/18 June 2009
From 2009.igem.org
(Difference between revisions)
Prima.moinul (Talk | contribs) |
|||
| (17 intermediate revisions not shown) | |||
| Line 137: | Line 137: | ||
<br> | <br> | ||
<div class="heading"> | <div class="heading"> | ||
| - | + | JUNE 18, 2009 | |
</div> | </div> | ||
<br> | <br> | ||
| Line 157: | Line 157: | ||
<br> | <br> | ||
<div class="heading"> | <div class="heading"> | ||
| - | + | Colony PCR | |
</div> | </div> | ||
<br> | <br> | ||
<div class="desc"> | <div class="desc"> | ||
</html> | </html> | ||
| - | + | * The transformation and the mutagenesis resulted in many colonies. Selected several colonies and continued with colony PCR with gene specific primers. | |
| - | + | * Results showed that the plasmid contained the 6KB sequence. | |
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
<html> | <html> | ||
| Line 235: | Line 184: | ||
<br> | <br> | ||
<div class="heading"> | <div class="heading"> | ||
| - | + | Campus Fair Follow-Up Meeting June 18th 2009 | |
</div> | </div> | ||
<br> | <br> | ||
<div class="desc"> | <div class="desc"> | ||
</html> | </html> | ||
| - | + | Today, I attended the our campus fair follow up meeting. Our sponsorship package also got rejected by Leica Inc. and Beckman Coulter. | |
<html> | <html> | ||
| Line 254: | Line 203: | ||
<tr> | <tr> | ||
<td> | <td> | ||
| - | <a name=" | + | <a name="Jamie"></a> |
<br> | <br> | ||
<div class="heading"> | <div class="heading"> | ||
| - | + | JAMIE | |
</div> | </div> | ||
<br> | <br> | ||
<div class="heading"> | <div class="heading"> | ||
| - | + | Biobricked LuxOU Restriction Digest | |
</div> | </div> | ||
<br> | <br> | ||
<div class="desc"> | <div class="desc"> | ||
</html> | </html> | ||
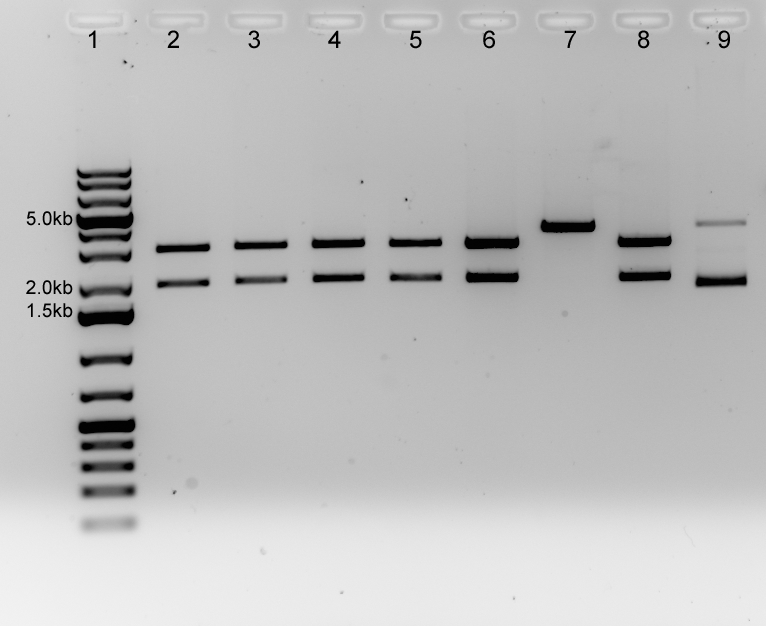
| - | + | Initial 1% agarose gel indicated unsuccesful cloning (data not shown). ''Not''I digest of other plasmids for 2h, 37oC and visualization with 1% agarose gel. | |
| - | + | [[Image:Calgary_luxOU_BBKVeri.jpg | 500px]] | |
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | Figure. 1% agarose gel (100V) for NotI verification digest of luxOU in psB1AC3 (XbaI and EcoRI construction). 200ng of plasmid was cut with NotI for 2 hours at 37oC. Lanes 2-5 are C1, C2, C4 and C5 respectively from the XbaI and PstI construction. Lanes 6-9 are C1-C4 respectively from the EcoRI and PstI construction. Expected band sizes were: 3kb (psB1AC3) and 2kb (luxOU) 5μL of GeneRuler 1kb Plus DNA Ladder was loaded. 20μL was loaded of the restriction digest reaction mixture with 4μL 10x Orange G to a total volume of 40μL. | |
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
<html> | <html> | ||
| Line 313: | Line 240: | ||
<br> | <br> | ||
<div class="heading"> | <div class="heading"> | ||
| - | + | Isolating LuxPQ in psB1AK3 from TOP10 overnights; restriction digest to verify presence of LuxPQ in BBk vector | |
</div> | </div> | ||
<br> | <br> | ||
<div class="desc"> | <div class="desc"> | ||
</html> | </html> | ||
| - | + | Purpose: isolate LuxPQ in psB1AK3 from TOP10 overnights in order to continue construction of signaling circuit. Plasmid was isolated from overnight cultures of luxPQ (6 colonies that were initially cut with EcoRI/PstI and 1 colony initially cut with XbaI/PstI). This was done using the QIAprep Spin MiniPrep Kit (QIAGEN). Plasmid purity and concentration were measured using the NanoDrop Spectrophotometer. A restriction digest was set up using NotI (in REact 3 buffer) in order to verify the presence of LuxPQ in psB1AK3. The products were then run on a 0.8% agarose gel (although no ‘uncut’ plasmid was run as a control). The gel revealed no NotI cutting, or simply on NotI site, as there was only one band in each lane. The bands were of various sizes, none necessarily revealing the presence of LuxPQ. The next step will be to redo this restriction digest. | |
<html> | <html> | ||
| Line 339: | Line 266: | ||
<br> | <br> | ||
<div class="heading"> | <div class="heading"> | ||
| - | + | Back to the Lab after Completing Initial Version of Quorum Sensing | |
</div> | </div> | ||
<br> | <br> | ||
<div class="desc"> | <div class="desc"> | ||
</html> | </html> | ||
| - | + | Quorum sensing bacteria are complete and I made bacteria with RFP and placed them beside other bacteria for display. I will probably return to work on this later, but it is now more important to finish the lab. I learned that the order in which you add materials during restriction digest is important so using the function (initially from cleaning script) created for storing the names of items in an inventory will be important to make sure that each time something is added, it is the correct item. I believe for the rest of the activity I may be able to use a modified version of my find item script. | |
| + | <br> | ||
| + | <br> | ||
| + | However, I have yet to determine a new way to move things to the water bath, heating block etc. in a creative way and still keep avatars involved. | ||
<html> | <html> | ||
| Line 372: | Line 302: | ||
R0040 (promoter), B0015 (terminator), and J13002 (RBS+Promoter) were isolated in order to verify the presence of them with Restriction digest. These parts are needed for general construction of our circuits. | R0040 (promoter), B0015 (terminator), and J13002 (RBS+Promoter) were isolated in order to verify the presence of them with Restriction digest. These parts are needed for general construction of our circuits. | ||
<html> | <html> | ||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
</div> | </div> | ||
<br> | <br> | ||
<div class="heading"> | <div class="heading"> | ||
| - | + | 2. Restriction digest of R0040, B0015, and J13002 | |
</div> | </div> | ||
<br> | <br> | ||
<div class="desc"> | <div class="desc"> | ||
</html> | </html> | ||
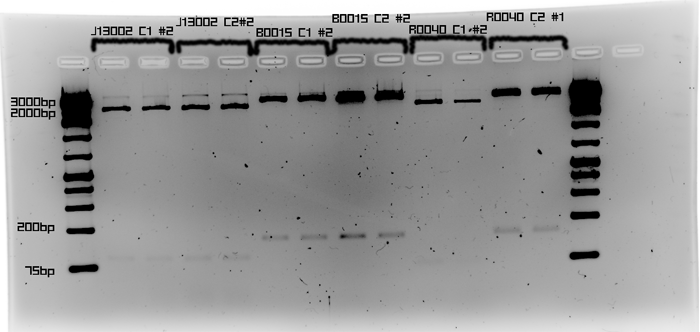
| - | + | To verify if B0015, R0040, and J13002 are present within the plasmid/cell, restriction digest was performed using NotI enzyme. | |
| + | [[Image:Calgary_2009.06.18.J13002.png|700px]] | ||
| + | <br> | ||
| + | The band in R0040 C2 #1 is the only one that did not match the expected size of each. Only one colony is needed to move on. | ||
| + | |||
<html> | <html> | ||
| Line 409: | Line 328: | ||
<tr> | <tr> | ||
<td> | <td> | ||
| - | <a name=" | + | <a name="Mandy"></a> |
<br> | <br> | ||
<div class="heading"> | <div class="heading"> | ||
| - | + | MANDY | |
</div> | </div> | ||
<br> | <br> | ||
<div class="heading"> | <div class="heading"> | ||
| - | + | Biosafety and Fumehoods in Second Life | |
</div> | </div> | ||
<br> | <br> | ||
<div class="desc"> | <div class="desc"> | ||
</html> | </html> | ||
| - | + | This morning was spent in biosafety training. Afterwards, I completed the fumehood as well as the necessary components inside the fume hood to illustrate the making of a gel for gel electrophoresis. It is now scripted to give users a new gel, which they will then take to the future gel electrophoresis station along with their amplified DNA to run a gel. | |
| - | + | ||
<html> | <html> | ||
</div> | </div> | ||
| + | <br> | ||
</td> | </td> | ||
</tr> | </tr> | ||
| Line 442: | Line 361: | ||
<br> | <br> | ||
<div class="heading"> | <div class="heading"> | ||
| - | + | June Newsletter | |
</div> | </div> | ||
<br> | <br> | ||
<div class="desc"> | <div class="desc"> | ||
</html> | </html> | ||
| - | + | In the moring, I was at the Biosafety training on main campus. (9:00am- 12:00 noon). | |
| - | + | Later that afternoon, Jamie, Fahd and I continued to work on the June newsletter. It was our goal to have it done by today. We have sent it to Sonja and Thane to double check it before we send it out. We are hoping to begin sending it out starting next week. | |
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | I made 20 colorful bake sale posters to put around the health science building. Fahd and I also made the bake sale event on Facebook and sent it to all of our friends and encouraged them to bring their friends. We also emailed the Health Science Communications coordinator to send an email out to all Health Science students to advertise our bake sale. Her generous efforts are greatly appreciated by the team. | |
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
<html> | <html> | ||
| Line 494: | Line 391: | ||
<br> | <br> | ||
<div class="heading"> | <div class="heading"> | ||
| - | + | NotI restriction digest of isolated LuxOD47A on psB1AC3 constructs. | |
</div> | </div> | ||
<br> | <br> | ||
<div class="desc"> | <div class="desc"> | ||
</html> | </html> | ||
| - | + | Purpose: Even though the colony PCR from 2 days ago had negative control contamination, we’re going to run a NotI restriction digest anyway. If we can excise a gene of the appropriate size, it is likely that the sample in question actually contains LuxOD47A on psB1AC3, as opposed to mere master mix contamination. | |
| + | |||
| + | Protocol: NotI restriction enzymes were used with REact I buffer, in accordance with the restriction digest procedure outlined on the protocol page. | ||
| + | |||
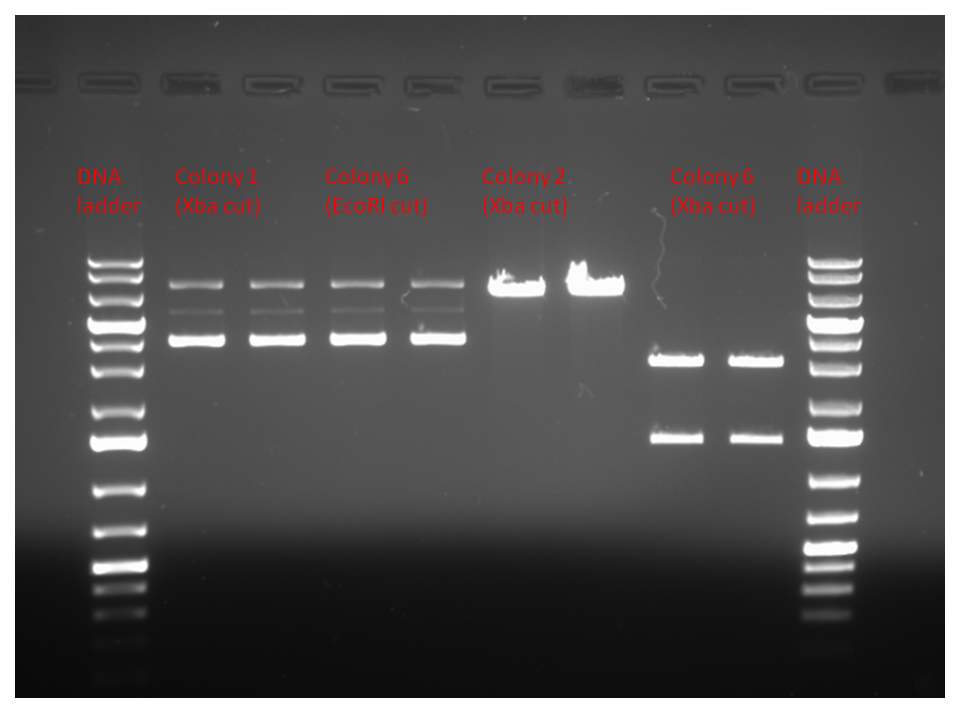
| + | Results: As shown below, the 4th set of digested genes displayed the set of bands that we expected: a band around 1400 bp (accounting for LuxOD47A) and a band around 3000 bp (accounting for psB1AC3). We will use this sample in our later constructions. | ||
| + | |||
| + | <br><br> | ||
| + | <center> | ||
| + | [[Image:June18.png|700px]] | ||
| + | </center> | ||
<html> | <html> | ||
Latest revision as of 03:06, 22 October 2009
UNIVERSITY OF CALGARY

 "
"