Team:Imperial College London/Chemoinduction
From 2009.igem.org
(→Project Tour) |
|||
| (7 intermediate revisions not shown) | |||
| Line 3: | Line 3: | ||
<center> | <center> | ||
<div class="highslide-gallery"> | <div class="highslide-gallery"> | ||
| - | <a href="https://static.igem.org/mediawiki/2009/5/5c/II09_MapIndicator_Chemoinduction.png" class="highslide" onclick="return hs.expand(this, config1) title="When the population of cells increases to a sufficient density, module 1 is induced by IPTG addition"> | + | <a href="https://static.igem.org/mediawiki/2009/5/5c/II09_MapIndicator_Chemoinduction.png" class="highslide" onclick="return hs.expand(this, config1)" title="When the population of cells increases to a sufficient density, module 1 is induced by IPTG addition"><img src="https://static.igem.org/mediawiki/2009/5/5c/II09_MapIndicator_Chemoinduction.png" alt="" title="Click to enlarge" width="75%"/></a> |
| - | <img src="https://static.igem.org/mediawiki/2009/5/5c/II09_MapIndicator_Chemoinduction.png" alt="" title="Click to enlarge" width="75%"/> | + | |
| - | </a> | + | |
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
</div> | </div> | ||
</center> | </center> | ||
| Line 17: | Line 11: | ||
<center> | <center> | ||
<div class="highslide-gallery"> | <div class="highslide-gallery"> | ||
| - | + | ||
<a href="https://static.igem.org/mediawiki/2009/7/70/II09_M1_Chart.png" class="highslide" onclick="return hs.expand(this, config1)"> | <a href="https://static.igem.org/mediawiki/2009/7/70/II09_M1_Chart.png" class="highslide" onclick="return hs.expand(this, config1)"> | ||
<img src="https://static.igem.org/mediawiki/2009/7/70/II09_M1_Chart.png" alt="" title="Click to enlarge" width="75%"/> | <img src="https://static.igem.org/mediawiki/2009/7/70/II09_M1_Chart.png" alt="" title="Click to enlarge" width="75%"/> | ||
| Line 90: | Line 84: | ||
<Center> | <Center> | ||
===Project Tour=== | ===Project Tour=== | ||
| + | <html><center><a href="https://2009.igem.org/Team:Imperial_College_London/Project_Overview"><img width=150px src="http://i691.photobucket.com/albums/vv271/dk806/ProjectOverviewL.jpg"></a><a href="https://2009.igem.org/Team:Imperial_College_London/M1"><img width=150px src="http://i691.photobucket.com/albums/vv271/dk806/Module1R.jpg"></a></center> | ||
| + | </html> | ||
| + | <br> | ||
| + | <hr> | ||
| + | ===Temporal Control Contents=== | ||
| + | <html><center><a href="https://2009.igem.org/Team:Imperial_College_London/Temporal_Control/Chemical_Induction"><img style="vertical-align:bottom;" width="20%" src="http://i691.photobucket.com/albums/vv271/dk806/II09_chemicalinduction.png"></a><a href="https://2009.igem.org/Team:Imperial_College_London/Temporal_Control/Autoinduction"><img style="vertical-align:bottom;" width="20%" src="http://i691.photobucket.com/albums/vv271/dk806/II09_Drylabmainimage1.png"></a><a href="https://2009.igem.org/Team:Imperial_College_London/Temporal_Control/Thermoinduction"><img style="vertical-align:bottom;" width="20%" src="http://i691.photobucket.com/albums/vv271/dk806/II09_Thermoinduction1.png"></a><a href="https://2009.igem.org/Team:Imperial_College_London/Wetlab/Results#Temporal_Control"><img style="vertical-align:bottom;"width="20%"src="http://i691.photobucket.com/albums/vv271/dk806/II09_Wetlabmainimage9.png"></a><a href="https://2009.igem.org/Team:Imperial_College_London/Drylab/Autoinduction"><img style="vertical-align:bottom;" width="20%" src="http://i691.photobucket.com/albums/vv271/dk806/II09_Drylabmainimage6.png"></a></center></html> | ||
| - | <html><center><a href="https://2009.igem.org/Team:Imperial_College_London/ | + | <html><table border="0" style="background-color:transparent;" width="100%"> |
| - | </ | + | <tr><td width="0%"></td> |
| + | <td width="20%"><center><a href="https://2009.igem.org/Team:Imperial_College_London/Temporal_Control/Chemical_Induction"><b>Chemoinduction</b></a></center></td> | ||
| + | <td width="20%"><center><a href="https://2009.igem.org/Team:Imperial_College_London/Temporal_Control/Autoinduction"><b>Autoinduction</b></a></center></td> | ||
| + | <td width="20%"><center><a href="https://2009.igem.org/Team:Imperial_College_London/Temporal_Control/Thermoinduction"><b>Thermoinduction</b></a></center></td> | ||
| + | <td width="20%"><center><a href="https://2009.igem.org/Team:Imperial_College_London/Wetlab/Results#Temporal_Control"><b>Wet Lab</b></a></center></td> | ||
| + | <td width="20%"><center><a href="https://2009.igem.org/Team:Imperial_College_London/Drylab/Autoinduction"><b>Modelling</b></a></center></td></table></html> | ||
{{Imperial/09/TemplateBottom}} | {{Imperial/09/TemplateBottom}} | ||
Latest revision as of 03:48, 22 October 2009

Contents |
Module Integration - Chemoinduction
Chemoinduction
Overview
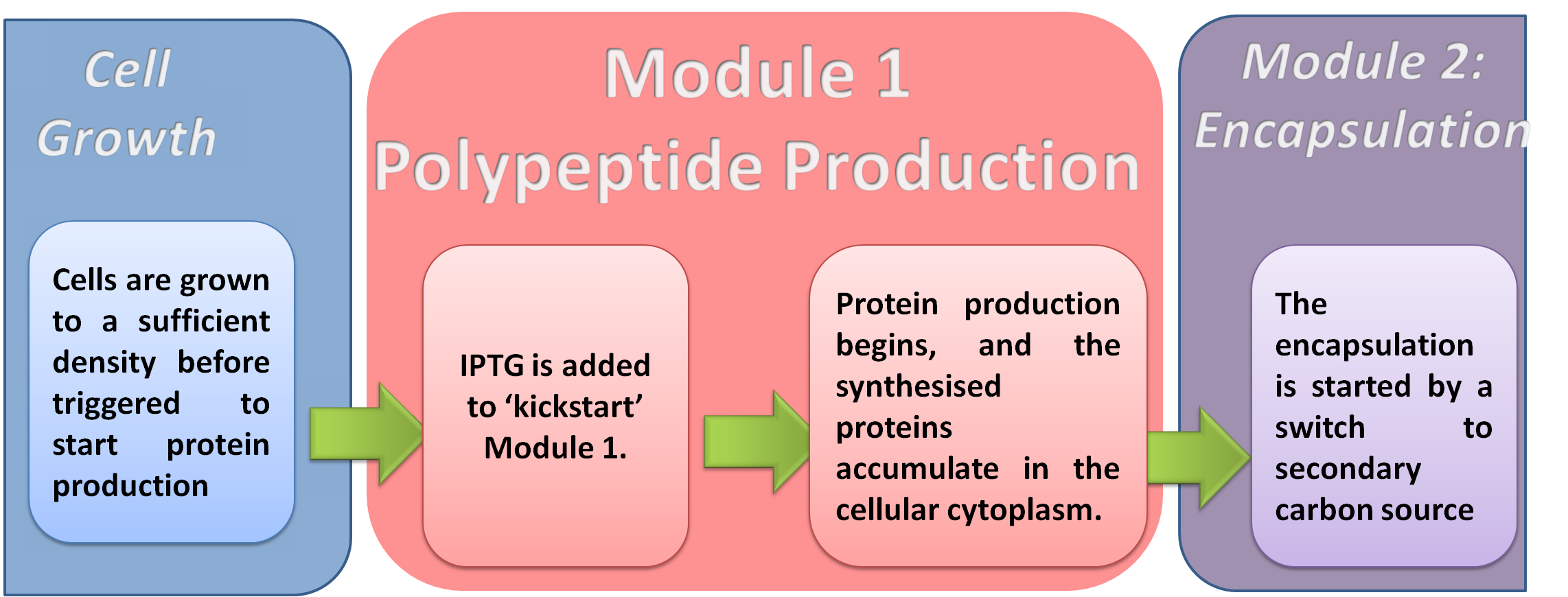
Our system is 'kick started' by the addition of a compound, IPTG, which activates protein production (module 1) of our system. This is achieved by putting the genes for protein production (module 1) under control of a chemically inducible genetic part called a promoter.
Rationale
In designing our system, we chose to use an inducible (switchable) promoter rather than a constitutive (always ON) promoter. This is to allow the cells to grow to a certain cell density before forcing them to produce protein. The production of protein can be a burden on the cell's resources, especially at the early stages of cell growth. Therefore, the culture should only be induced by the addition of IPTG once it has reached a critical cell density.

About our reasons for choosing chemical induction
Theory
The Lac Operon
The Lac Operon is a prime example of a genetic operon (several genes under control of one promoter), and has been widely studied in the field of genetics.
The Lac Operon is a set of genes found in E. Coli that control lactose metabolism in times when preferable sugars (ie. glucose) are scarce. When glucose levels are high, the E.coli feed on the glucose. However, when glucose is not available, E.coli has a set of genes coding for the enzymes responsible for the metabolism of the secondary carbon source, in this case, lactose.
In order to control the activation of Module 1, the team is using the same promoter as that found in the lac operon.
Genetic Circuit
These genetic circuits are the designs upon which the functionality of The E.ncapsulator is based. We will go through the genetic design of the system, module by module. If you are unfamiliar with the symbols used, then check out our key to familiarise yourself with each circuit symbol and what it means.
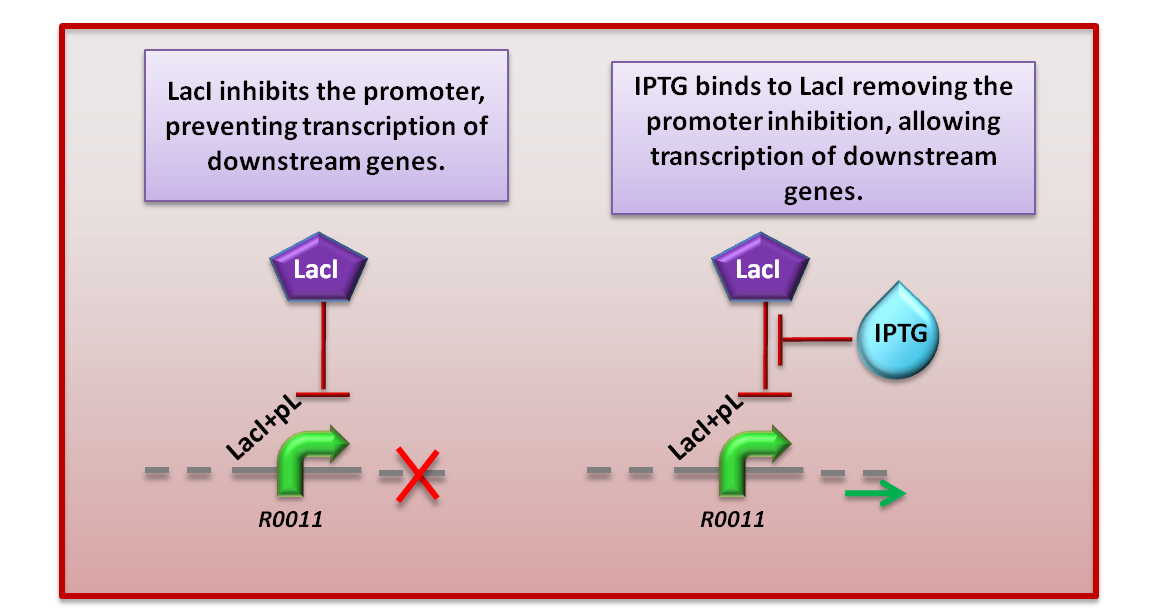
This diagram makes up part of the Module 1 genetic circuit
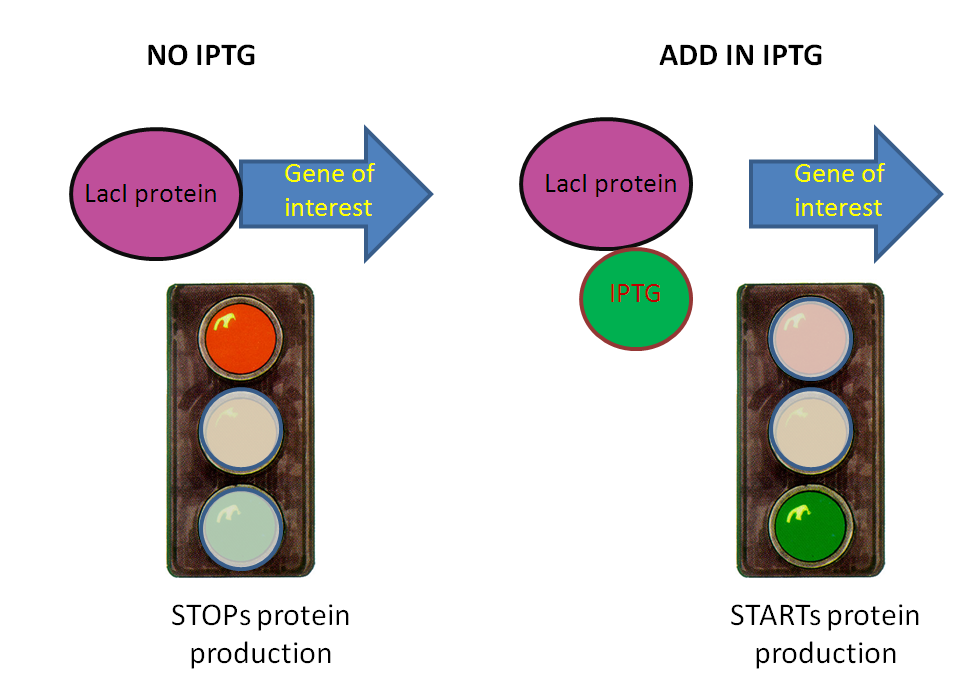
Under normal circumstances (without IPTG), the Lac inhibitor (LacI) is constitutively present in the system. This means that the promoter is repressed, and blocks the expression of the downstream genes.
When IPTG is added, it binds to LacI and cancels the repression on the promoter. This allows for the expression of the downstream genes, and therefore for the start of Module 1.
Induction with IPTG
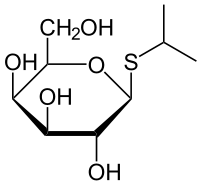
IPTG is an allolactose analogue. Allolactose is the lactose metabolite that triggers the transcription of the lactose operon in natural systems. Allolactose and IPTG activate transcription of the lac operon by binding to the LacI protein. When such binding occurs, the LacI protein changes confirmation, which renders it unable to bind to the operator region. This allows the RNA polymerase to continue reading beyond the operator and transcribe the downstream genes, thus removing the inhibition.
Results
Wet Lab
Different experiments were run using the chemoinduction module.
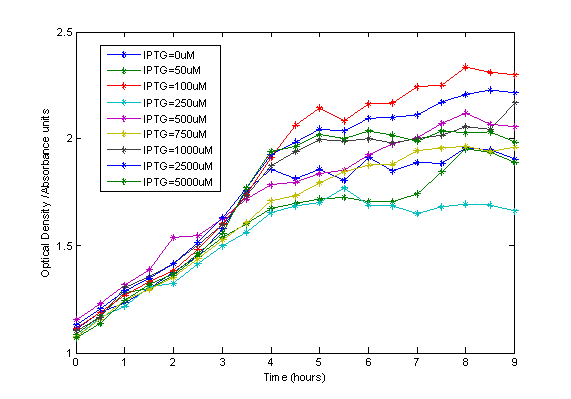
Firstly, we did a preliminary test on the toxicity of IPTG, and whether this affected cell growth. We grew our cells in varying concentrations of IPTG and monitored the growth by measuring optical density.
IPTG, as expected, does not have a negative effect on cell growth.
This provided us with a good starting point for our experiment. We then ligated the IPTG induced promoter to RFP, and tried to investigate the effects of IPTG on red flourescence. Unfortunately, we had problems with getting this construct to function, and we are still troubleshooting as of the wiki-freeze deadline.
Dry Lab
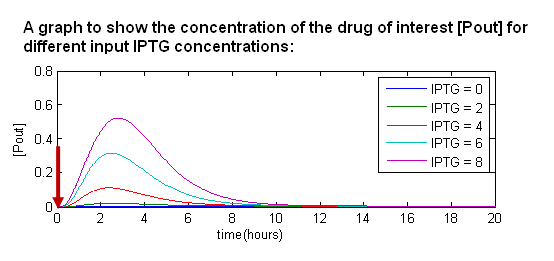
Taking into account the experimental data from the wet lab, we can then draw a relationship between different input IPTG concentrations and the amount of polypeptide produced.
- When we introduce IPTG into the system, it temporarily removes LacI from the system. Hence, during this period of time, we produce the drug of interest.
- When the effects of IPTG wear off, the system returns to equilibrium.
- The more IPTG we add in, the higher the amount of output protein.
Conclusion
Chemoinduction is the most widely used means of inducing a promoter. By using this well characterised IPTG inducible promoter as the starting point for our system, we give the user full control as to when to kick-start the system.
Project Tour


Temporal Control Contents





 "
"