Team:British Columbia/pBAD
From 2009.igem.org
m (→Overview) |
|||
| (17 intermediate revisions not shown) | |||
| Line 2: | Line 2: | ||
=Arabinose sensor: the pBAD promoter= | =Arabinose sensor: the pBAD promoter= | ||
==Overview== | ==Overview== | ||
| + | |||
| + | [[Image:E_coli_Traffic_Light_Sensitive_Promoters.png|thumb|center|500px]] | ||
| + | |||
| + | |||
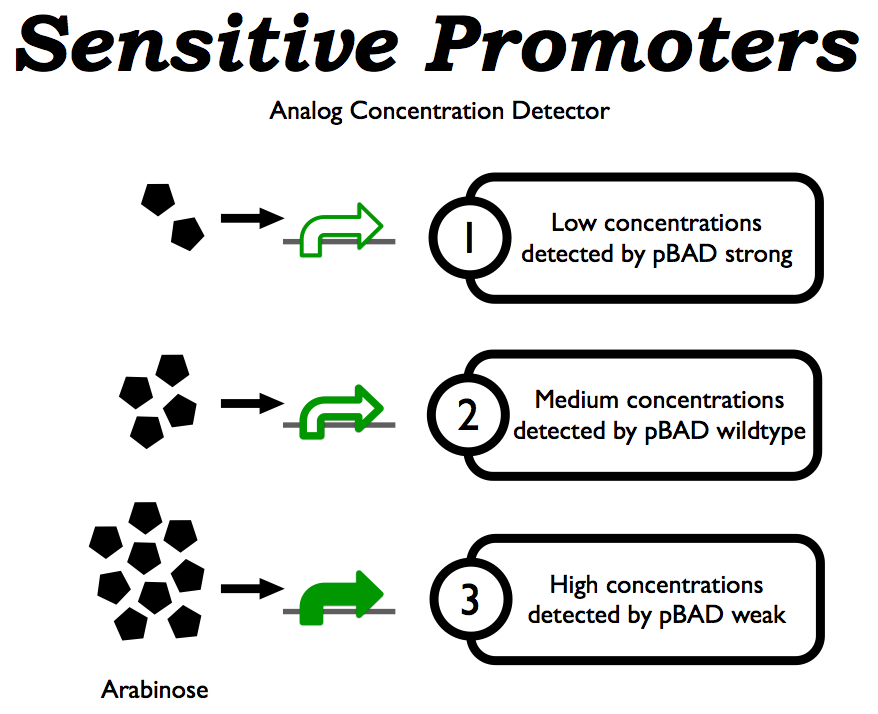
In order to make an analog biosensor, we need our traffic light to produce distinct, unique responses to a range of concentrations of an input. However, the Registry is lacking in variable strength inducible promoters. We designed two variants of the <partinfo>I13453</partinfo> pBAD promoter, one weaker and one stronger than wild type, based on AraC binding experiments performed by [[Team:British_Columbia/Bibliography|Niland et al.]] | In order to make an analog biosensor, we need our traffic light to produce distinct, unique responses to a range of concentrations of an input. However, the Registry is lacking in variable strength inducible promoters. We designed two variants of the <partinfo>I13453</partinfo> pBAD promoter, one weaker and one stronger than wild type, based on AraC binding experiments performed by [[Team:British_Columbia/Bibliography|Niland et al.]] | ||
| - | [[Image:E_coli_Traffic_Light_Mutated_Promoters.png| | + | We did the following: |
| + | # Mutagenesis of pBAD promoter sequence to create a stronger promoter (Strong pBAD) and a weaker promoter (Weak pBAD). | ||
| + | # Quantification of mutant promoter-driven RFP fluorescence. | ||
| + | # BioBrick submission. | ||
| + | |||
| + | ==pBAD Mutagenesis== | ||
| + | |||
| + | [[Image:E_coli_Traffic_Light_Mutated_Promoters.png|center|thumb||500px|A diagram of the changed nucleotide sequences of arabinose-inducible promoters. Green nucleotides of the wildtype show all sequences that were mutated from the wild type to form strong and weak.]] | ||
<!--- in case we want the text version: | <!--- in case we want the text version: | ||
| Line 14: | Line 25: | ||
--> | --> | ||
| - | Above is a sequence alignment of our promoter variants with the wild type pBAD, truncated for space. For the full sequences, see our | + | Above is a sequence alignment of our promoter variants with the wild type pBAD, truncated for space. For the full sequences, see our pBAD strong [http://partsregistry.org/wiki/index.php?title=Part:BBa_K206000 BBa_K206000] and pBAD weak [http://partsregistry.org/wiki/index.php?title=Part:BBa_K206001 BBa_K206001]. |
| + | |||
| + | ==Quantification== | ||
| + | |||
| + | <html>We assembled each promoter with the RFP reporter part <partinfo>I13507</partinfo> in order to test the relative activity of the mutated promoters</html> | ||
| + | |||
| + | <center>[[Image:Timecourse.jpg]]</center> | ||
| + | We have been able to successfully show that at low arabinose concentrations, the activity of the Strong pBAD promoter and Weak pBAD promoter following arabinose induction is, as expected, greater and lesser respectively then the Wild Type pBAD promoter. By examining the development of a RFP reporter, it is observed that the Strong pBAD promoter has both a faster rate of development and reaches a higher maximum value compared to the Wild Type sequence. Similarly, the Weak pBAD promoter develops slower and to a lower maximum intensity. | ||
| + | <center>[[Image:BW_Growth.png]]</center> | ||
| + | <center>[[Image:PBAD family - Transfer Function.png]]</center> | ||
| + | |||
| + | <br> | ||
| + | |||
| + | |||
| + | The three promoter activites were then tested using various arabinose concentrations and the data was fit to a hill equation. It is observed that Strong pBAD and Weak pBAD are more and less responsive respectively to arabinose then the Wild Type promoter. Therefore, they could be suitable to be used in conjunction as a bio-sensor. | ||
| - | + | ==BioBrick Submission== | |
| - | [ | + | Here you can find our pBAD strong [http://partsregistry.org/wiki/index.php?title=Part:BBa_K206000 BBa_K206000] and pBAD weak [http://partsregistry.org/wiki/index.php?title=Part:BBa_K206001 BBa_K206001]. |
| - | [ | + | |
| - | + | ||
Latest revision as of 04:00, 22 October 2009
Home Team Traffic Light Sensor Lock&Key Jammer [http://partsregistry.org/cgi/partsdb/pgroup.cgi?pgroup=iGEM2009&group=British_Columbia Parts] Safety Sponsors Notebook Bibliography
Contents |
Arabinose sensor: the pBAD promoter
Overview
In order to make an analog biosensor, we need our traffic light to produce distinct, unique responses to a range of concentrations of an input. However, the Registry is lacking in variable strength inducible promoters. We designed two variants of the pBAD promoter, one weaker and one stronger than wild type, based on AraC binding experiments performed by Niland et al.
We did the following:
- Mutagenesis of pBAD promoter sequence to create a stronger promoter (Strong pBAD) and a weaker promoter (Weak pBAD).
- Quantification of mutant promoter-driven RFP fluorescence.
- BioBrick submission.
pBAD Mutagenesis
Above is a sequence alignment of our promoter variants with the wild type pBAD, truncated for space. For the full sequences, see our pBAD strong [http://partsregistry.org/wiki/index.php?title=Part:BBa_K206000 BBa_K206000] and pBAD weak [http://partsregistry.org/wiki/index.php?title=Part:BBa_K206001 BBa_K206001].
Quantification
We assembled each promoter with the RFP reporter part
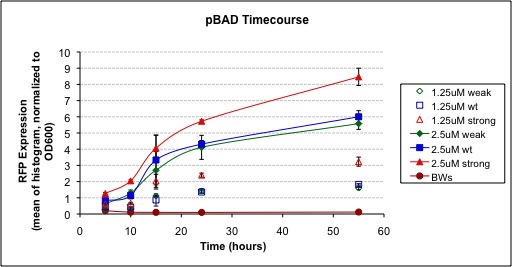
We have been able to successfully show that at low arabinose concentrations, the activity of the Strong pBAD promoter and Weak pBAD promoter following arabinose induction is, as expected, greater and lesser respectively then the Wild Type pBAD promoter. By examining the development of a RFP reporter, it is observed that the Strong pBAD promoter has both a faster rate of development and reaches a higher maximum value compared to the Wild Type sequence. Similarly, the Weak pBAD promoter develops slower and to a lower maximum intensity.
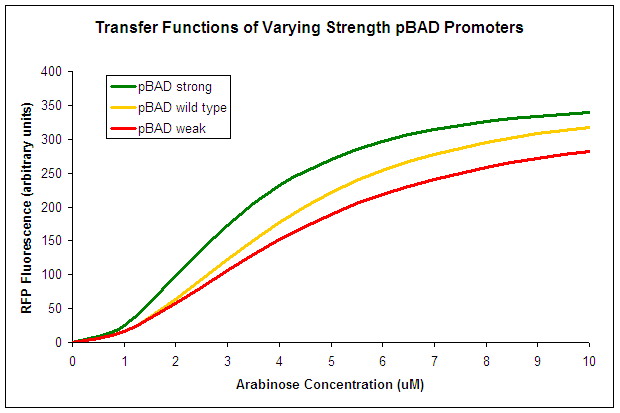
The three promoter activites were then tested using various arabinose concentrations and the data was fit to a hill equation. It is observed that Strong pBAD and Weak pBAD are more and less responsive respectively to arabinose then the Wild Type promoter. Therefore, they could be suitable to be used in conjunction as a bio-sensor.
BioBrick Submission
Here you can find our pBAD strong [http://partsregistry.org/wiki/index.php?title=Part:BBa_K206000 BBa_K206000] and pBAD weak [http://partsregistry.org/wiki/index.php?title=Part:BBa_K206001 BBa_K206001].
 "
"