Team:Imperial College London/M3
From 2009.igem.org
(→Module 3: Genome Deletion) |
|||
| (43 intermediate revisions not shown) | |||
| Line 1: | Line 1: | ||
{{Imperial/09/TemplateTop}} | {{Imperial/09/TemplateTop}} | ||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
<html> | <html> | ||
<center> | <center> | ||
| - | <img src=" | + | <div class="highslide-gallery"> |
| + | <a href="https://static.igem.org/mediawiki/2009/8/8d/II09_MapIndicator_Module3.png" class="highslide" onclick="return hs.expand(this, config1)" title="After thermoinduction, restriction enzymes are expressed that remove the genetic material"> | ||
| + | <img src="https://static.igem.org/mediawiki/2009/8/8d/II09_MapIndicator_Module3.png" alt="" title="Click to enlarge" width="75%"/> | ||
| + | </a> | ||
| + | <div class="highslide-caption"> | ||
| + | Module 2: Encapsulation | ||
| + | </div> | ||
| + | </div> | ||
</center> | </center> | ||
</html> | </html> | ||
| + | =<!--[[Image:II09_Thumb_m3.png|40px]]--><font size='5'><b>Module 3: Genome Deletion</b></font>= | ||
<br> | <br> | ||
[[Image:II09_transition_module3.jpg|center|400px]] | [[Image:II09_transition_module3.jpg|center|400px]] | ||
| - | + | <b>Module 3</b> is the final module of the system. <b><i>The E.ncapsulator</i></b> has successfully completed its job of protein production (module 1) and encapsulation (module 2). Now, it needs to be prepared to be converted into a safe pill carrying the protein of interest. This is done by removing the genetic material which renders the cell inanimate. <br> | |
| - | + | ||
| - | <b>Module 3</b> is the final module of the system. <b><i>The E.ncapsulator</i></b> has successfully completed its job of | + | |
| - | + | ||
<br> | <br> | ||
| + | [[Image:II09_Module3reusable.jpg|right|200px]] | ||
| - | ==== | + | ==Rationale== |
| + | Module 3 acts as a <b>reusable</b> module for <b>removal of genetic material</b> without toxic effects.<br> | ||
<br> | <br> | ||
| - | Removal of genetic material by the use of restriction enzymes prevents the accidental transfer of DNA to other gut microflora, which could lead to development of virulence. This module is a highly reusable | + | Removal of genetic material by the use of restriction enzymes prevents the accidental transfer of DNA to other gut microflora, which could lead to development of virulence. This module is a highly reusable for any chassis system where there is a need to remove genetic material after genes are expressed. <br> |
| + | <br> | ||
| + | Our pill is to be <b>consumed</b> within the human body. This rules out the <b>toxin-generating</b> methods to induce cell death. Restriction enzymes are the preferred method for inducing cell death as they ar relatively harmless outside of the cell. <br> | ||
<br> | <br> | ||
<br> | <br> | ||
| - | |||
| - | |||
| - | |||
| - | ==== | + | ==Theory== |
| - | + | ===Engineering cell death=== | |
| - | + | Due to the possible <b>pathogenicity and health concerns</b>, cell death must occur before the pill is ready for consumption. Therefore, the method chosen needs to be foolproof and have <b>failsafe mechanism</b>. <br> | |
| - | + | <br> | |
| + | [[Image:M3gci2.jpg|600px]] | ||
| + | <br><br> | ||
<html><a href="https://2009.igem.org/Team:Imperial_College_London/M3/Genetic | <html><a href="https://2009.igem.org/Team:Imperial_College_London/M3/Genetic | ||
| - | "><img style="vertical-align:bottom;" width=50px align="left" src="http://i691.photobucket.com/albums/vv271/dk806/II09Learnmore.png"></a></html | + | "><img style="vertical-align:bottom;" width=50px align="left" src="http://i691.photobucket.com/albums/vv271/dk806/II09Learnmore.png"></a></html><b> About our genetic circuit</b> |
| - | + | <br><br> | |
| + | <br> | ||
| + | Under the control of a thermoinducible promoter system ([http://partsregistry.org/Part:BBa_K098995 K098995]), when the temperature is raised, the promoter is activated and restriction enzymes are produced. There is a safeguard here as the temperature of the human body is around 37°C, so that even if the bacteria are not killed by the heat pulse, they will be killed after they enter the human body. <br> | ||
| + | <br> | ||
| + | The restriction enzymes DpnII ([http://partsregistry.org/Part:BBa_K200009 K200009]) and TaqI ([http://partsregistry.org/Part:BBa_K200010 K200010]) are produced. This duplicity of restriction enzymes ensures that even when one enzyme becomes mutated and dysfunctional, the other restriction enzyme still works well by itself. Therefore, by using two restriction enzymes, we can be more certain that our DNA has been digested. | ||
| + | <br> | ||
<br> | <br> | ||
| - | |||
<html><a href="https://2009.igem.org/Team:Imperial_College_London/M3/RestrictionEnzymes | <html><a href="https://2009.igem.org/Team:Imperial_College_London/M3/RestrictionEnzymes | ||
"><img style="vertical-align:bottom;" width=50px align="left" src="http://i691.photobucket.com/albums/vv271/dk806/II09Learnmore.png"></a></html><b> About Restriction Enzymes</b> | "><img style="vertical-align:bottom;" width=50px align="left" src="http://i691.photobucket.com/albums/vv271/dk806/II09Learnmore.png"></a></html><b> About Restriction Enzymes</b> | ||
| - | + | <br><br> | |
<br> | <br> | ||
| + | Dam methylase ([http://partsregistry.org/Part:BBa_K200001 K200001]) is constitutively produced at a low amount. This prevents leaky expression of restriction enzymes from damaging the genome prematurely. Consequently, a balance exists between Dam methylation and restriction enzyme activity. <br><br> | ||
<html><a href="https://2009.igem.org/Team:Imperial_College_London/M3/DamMethylation | <html><a href="https://2009.igem.org/Team:Imperial_College_London/M3/DamMethylation | ||
"><img style="vertical-align:bottom;"width=50px align="left" src="http://i691.photobucket.com/albums/vv271/dk806/II09Learnmore.png"></a></html><b> About Methylation</b> | "><img style="vertical-align:bottom;"width=50px align="left" src="http://i691.photobucket.com/albums/vv271/dk806/II09Learnmore.png"></a></html><b> About Methylation</b> | ||
| - | + | ==Results== | |
| - | + | ===Wet Lab=== | |
| - | ==== | + | |
| - | + | ||
| - | + | ||
| - | + | ||
[[Image:II09 DpnII Digest.png|right|400px]] | [[Image:II09 DpnII Digest.png|right|400px]] | ||
The activity of the restriction enzymes is critical to module 3. We have tested this using a genomic digest assay. <br> | The activity of the restriction enzymes is critical to module 3. We have tested this using a genomic digest assay. <br> | ||
| Line 79: | Line 65: | ||
<html><a href="https://2009.igem.org/Team:Imperial_College_London/Wetlab/Results#Module_3 | <html><a href="https://2009.igem.org/Team:Imperial_College_London/Wetlab/Results#Module_3 | ||
| - | "><img style="vertical-align:bottom;" width=50px align="left" src="http://i691.photobucket.com/albums/vv271/dk806/II09Learnmore.png"></a></html> | + | "><img style="vertical-align:bottom;" width=50px align="left" src="http://i691.photobucket.com/albums/vv271/dk806/II09Learnmore.png"></a></html> <b> About our wet lab results</b> |
<br> | <br> | ||
| Line 85: | Line 71: | ||
<br> | <br> | ||
| - | + | ===Dry Lab=== | |
| - | + | ||
We have also attempted to link our restriction enzymes with cell death using a model.<br> | We have also attempted to link our restriction enzymes with cell death using a model.<br> | ||
<br> | <br> | ||
| Line 93: | Line 78: | ||
<html><a href="https://2009.igem.org/Team:Imperial_College_London/Drylab/Genome_deletion | <html><a href="https://2009.igem.org/Team:Imperial_College_London/Drylab/Genome_deletion | ||
| - | "><img style="vertical-align:bottom;" width=50px align="left" src="http://i691.photobucket.com/albums/vv271/dk806/II09Learnmore.png"></a></html> | + | "><img style="vertical-align:bottom;" width=50px align="left" src="http://i691.photobucket.com/albums/vv271/dk806/II09Learnmore.png"></a></html> <b> About our dry lab results</b><br> |
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
<br> | <br> | ||
| - | + | ===Results summary=== | |
| - | + | We have shown that cells can be protected from low concentrations of the restriction enzymes DpnII and TaqI by Dam methylation, and how the cell population rapidly decreases with thermoinduction of restriction enzymes. <br> | |
<br> | <br> | ||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
<html><center></html> | <html><center></html> | ||
| Line 139: | Line 113: | ||
</tr></table></html> | </tr></table></html> | ||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
{{Imperial/09/TemplateBottom}} | {{Imperial/09/TemplateBottom}} | ||
Latest revision as of 03:54, 22 October 2009

Contents |
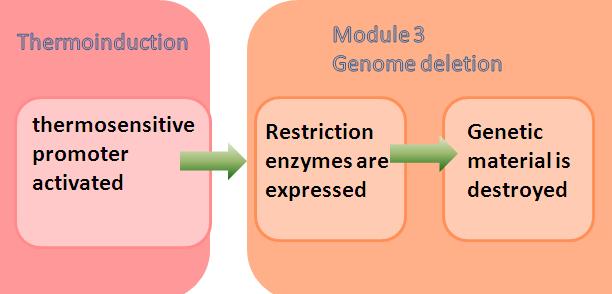
Module 3: Genome Deletion
Module 3 is the final module of the system. The E.ncapsulator has successfully completed its job of protein production (module 1) and encapsulation (module 2). Now, it needs to be prepared to be converted into a safe pill carrying the protein of interest. This is done by removing the genetic material which renders the cell inanimate.
Rationale
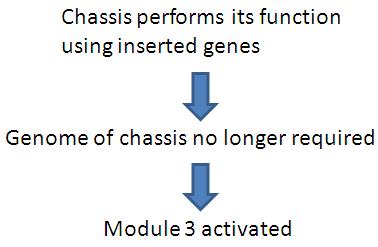
Module 3 acts as a reusable module for removal of genetic material without toxic effects.
Removal of genetic material by the use of restriction enzymes prevents the accidental transfer of DNA to other gut microflora, which could lead to development of virulence. This module is a highly reusable for any chassis system where there is a need to remove genetic material after genes are expressed.
Our pill is to be consumed within the human body. This rules out the toxin-generating methods to induce cell death. Restriction enzymes are the preferred method for inducing cell death as they ar relatively harmless outside of the cell.
Theory
Engineering cell death
Due to the possible pathogenicity and health concerns, cell death must occur before the pill is ready for consumption. Therefore, the method chosen needs to be foolproof and have failsafe mechanism.
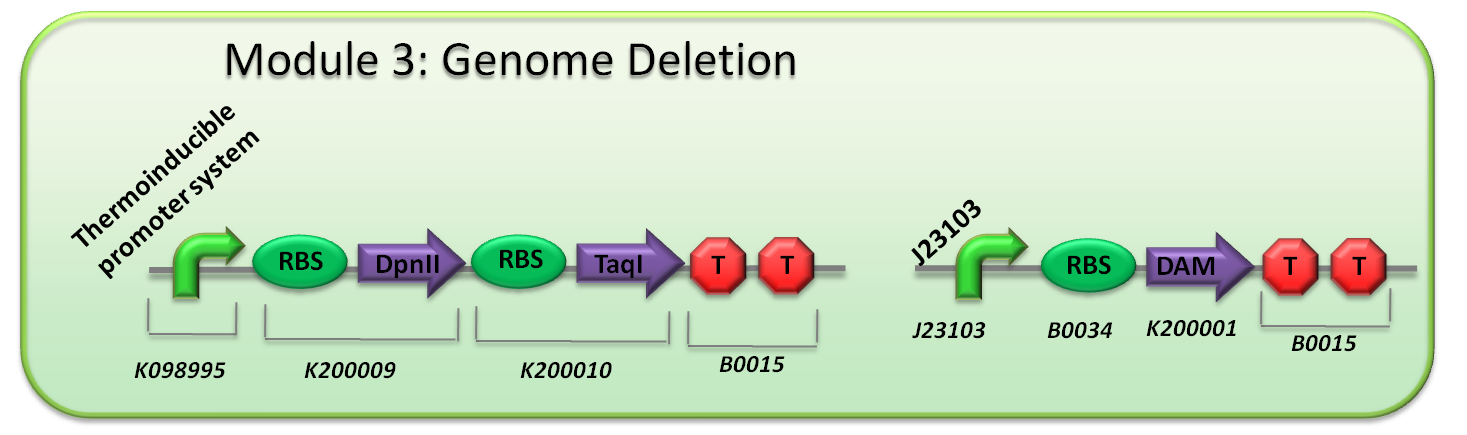
 About our genetic circuit
About our genetic circuit
Under the control of a thermoinducible promoter system ([http://partsregistry.org/Part:BBa_K098995 K098995]), when the temperature is raised, the promoter is activated and restriction enzymes are produced. There is a safeguard here as the temperature of the human body is around 37°C, so that even if the bacteria are not killed by the heat pulse, they will be killed after they enter the human body.
The restriction enzymes DpnII ([http://partsregistry.org/Part:BBa_K200009 K200009]) and TaqI ([http://partsregistry.org/Part:BBa_K200010 K200010]) are produced. This duplicity of restriction enzymes ensures that even when one enzyme becomes mutated and dysfunctional, the other restriction enzyme still works well by itself. Therefore, by using two restriction enzymes, we can be more certain that our DNA has been digested.
 About Restriction Enzymes
About Restriction Enzymes
Dam methylase ([http://partsregistry.org/Part:BBa_K200001 K200001]) is constitutively produced at a low amount. This prevents leaky expression of restriction enzymes from damaging the genome prematurely. Consequently, a balance exists between Dam methylation and restriction enzyme activity.
Results
Wet Lab
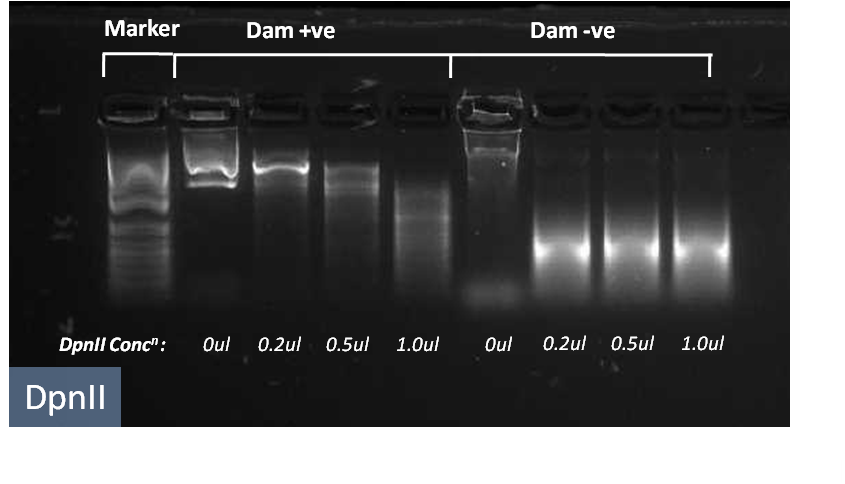
The activity of the restriction enzymes is critical to module 3. We have tested this using a genomic digest assay.
The restriction enzymes DpnII and TaqI are shown to cut genomic DNA into small fragments, shown on the right by a smear of bands. We have further tested the restriction enzymes in DNA which have been methylated by Dam enzymes and shown that there is essentially no cleavage at low concentrations of restriction enzymes.
Dry Lab
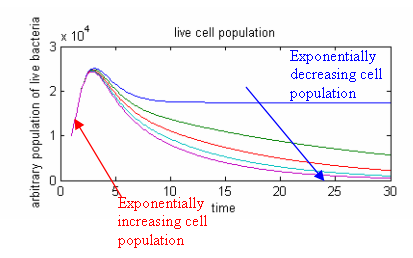
We have also attempted to link our restriction enzymes with cell death using a model.
The population increase is initially exponential as the restriction enzymes have a delay in production. As the restriction enzymes accumulate in the cell, the cell growth starts to slow down. If the lambda cI promoter is strong enough, killing rate will greatly exceed cell division rate, and there will be an exponential decrease in cell population.
Results summary
We have shown that cells can be protected from low concentrations of the restriction enzymes DpnII and TaqI by Dam methylation, and how the cell population rapidly decreases with thermoinduction of restriction enzymes.
Project Tour


Module 3 Contents





 "
"