Team:Calgary/4 June 2009
From 2009.igem.org
(Difference between revisions)
| (30 intermediate revisions not shown) | |||
| Line 113: | Line 113: | ||
<br> | <br> | ||
<div class="heading"> | <div class="heading"> | ||
| - | + | Modelling Papers | |
</div> | </div> | ||
<br> | <br> | ||
<div class="desc"> | <div class="desc"> | ||
</html> | </html> | ||
| - | + | The modelling team continued exploring Simbiology and looking at several papers in how genetic modelling can work and what areas should our iGEM team should explore. | |
| + | |||
<html> | <html> | ||
</div> | </div> | ||
</td> | </td> | ||
</tr> | </tr> | ||
| + | |||
| + | |||
<tr> | <tr> | ||
| Line 131: | Line 134: | ||
<tr> | <tr> | ||
<td> | <td> | ||
| - | <a name=" | + | <a name="Emily"></a> |
<br> | <br> | ||
<div class="heading"> | <div class="heading"> | ||
| - | + | EMILY | |
</div> | </div> | ||
<br> | <br> | ||
<div class="heading"> | <div class="heading"> | ||
| - | + | Biobricking LuxOD47A | |
</div> | </div> | ||
<br> | <br> | ||
<div class="desc"> | <div class="desc"> | ||
</html> | </html> | ||
| - | + | *Objective: To form linear copies of LuxOD47A (currently inside TOPO T/A) with BioBrick ends. | |
| - | + | Materials and methods: | |
| - | + | ||
| - | + | ||
| - | + | ||
| + | * Template: LuxOD47E C4-8 in TOPO T/A [40 ng/uL]- original concedntration = [409.5 ng/uL] | ||
| + | * Forward primer: Lux0-F-RS star(Tm = 60 degrees C) | ||
| + | * Reverse primer: LuxO-R-RS primer (Tm = 60 degrees C) | ||
| - | + | Master Mix contents (for 15 tubes) | |
| - | + | * 10X Pfx Amplification buffer (75 uL) | |
| - | + | * 10mM dNTPs mixture (15 uL) | |
| - | + | * 50mM MgSO4 (22.5 uL) | |
| + | * Forward primer [10 uM] (15 uL) | ||
| + | * Reverse primer [10 uM] (15 uL) | ||
| + | * Platinum Pfx DNA polymerase [2.5 units] (10 uL) | ||
| + | * Autoclaved distilled HOH (577.5 uL) | ||
| - | + | * TOTAL: 735 uL | |
| - | + | * PCR reaction volume for Master Mix: 49 uL | |
| - | + | ||
| - | + | PCR steps: | |
| - | + | * Denaturation: 94 degrees C, 5 minutes | |
| - | + | * Amplification: 36 cycles of { | |
| - | + | ** Denaturation (94 degrees C, 15 seconds); | |
| - | + | ** Annealing (58-66 degrees C, 30 seconds); | |
| - | + | ** Extension (68 degrees C, 1.5 minutes)} | |
| - | + | * Final extension: 68 degrees C, 15 minutes | |
| - | + | * Hold temperature: 4 degrees C | |
| - | + | ||
| - | + | 12 tubes were prepared to span the gradient, each of which contained 1 uL of template DNA and 49 uL master mix. A negative control with ddH2O in lieu of template DNA was also prepared. | |
| - | + | ||
| - | + | Results: | |
| + | |||
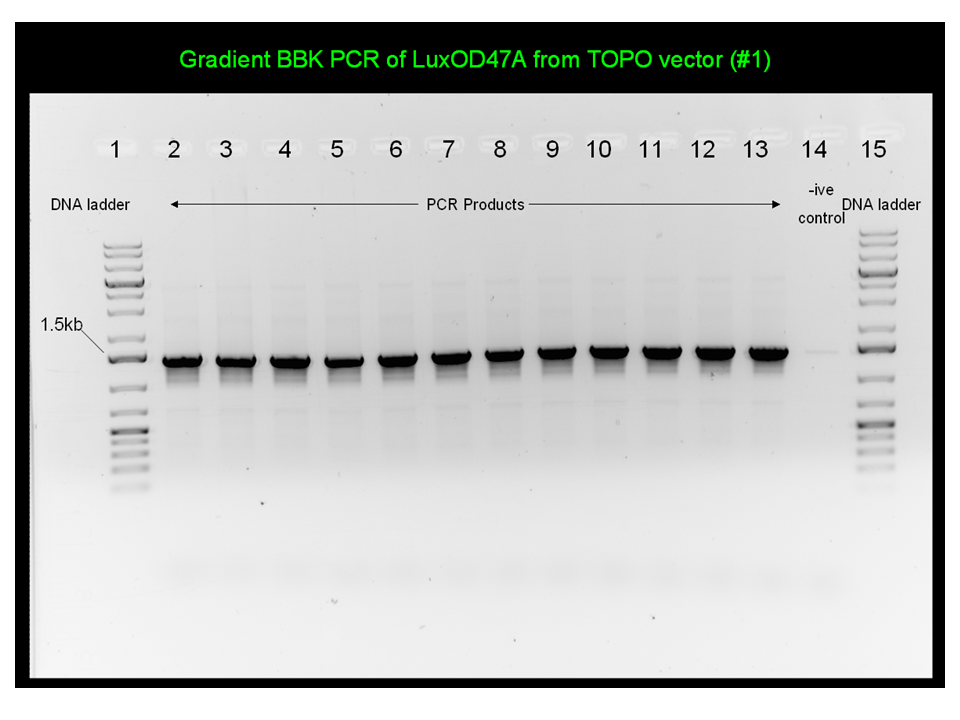
| + | The gradient PCR products were visualised on a 1% agarose gel, as shown below. Lanes 1-12 contain template and labe 13 is a negative control with ddH20 which shows some contamination. The bands are at the right size (~1.4 kb) to suggest that LuxOD47E is in fact in the vector. We will proceed by isolating plasmid and sending it down for DNA sequencing. | ||
| + | |||
| + | [[Image:2009.06.04.LuxOD47E_BBKAmf+PCR2.jpg|350px]] | ||
<html> | <html> | ||
| Line 191: | Line 202: | ||
<br> | <br> | ||
<div class="heading"> | <div class="heading"> | ||
| - | + | Marketing and Outreach for June 4th 2009 | |
</div> | </div> | ||
<br> | <br> | ||
<div class="desc"> | <div class="desc"> | ||
</html> | </html> | ||
| - | + | Today, I continued filling out funding applications for some pharmaceutical companies. Some of these companies include Pfizer Canada, Merck Frosst Canada and Glaxosmithkline Canada. I almost got them completed and once they are done, I will run them through | |
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
<br> | <br> | ||
| - | |||
| - | |||
| - | |||
<br> | <br> | ||
| - | + | I continued working on the Campus Fair agenda as the Campus Fair Day way coming close. | |
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
<html> | <html> | ||
| Line 228: | Line 216: | ||
</td> | </td> | ||
</tr> | </tr> | ||
| + | |||
<tr> | <tr> | ||
| Line 243: | Line 232: | ||
<br> | <br> | ||
<div class="heading"> | <div class="heading"> | ||
| - | + | Gradient PCR amplification <i>luxOU</i> with Biobrick suffix/prefix from TOPO vector | |
</div> | </div> | ||
<br> | <br> | ||
<div class="desc"> | <div class="desc"> | ||
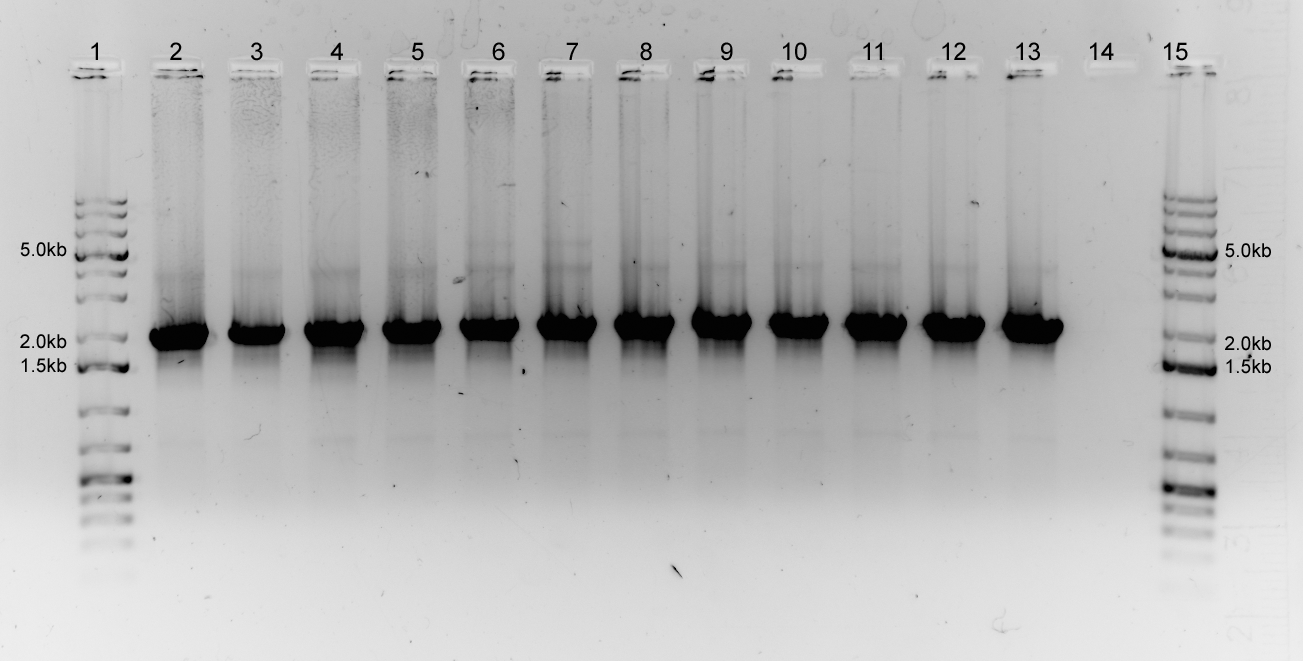
| - | </html> | + | pPFX (Invitrogen, CA) was used to amplify <i>luxOU</i> from the TOPO BL plasmid with luxOU gene-specific primers with the appropriate suffix or prefix (T<sub>m</sub>: 68<sup>o</sup>C). Denaturation at 94<sup>o</sup>C for 5 minutes followed by 36 cycles of 94<sup>o</sup>C for 15sec, 58-66<sup>o</sup>C for 30sec , 68<sup>o</sup>C for 2 minutes. Final extension at 68<sup>o</sup>C for 15 minutes. Visualization of 3uL of product on 1% agarose gel, 120V. |
| - | + | <br /></html> | |
| - | + | <center>[[Image:Calgary_luxOU_GradPCR.jpg | 500px]]</center> | |
| - | <html> | + | <br /><html> |
| + | Figure. 1% agarose gel (100V) for gradient PCR amplification of luxOU with Biobrick suffix and prefix. pPFX amplified 40ng of TOPO BL with luxOU. Gene specific primers with flanking Biobrick restriction sites were used at annealing temperatures ranging from 58oC (left) to 66oC (right). Lanes 2-13 are samples from each reaction carried out at different temperatures. Lane 14 is a negative control. Expected band size is 2kb. 5μL of GeneRuler 1kb Plus DNA Ladder was loaded. | ||
</div> | </div> | ||
| + | <br> | ||
</td> | </td> | ||
</tr> | </tr> | ||
| Line 269: | Line 260: | ||
<br> | <br> | ||
<div class="heading"> | <div class="heading"> | ||
| - | + | Preparation of TOP10 Competent Cells | |
</div> | </div> | ||
<br> | <br> | ||
<div class="desc"> | <div class="desc"> | ||
</html> | </html> | ||
| - | + | In order to replicate plasmid and ensure successful transformation in the summer, today’s work was devoted to making TOP10 competent cells. | |
| + | <br> | ||
| + | To test whether the cells were competent, pBluescript was used as a positive control, and grown on LB-Ampicillin plates. Colonies were seen the next day, verifying that the cells were made competent. | ||
<html> | <html> | ||
| Line 295: | Line 288: | ||
<br> | <br> | ||
<div class="heading"> | <div class="heading"> | ||
| - | + | Improving Usability | |
</div> | </div> | ||
<br> | <br> | ||
<div class="desc"> | <div class="desc"> | ||
</html> | </html> | ||
| - | + | I have found a decent way to ask questions and allow users to answer in a better way then entering things into chat. By using the llDialog function we can also communicate on negative channels, which is not possible in the chat window and helps to divide communication happening between people and between people and objects or objects and other objects. | |
| + | <br> | ||
| + | <br> | ||
| + | So, I will be making everything entered in chat for the PCR machine into dialog prompt where avatars may pick from specific choices. This also gets rid of the risk of spelling errors and increases usability. | ||
<html> | <html> | ||
| Line 333: | Line 329: | ||
</tr> | </tr> | ||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
<tr> | <tr> | ||
| Line 373: | Line 344: | ||
<br> | <br> | ||
<div class="heading"> | <div class="heading"> | ||
| - | + | Early Development | |
</div> | </div> | ||
<br> | <br> | ||
<div class="desc"> | <div class="desc"> | ||
</html> | </html> | ||
| - | + | Retrospective Notebook: This entry was not written on this day, but derived later from working notes I made that day. | |
| + | |||
| + | First began charting out the differences between the types of biobrick part that I wanted to support. Initially thought each type would get its own type of (ie a Promoter Script existing only in promoters), but later resolved to create a single script for to run all of the types for maintainability; and to specify the part's type in the same way internal name and next part in the sequence's names are specified. | ||
| + | |||
| + | Ran some more tests of how the messaging system works in SL, and created six different versions of the device over the course of the day. I began using version numbers to track how my scripts had changed, and was already up to versions 4 and 3 for some scripts, things were changing very quickly early on. | ||
<html> | <html> | ||
| Line 399: | Line 374: | ||
<br> | <br> | ||
<div class="heading"> | <div class="heading"> | ||
| - | + | Marketing and Outreach | |
</div> | </div> | ||
<br> | <br> | ||
<div class="desc"> | <div class="desc"> | ||
</html> | </html> | ||
| - | + | Today, Fahd and I were planning our work for the Campus Fair this Saturday. We decided that we wanted to do the following: | |
| - | + | Pipetting competition - for children aged 8 - 13 using colored dye - gloves, lab coats and goggles were to be used. | |
| - | + | Bacteria crafts - design your own bacteria using foam and pipe cleaners and glitter - for children aged 3 - 8 | |
| - | + | Draw - pretending tube caps were bacteria and people had to guess how many bacteria are in the bag. The answer was 500! | |
| - | + | ||
| - | + | I followed up and emailed almost all the companies that I was in charge of contacting. I haven't heard back from most of them but I'll continue to contact them. | |
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
<html> | <html> | ||
| Line 451: | Line 406: | ||
<br> | <br> | ||
<div class="heading"> | <div class="heading"> | ||
| - | + | Gradient PCR for biobrick cloning of LuxOD47A pieces | |
</div> | </div> | ||
<br> | <br> | ||
<div class="desc"> | <div class="desc"> | ||
</html> | </html> | ||
| - | + | Note: I have switched with Jeremy: I will now be dealing with LuxOD47A and he will be handling LuxPQ | |
| + | |||
| + | Purpose: To form linear copies of LuxOD47A (currently inside TOPO T/A) with BioBrick ends. | ||
| + | |||
| + | Materials and methods: | ||
| + | |||
| + | * Template: LuxOD47A in TOPO T/A [40 ng/uL] | ||
| + | * Forward primer: LuxOD47A BBk forward primer (Tm = 60 degrees C) | ||
| + | * Reverse primer: LuxOD47A BBk reverse primer (Tm = 60 degrees C) | ||
| + | |||
| + | Master Mix contents (for 15 tubes) | ||
| + | * 10X Pfx Amplification buffer (75 uL) | ||
| + | * 10mM dNTPs mixture (15 uL) | ||
| + | * 50mM MgSO4 (22.5 uL) | ||
| + | * Forward primer [10 uM] (15 uL) | ||
| + | * Reverse primer [10 uM] (15 uL) | ||
| + | * Platinum Pfx DNA polymerase [2.5 units] (10 uL) | ||
| + | * Autoclaved distilled HOH (577.5 uL) | ||
| + | |||
| + | * TOTAL: 735 uL | ||
| + | * PCR reaction volume for Master Mix: 49 uL | ||
| + | |||
| + | PCR steps: | ||
| + | * Denaturation: 94 degrees C, 5 minutes | ||
| + | * Amplification: 36 cycles of { | ||
| + | ** Denaturation (94 degrees C, 15 seconds); | ||
| + | ** Annealing (58-66 degrees C, 30 seconds); | ||
| + | ** Extension (68 degrees C, 1.5 minutes)} | ||
| + | * Final extension: 68 degrees C, 15 minutes | ||
| + | * Hold temperature: 4 degrees C | ||
| + | |||
| + | 12 tubes were prepared to span the gradient, each of which contained 1 uL of template DNA and 49 uL master mix. A negative control with ddHOH in lieu of template DNA was also prepared. | ||
| + | |||
| + | Results: | ||
| + | |||
| + | The gradient PCR products were visualised on a 1% agarose gel, as shown below. The right-most lane is the negative control, although it can’t really be distinguished in the results. This doesn’t seem to have worked, so it will be repeated later. | ||
| + | <br><br> | ||
| + | <center> | ||
| + | [[Image:June4.png|700px]] | ||
| + | </center> | ||
| + | |||
<html> | <html> | ||
</div> | </div> | ||
Latest revision as of 04:01, 19 October 2009
UNIVERSITY OF CALGARY

 "
"