Team:Kyoto/GSDD/Results
From 2009.igem.org
(→Experimental Results) |
|||
| (24 intermediate revisions not shown) | |||
| Line 21: | Line 21: | ||
#[[Team:Kyoto/GSDD|GSDD]] | #[[Team:Kyoto/GSDD|GSDD]] | ||
#Reslts & Discussion | #Reslts & Discussion | ||
| + | {{:Team:Kyoto/topicpath}} | ||
</div><!-- /#topicpath --> | </div><!-- /#topicpath --> | ||
{{:Team:Kyoto/Leftnavi}} | {{:Team:Kyoto/Leftnavi}} | ||
| Line 28: | Line 29: | ||
==Experimental Results== | ==Experimental Results== | ||
| - | |||
| - | + | ===Result of subgoal-1; Microgene Polymerization Reaction (MPR) for constructing repetitive LacI binding sites=== | |
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | === | + | |
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
We achieved constructing repetitive sequence of LacI binding site. A certain length of LacI binding site is required in both ends of Timer vector. We expected that LacI binds to this site and protects the both ends of Timer vector from the degradation caused by exonuclease, so the both ends of Timer vector become shorter per duplication by the end replication problem. In short, the repetitive sequence counts the time. Also, constructing various length of repetitive sequence was necessary because we did not evaluate the DNA stability by binding LacI. | We achieved constructing repetitive sequence of LacI binding site. A certain length of LacI binding site is required in both ends of Timer vector. We expected that LacI binds to this site and protects the both ends of Timer vector from the degradation caused by exonuclease, so the both ends of Timer vector become shorter per duplication by the end replication problem. In short, the repetitive sequence counts the time. Also, constructing various length of repetitive sequence was necessary because we did not evaluate the DNA stability by binding LacI. | ||
As a result, we constructed a set of repetitive sequence that contains multiple LacI binding site by MPR (Microgene Polymerization Reaction). MPR suits our experiment because various length of DNA are constructed at once without ligation. | As a result, we constructed a set of repetitive sequence that contains multiple LacI binding site by MPR (Microgene Polymerization Reaction). MPR suits our experiment because various length of DNA are constructed at once without ligation. | ||
| Line 48: | Line 37: | ||
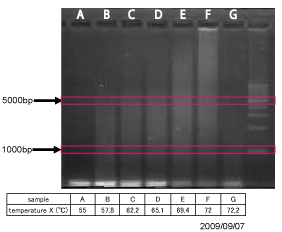
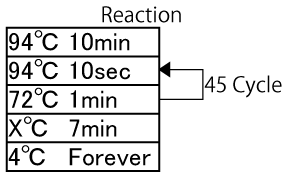
====1) Experimental conditions in MPR==== | ====1) Experimental conditions in MPR==== | ||
| - | [[Image:figure(reaction2).png| | + | [[Image:figure(reaction2).png|220px|center|Fig.1]] |
| - | [[Image:figure(MPR1).png| | + | [[Image:figure(MPR1).png|420px|center|Fig.1]] |
| - | [[Image:figure(MPR2).png| | + | [[Image:figure(MPR2).png|420px|center|Fig.2]] |
We tried various conditions in the temperature X. The result indicated that the temperature during MPR influences the length of the acquired sequence. At last, we programmed 72 o C of the temperature X. | We tried various conditions in the temperature X. The result indicated that the temperature during MPR influences the length of the acquired sequence. At last, we programmed 72 o C of the temperature X. | ||
Fig shows the repetitive sequence that we used for designing parts. As shown in Fig. 4, we succeeded in constructing the polymerized microgenes with different number of LacI binding sites. | Fig shows the repetitive sequence that we used for designing parts. As shown in Fig. 4, we succeeded in constructing the polymerized microgenes with different number of LacI binding sites. | ||
| Line 59: | Line 48: | ||
In conclusion, we successfully designed and constructed the repetitive sequences with the multiple protein binding sites by MPR. This is the simple method to construct any repetitive DNA sequence. Thus, it might be applicable to synthesize other repetitive sequences with different lengths in a tailor-made manner. | In conclusion, we successfully designed and constructed the repetitive sequences with the multiple protein binding sites by MPR. This is the simple method to construct any repetitive DNA sequence. Thus, it might be applicable to synthesize other repetitive sequences with different lengths in a tailor-made manner. | ||
| - | ===Result-2 Construction of Timer vector that contains LacI generator and TIMER=== | + | '''blunt end clonig by Stu1 (BBa_K210014)''' |
| - | + | ||
| + | [[Image:Length_check_blunt_end_clonig_by_Stu1_(BBa_K210014).png]] | ||
| + | |||
| + | |||
| + | |||
| + | ===Parts sequence=== | ||
| + | We sequenced features and parts which we constructed. The results of sequences are matched as we expect. | ||
| + | If you see the detailed data, click [[Team:Kyoto/sequence|here]]. | ||
| + | |||
| + | |||
| + | |||
| + | ===Result of subgoal-2 Construction of Timer vector that contains LacI generator and TIMER=== | ||
| + | |||
| + | '''Timer vector #1600 for measurement (BBa_K210008)''' | ||
| + | |||
| + | [[Image:Timer_vector_check.png]] | ||
| + | |||
| + | ===Result of subgoal-3 Construction of the constitutive LacI expression vector=== | ||
| + | |||
| + | '''lacI generator for yeast (BBa_K210005)''' | ||
| - | + | [[Image:32.png]] | |
| - | + | ||
===Result-4 Analysis of the constructed TIMER vector in yeast=== | ===Result-4 Analysis of the constructed TIMER vector in yeast=== | ||
| - | + | No data. | |
==Discussion== | ==Discussion== | ||
Latest revision as of 02:55, 22 October 2009
Experimental Results
Result of subgoal-1; Microgene Polymerization Reaction (MPR) for constructing repetitive LacI binding sites
We achieved constructing repetitive sequence of LacI binding site. A certain length of LacI binding site is required in both ends of Timer vector. We expected that LacI binds to this site and protects the both ends of Timer vector from the degradation caused by exonuclease, so the both ends of Timer vector become shorter per duplication by the end replication problem. In short, the repetitive sequence counts the time. Also, constructing various length of repetitive sequence was necessary because we did not evaluate the DNA stability by binding LacI. As a result, we constructed a set of repetitive sequence that contains multiple LacI binding site by MPR (Microgene Polymerization Reaction). MPR suits our experiment because various length of DNA are constructed at once without ligation.
It was suggested that the temperature during MPR influences the length of the acquired sequence. Because we aimed to construct a set of repetitive DNA with different lengths, we investigated and optimized the temperature condition during MPR. After getting repetitive sequences, we inserted them into plasmid by flush end ligation, and designed the part that has repetitive sequence.
1) Experimental conditions in MPR
We tried various conditions in the temperature X. The result indicated that the temperature during MPR influences the length of the acquired sequence. At last, we programmed 72 o C of the temperature X. Fig shows the repetitive sequence that we used for designing parts. As shown in Fig. 4, we succeeded in constructing the polymerized microgenes with different number of LacI binding sites.
2) Designing the repetitive part of LacI binding site
We can not embed restriction site in the both ends of repetitive sequence part because it is consists of repeating short DNA. Embedding restriction site in each short primer leads to the degradation into small parts when digesting by enzyme. Therefore, we inserted it into plasmid by blunt end ligation. The repetitive part was inserted into PSB1A2 which has blunt end. In this process, we phosphorylated PSB1A2, and dephosphorylated the repetetitive. We purified them before ligation. In conclusion, we successfully designed and constructed the repetitive sequences with the multiple protein binding sites by MPR. This is the simple method to construct any repetitive DNA sequence. Thus, it might be applicable to synthesize other repetitive sequences with different lengths in a tailor-made manner.
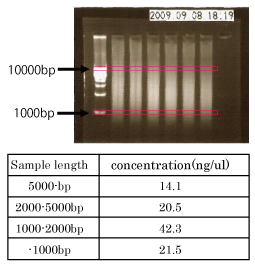
blunt end clonig by Stu1 (BBa_K210014)
Parts sequence
We sequenced features and parts which we constructed. The results of sequences are matched as we expect. If you see the detailed data, click here.
Result of subgoal-2 Construction of Timer vector that contains LacI generator and TIMER
Timer vector #1600 for measurement (BBa_K210008)
Result of subgoal-3 Construction of the constitutive LacI expression vector
lacI generator for yeast (BBa_K210005)
Result-4 Analysis of the constructed TIMER vector in yeast
No data.
Discussion
The difference between our designed device and YAC
The difference between our designed TIMER and YAC In the case of YAC, both ends is composed of teromeric site. Thus, when some of the end sequences of them are deleted gradually by the end replication problem, the deleted sequences will be repaired. But in the case of our system, the deleted sequence will not be repaired because it does not contain the telomeric sites. Therefore, the length of the TIMER vector will become shorter as cell division proceeds. We assume that the most critical problem in this system is that we do not know exactly how many DNA bases are removed during each cell division and how differently the system works between each cell. We assume that about 100-200 base pairs (this length is same with the average primer sequence in the ragging chain replication of yeast cells) are removed during one cell division. Thus, we constructed a set of repetitive DNA sequences with the different number of LacI binding sites to optimize the condition in our system.
Evaluation for the inhibition of degradation
If we model Timer vector, we can upgrade the timer function, and apply Timer vector to more various uses. In order to model it, it is necessary to evaluate the time to that the repetitive sequence is completely lost and Timer vector is degraded. We need to analyze the length of DNA which becomes shorter during each cell division.
The both ends of Timer vector become short because of the end replication problem as already discussed. However, the both ends are also degraded by exonuclease because the LacI that binds to the repetitive sequences does not completely protect and stabilize the both end of Timer. Therefore, evaluation for the inhibition of degradation by LacI is important.
Thus, to evaluate the inhibition by LacI binding, we need to verify the length of linear DNA which becomes short not by the end replication problem but by exonuclease, after the Timer Vector is transformed.
In previous work, it has been reported that a designed linear DNA was stabilized in E. coli. by the binding of lactose repressor to the both ends of the designed repeated binding sequences (REFERENCE). They quantified the inhibition efficiency by the protein expression level, by majoring the amount of fluorescence of GFP expressed from the linear DNA. They compared (a) and (b) below. They observed them in cell-free gene expression system, RTS (rapid translation system, Roche diagnostics), in the presence of the exonuclease.
- (a) linear DNA which can express GFP
- (b) linear DNA which can express GFP and have LacI binding site in both ends
After (a) and (b) were expressed, the amount of fluorescence in (b) was more than (a). In addition, when they increased the concentration of LacI in RTS, the difference of them became bigger. This work indicated that LacI in ends inhibit the degradation by exonuclease. In previous work, they quantified the inhibition not by the length of degraded DNA but by the expression level, and they used E coli. No studies have ever evaluated the length of DNA degraded by exonuclease and the length protected by LacI in Saccharomyces cerevisiae. We thought of two following ideas to investigate them.
The first idea is to observe linear DNA which have various length of LacI binding site in both ends, and measure the time to stop the expression of GFP. In this idea, much data is necessary to estimate the parameters to model it. It is thought to be difficult to estimate the parameters, because this data is supposed to vary because the end replication problem itself doesn’t happen evenly. So the second idea is supposed to be more realistic. It is to stop the duplication, and measure the length of linear DNA which is shortened only by exonuclease. After the Timer vector is transformed, we stop the duplication in some way. We can observe the expression of GFP, and the time to stop the expression. This enables us to quantify the inhibition by the length of degraded DNA, where the end replication problem doesn’t occur.
To sum up, in order to model the Timer vector, we need to evaluate the length of DNA degraded by exonuclease besides the end replication problem. This is expected to be possible by some experiments noted above.
The instability aspect of linear vector
The ends of DNA double-strand break are repaired by homologous recombination or nonhomologous end-joining after they are degraded by exonuclease inside the cell. Therefore, linear vector is supposed to be instable because there is a possibility of recombination with other DNA or circularization in addition to DNA degradation. Binding LacI or other DNA binding protein in both ends is expected to inhibit degradation, recombination and circularization. For example, telomeric DNA inside the cell has such a mechanism (REFERENCE). Timer vector has LacI binding sites in both ends. However, there still may be the instability aspect of it. In order to reinforce this end protection, the following ideas are likely to be effective.
- Use other binding protein which has stronger binding force, instead of LacI
- Use mutant yeast with loss of function of a protein related DNA degradation
By adopting these ideas, suppression of the instability is supposed to be stronger.
 "
"