|
|
| (12 intermediate revisions not shown) |
| Line 242: |
Line 242: |
| | <br> | | <br> |
| | <div class="heading"> | | <div class="heading"> |
| - | Descriptive Title of What You're Doing
| + | Marketing and Ethics for July 31st 2009 |
| | </div> | | </div> |
| | <br> | | <br> |
| | <div class="desc"> | | <div class="desc"> |
| | </html> | | </html> |
| - | WIKI CODING HERE
| + | Today I worked more on the ethcial aspect of our project. I read more papers on the environmental and economical issues of our team calgary project. |
| | + | <br> |
| | + | <br> |
| | + | I also made follow up calls to a couple of comapnies. |
| | | | |
| | <html> | | <html> |
| Line 268: |
Line 271: |
| | <br> | | <br> |
| | <div class="heading"> | | <div class="heading"> |
| - | Descriptive Title of What You're Doing
| + | More Educational Aspects: Rules Animations |
| | </div> | | </div> |
| | <br> | | <br> |
| | <div class="desc"> | | <div class="desc"> |
| | </html> | | </html> |
| - | WIKI CODING HERE
| + | The next step to enrich the educational aspects of our model is to animate each rule. With doing so, biologists can make connection with the rules as well as the notations used to implement these rules. It is really important for us that make our model convenient to understand for biologists as they are the main users of it. that is why we have designed these animation so that biology and computer science could be connected. |
| | + | |
| | + | [[Image:Rules2.png]] |
| | + | |
| | + | That is what I have been working on this week. |
| | + | |
| | + | In addition we had our meeting, and here is a summary for modeling team: |
| | | | |
| | + | The team members involved in modelling need to meet more and learn more about what the others are doing. We will address this by meeting on Tuesdays and Thursdays at 10h00 am. |
| | + | On the differential side, we have a lot of breadth to our approach but it's still fairly superficial. We have established a differential reaction-based model on SimBiology, but finding values for phosphorylation constants has been a much greater challenge than we had anticipated. We will need to set up a model anyway and collect some input/output data, and from that we can approximate a set of parameters. It's highly unlikely that they'll be unique, but they will give us something to work with and help us evaluate which ones are limiting steps and would be worth tweaking in a real system |
| | + | The point of standardisation and characterisation is to have a set of parts to work with and choose from, based on characteristics that can be compared among different parts. Endy's 2006 paper is interesting, but it only presents characterisation for one part. It would be useful to have more parts characterised, especially if they actually work. We are submitting a few parts to the Registry: a signalling circuit, a reporter, mutants - all with various combinations of promoter and terminator sequences. It would be useful to characterise as many of them as possible, even if it's just for static and dynamic respones, so that future users have a quantifiable way of comparing performance. |
| | + | We are testing a signalling-reporter combination, but we don't know how much the reporter will affect our model of the signalling circuit. For example, if we consider response time, we don't know if our circuit response is limited by the signalling circuit, the reporter circuit, or a combination of both. If the circuit response is slowed mostly by the reporter circuit, it would cast doubt on the validity of the model as soon as a different reporter or sexy circuit were implemented. If there is a way to characterise the reporter by itself along with the signalling + reporter (and if we could do the same with the mutants, that would be interesting too), it would help us assess how good our model is for the signalling circuit by itself, and how it will perform when used with a different reporter or sexy circuit at the end. |
| | <html> | | <html> |
| | </div> | | </div> |
| Line 302: |
Line 315: |
| | <li>Vacufuge to concentrate.</li> | | <li>Vacufuge to concentrate.</li> |
| | <li> Verification digest with <i>Xba</i>I and <i>Pst</i>I. Control: cl lambda part from the Registry distribution plate.</li> | | <li> Verification digest with <i>Xba</i>I and <i>Pst</i>I. Control: cl lambda part from the Registry distribution plate.</li> |
| | + | <li>Results: Faint bands were present. Looked into vector characteristics and it turns out it is high-copy number with IPTG induction (Thanks Thane!) |
| | </ul> | | </ul> |
| | | | |
| Line 352: |
Line 366: |
| | <br> | | <br> |
| | <div class="heading"> | | <div class="heading"> |
| - | Descriptive Title of What You're Doing
| + | Modifications to DNA Extraction, Restriction Digest, Instructions and Write-ups |
| | </div> | | </div> |
| | <br> | | <br> |
| | <div class="desc"> | | <div class="desc"> |
| | </html> | | </html> |
| - | WIKI CODING HERE
| + | The movement of the restriction digest tubes have been modified so they are more interactive so the user is asked what piece of equipment is required once all required items have been added in the correct order to each tube. The three pieces of equipment to touch are the water bath, heating block and incubator (for the insert) respectively. I still am required to place my restriction digest instructions into a notecard on the lab table, which I will do next week. |
| | + | <br> |
| | + | <br> |
| | + | I have been able to modify the first five steps in DNA extraction so that the order in which a avatar proceeds is important. Now if the initial step of resuspension has not been accomplished, if you touch the lysis activity you will not be allowed to perform the activity. However, once a step has been unlocked an avatar may return to it at any time if they believe they missed something or just wish to redo a step. Once an avatar reaches the end of the activity a message is sent to all objects, which resets the entire activity. Now I will have to begin incorporating the containers that store: Wash Solution, High Salt Buffer, Isopropanol, and one containing a DNA buffer as well as one more centrifuge step. |
| | + | <br> |
| | + | <br> |
| | + | Some more instruction were added to bacterial transformation so there is a response for everything an avatar does in attempt to complete the activity. For example, if a user drops an item into the quiz’s inventory that is of the same type that has been submitted before it will alert them to the fact that they have answered that set of questions before and will not ask them again. I have been given the task of creating more products for this activity as well so I will have to add to my transformation script so that the plates with growth will give you the proper object to be used within the lab elsewhere. |
| | + | <br> |
| | + | <br> |
| | + | I also wrote an introduction to the virtual lab for our project description for the wiki, which generally explains the lab’s content as well as its purpose for the second life project. |
| | | | |
| | <html> | | <html> |
| Line 383: |
Line 406: |
| | <div class="desc"> | | <div class="desc"> |
| | </html> | | </html> |
| - | In order to verify cPCR, I have once again ran a cPCR of the reporter circuit, <i>Pqrr4</i>+B0034(RBS)+K082003(GFP+LVA) with <i>Pqrr4</i> Forward and Biobrick CP Reverse primers. Of course, pTaq was used to amplify the DNA. The product was then ran on 2% agarose gel at 90V. ----the gel is being run right now, will edit--- | + | In order to verify cPCR, I have once again ran a cPCR of the reporter circuit, <i>Pqrr4</i>+B0034(RBS)+K082003(GFP+LVA) with <i>Pqrr4</i> Forward and Biobrick CP Reverse primers. Of course, pTaq was used to amplify the DNA. The product was then ran on 2% agarose gel at 90V. The following is a picture of the gel.<br><br> |
| - | | + | [[Image:Calgary_2009.07.31.reporter_cPCR.png|700px]] |
| - | <html> | + | <br><br> |
| - | <div class="heading"> | + | As you can see, the bands in most of the colony lanes match the positive control of <i>Pqrr4</i>+B0034, which means that the cells contain those pieces, not the one of interest. It seems that only the colony 9 is between 1000bp and 1500bp, which is what was expected. However, even this band is suspicious. The size of the circuit is about 1043bp, and with about 125bp addition due to Biobrick CP reverse primer annealing outside of the circuit; however, the band on colony 9 lane is at about 1250bp. Another problem can be noticed in the negative control lane, which is another contamination. It is very disappointing to see a whole day of work gone to waste :(! Yet, it is too early to give up hope. I can try to isolate the plasmid of the Colony 9 and perform a Restriction digest on it to see if I still do not see the expected band, which is at this point unlikely. |
| - | Distribution of our baking sale posters | + | <br><br> |
| - | </div> | + | <b>Distribution of our baking sale posters</b><br><br> |
| - | <br> | + | |
| - | <div class="desc"> | + | |
| - | </html>
| + | |
| | Fahd left a stack of Baking sale posters for us to post throughout the building, and they were posted before 11am today. | | Fahd left a stack of Baking sale posters for us to post throughout the building, and they were posted before 11am today. |
| - |
| |
| | <html> | | <html> |
| | </div> | | </div> |
| Line 444: |
Line 463: |
| | <br> | | <br> |
| | <div class="heading"> | | <div class="heading"> |
| - | Descriptive Title of What You're Doing
| + | Automagic Biobricks |
| | </div> | | </div> |
| | <br> | | <br> |
| | <div class="desc"> | | <div class="desc"> |
| | </html> | | </html> |
| - | WIKI CODING HERE
| + | I spent most of today being very productive with the Biobricker portion of the Biobrick Simulator. This is a little user interface upgrade that will permit you to build any biobrick device you please, from a small library of parts implemented in Second Life. In SL terms, the UI is an object owned by the user's avatar, that can be attached to your display. The simulator's UI will have quite a few features to make your digital bioengineering easier, of which the Biobricker is an important one. |
| | + | |
| | + | I finished working on all of the controls for the biobricker, including some scroll buttons for the parts in the device under construction, and the ability to add parts from the UI's inventory, and delete them too. The Biobricker is all done except for one last control... the build button! This single button will be as much work as the rest of the Biobricker put together, though. The task is to create a copy of each individual Biobrick in world, inform each one where it belongs in the completed device, and link them all together. This will involve a lot of communication back and forth between the user interface object and the biobrick objects out in the world. |
| | + | |
| | + | The Biobricker is in a very primitive state right now, with letters standing in for where all the pretty icons will be in the final version. So instead here's a screenshot of the Main Menu, with the link to the Biobricker visible. |
| | + | |
| | + | <center>[[Image:Calgary Biobrick Simulator UI Preview.png|400px]]</center> |
| | + | |
| | + | The UI fits in at the top left corner of the screen. The red letters are buttons, that cause the UI to be redrawn with a new menu for whichever module you've chosen. I hope this gives you an idea of how the UI fits into the overall simulator. |
| | | | |
| | <html> | | <html> |
| Line 501: |
Line 528: |
| | <br> | | <br> |
| | <div class="heading"> | | <div class="heading"> |
| - | Descriptive Title of What You're Doing
| + | Continuing along the path |
| | </div> | | </div> |
| | <br> | | <br> |
| | <div class="desc"> | | <div class="desc"> |
| | </html> | | </html> |
| - | WIKI CODING HERE
| + | Construction of the path is almost complete and finals plans were mapped out but are subject to change to make room for future virtual demonstrations. In order to complete each station I've decided that a Hawaiian Bobtail Squid will be added to each that will tell visitors about what they see. It will preferably be very interactive (ie. move around and talk to you rather than give a notecard) so as to be fun and educational for the junior high and high school audience we are targeting. The squid that was already in the synthetic kingdom was slightly modified in order to match the physical characteristics of the Hawaiian Bobtail Squid. |
| | | | |
| | <html> | | <html> |
| Line 527: |
Line 554: |
| | <br> | | <br> |
| | <div class="heading"> | | <div class="heading"> |
| - | Descriptive Title of What You're Doing
| + | Discussion write-up, referencing and wiki posting |
| | </div> | | </div> |
| | <br> | | <br> |
| | <div class="desc"> | | <div class="desc"> |
| | </html> | | </html> |
| - | WIKI CODING HERE
| + | (This isn't terribly exciting. I continued with the discussion section of the technical report, I learned all about APA referencing, and I posted part of my notebook on the wiki). |
| | + | |
| | + | <html> |
| | + | </div> |
| | + | <div class=”heading”> |
| | + | Lab meeting |
| | + | </div> |
| | + | <br> |
| | + | <div class=”desc”> |
| | + | </html> |
| | + | |
| | + | We discussed characterisation elements for our signalling circuit, as described in Carol's section. I focussed on dynamic performance and response time, or, in other words, what happens when you add a lot of AI-2 into a system where it wasn't there previously. |
| | + | |
| | + | We also discussed how we could consolidate our modelling efforts and come up with something with a little depth to it in time for the competition. Some of the main points that came out of that are: |
| | + | *The team members involved in modelling need to meet more and learn more about what the others are doing. We will address this by meeting on Tuesdays and Thursdays at 10h00 am. |
| | + | *On the differential side, we have a lot of breadth to our approach but it's still fairly superficial. We have established a differential reaction-based model on SimBiology, but finding values for phosphorylation constants has been a much greater challenge than we had anticipated. We will need to set up a model anyway and collect some input/output data, and from that we can approximate a set of parameters. It's highly unlikely that they'll be unique, but they will give us something to work with and help us evaluate which ones are limiting steps and would be worth tweaking in a real system |
| | + | *The point of standardisation and characterisation is to have a set of parts to work with and choose from, based on characteristics that can be compared among different parts. Endy's 2006 paper is interesting, but it only presents characterisation for one part. It would be useful to have more parts characterised, especially if they actually work. We are submitting a few parts to the Registry: a signalling circuit, a reporter, mutants - all with various combinations of promoter and terminator sequences. It would be useful to characterise as many of them as possible, even if it's just for static and dynamic respones, so that future users have a quantifiable way of comparing performance. |
| | + | *We are testing a signalling-reporter combination, but we don't know how much the reporter will affect our model of the signalling circuit. For example, if we consider response time, we don't know if our circuit response is limited by the signalling circuit, the reporter circuit, or a combination of both. If the circuit response is slowed mostly by the reporter circuit, it would cast doubt on the validity of the model as soon as a different reporter or sexy circuit were implemented. If there is a way to characterise the reporter by itself along with the signalling + reporter (and if we could do the same with the mutants, that would be interesting too), it would help us assess how good our model is for the signalling circuit by itself, and how it will perform when used with a different reporter or sexy circuit at the end. |
| | + | |
| | + | |
| | | | |
| | <html> | | <html> |
|
|
CAROL
Transformation of R0040 (promoter) +pCS26 (vector) into XL Gold cells
- From the overnight digest of R0040 with XhoI and BamHI, the promoter was ligated with vector, pCS26.
- These plasmids were transformed into XL Gold cells and was plated on Kanamycin plates overnight.
Team meeting:
- Lab:discussed about promoter library and the failure to see the expression of luciferase. Also, discussed about the primer design of R0040, so that we can clone the constitutively ON promoter in front of luciferase. This will show us what the luciferase expression will look like.
- Modelling: Discussed about Robustness. The three things that we decided to look at for robustness is: 1. PQ expression (the selection of the 'best' sigma 70 promoter) 2. Temperature Alterations (to see if changes in temperature (from 37oC to another temperature would affect expression) 3. Type of media (LB vs. LM)
|
|
|
CHINMOYEE
Descriptive Title of What You're Doing
|
|
|
EMILY
Sequencing of J13002-LuxOD47E-B0015 Construct in psB1AC3
- Sent Colony 3 of my J13002-LuxOD47E-B0015 construct down for sequencing, results will come in on Monday hopefully.
- Explored Secondlife and met with Rob from UChicago.
- Prepared the Human Practices project write-up for the Wiki.
- Team Meeting- Talked about the exploration of seminars in Second Life and what ideas that has given us. Talked about the pending completion of the J13002-LuxOD47E-B0015 curcuit, results will come in on Monday.
|
|
|
FAHD
Marketing and Ethics for July 31st 2009
Today I worked more on the ethcial aspect of our project. I read more papers on the environmental and economical issues of our team calgary project.
I also made follow up calls to a couple of comapnies.
|
|
|
IMAN
More Educational Aspects: Rules Animations
The next step to enrich the educational aspects of our model is to animate each rule. With doing so, biologists can make connection with the rules as well as the notations used to implement these rules. It is really important for us that make our model convenient to understand for biologists as they are the main users of it. that is why we have designed these animation so that biology and computer science could be connected.
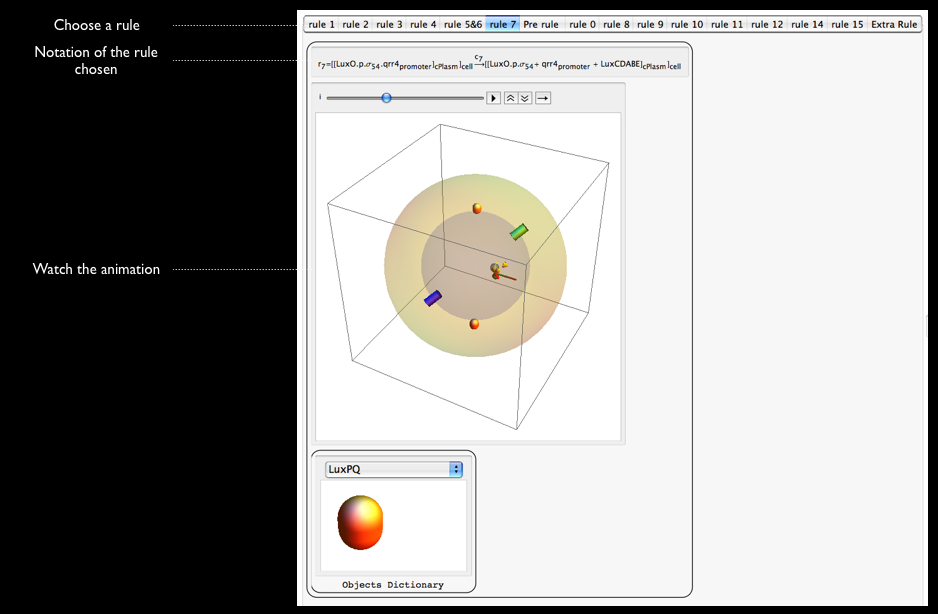
That is what I have been working on this week.
In addition we had our meeting, and here is a summary for modeling team:
The team members involved in modelling need to meet more and learn more about what the others are doing. We will address this by meeting on Tuesdays and Thursdays at 10h00 am.
On the differential side, we have a lot of breadth to our approach but it's still fairly superficial. We have established a differential reaction-based model on SimBiology, but finding values for phosphorylation constants has been a much greater challenge than we had anticipated. We will need to set up a model anyway and collect some input/output data, and from that we can approximate a set of parameters. It's highly unlikely that they'll be unique, but they will give us something to work with and help us evaluate which ones are limiting steps and would be worth tweaking in a real system
The point of standardisation and characterisation is to have a set of parts to work with and choose from, based on characteristics that can be compared among different parts. Endy's 2006 paper is interesting, but it only presents characterisation for one part. It would be useful to have more parts characterised, especially if they actually work. We are submitting a few parts to the Registry: a signalling circuit, a reporter, mutants - all with various combinations of promoter and terminator sequences. It would be useful to characterise as many of them as possible, even if it's just for static and dynamic respones, so that future users have a quantifiable way of comparing performance.
We are testing a signalling-reporter combination, but we don't know how much the reporter will affect our model of the signalling circuit. For example, if we consider response time, we don't know if our circuit response is limited by the signalling circuit, the reporter circuit, or a combination of both. If the circuit response is slowed mostly by the reporter circuit, it would cast doubt on the validity of the model as soon as a different reporter or sexy circuit were implemented. If there is a way to characterise the reporter by itself along with the signalling + reporter (and if we could do the same with the mutants, that would be interesting too), it would help us assess how good our model is for the signalling circuit by itself, and how it will perform when used with a different reporter or sexy circuit at the end.
|
|
|
JAMIE
Verification of cl lambda inverter in psB2K3
- Plasmid miniprep of overnight cultures with Sigma Genelute Miniprep kit. Standard manufacturer protocol used; elution in 40µL of ddH2O.
- Vacufuge to concentrate.
- Verification digest with XbaI and PstI. Control: cl lambda part from the Registry distribution plate.
- Results: Faint bands were present. Looked into vector characteristics and it turns out it is high-copy number with IPTG induction (Thanks Thane!)
|
|
|
JEREMY
Isolation of cl lambda; Sequencing for LuxPQ-LuxOU construct
Six overnight cultures of cl lambda colony 1 were prepared with LB-Kanamycin broth. Plasmid was then isolated using the Sigma GenElute MiniPrep kit. Two plasmids were not successfully isolated, but the other four tubes were pooled into two tubes and vacufuged to increase their concentration. A restriction digest with XbaI and PstI was then set up for 2 hours at 37ºC.
The LuxPQ-B-R-LuxOU-B construct in psB1AC3 was sent for sequencing today based on verification techniques performed over the past few days. Primers used include BBK CP F/R and R0040 Reverse.
|
|
|
KATIE
Modifications to DNA Extraction, Restriction Digest, Instructions and Write-ups
The movement of the restriction digest tubes have been modified so they are more interactive so the user is asked what piece of equipment is required once all required items have been added in the correct order to each tube. The three pieces of equipment to touch are the water bath, heating block and incubator (for the insert) respectively. I still am required to place my restriction digest instructions into a notecard on the lab table, which I will do next week.
I have been able to modify the first five steps in DNA extraction so that the order in which a avatar proceeds is important. Now if the initial step of resuspension has not been accomplished, if you touch the lysis activity you will not be allowed to perform the activity. However, once a step has been unlocked an avatar may return to it at any time if they believe they missed something or just wish to redo a step. Once an avatar reaches the end of the activity a message is sent to all objects, which resets the entire activity. Now I will have to begin incorporating the containers that store: Wash Solution, High Salt Buffer, Isopropanol, and one containing a DNA buffer as well as one more centrifuge step.
Some more instruction were added to bacterial transformation so there is a response for everything an avatar does in attempt to complete the activity. For example, if a user drops an item into the quiz’s inventory that is of the same type that has been submitted before it will alert them to the fact that they have answered that set of questions before and will not ask them again. I have been given the task of creating more products for this activity as well so I will have to add to my transformation script so that the plates with growth will give you the proper object to be used within the lab elsewhere.
I also wrote an introduction to the virtual lab for our project description for the wiki, which generally explains the lab’s content as well as its purpose for the second life project.
|
|
|
KEVIN
cPCR of the reporter circuit
In order to verify cPCR, I have once again ran a cPCR of the reporter circuit, Pqrr4+B0034(RBS)+K082003(GFP+LVA) with Pqrr4 Forward and Biobrick CP Reverse primers. Of course, pTaq was used to amplify the DNA. The product was then ran on 2% agarose gel at 90V. The following is a picture of the gel.

As you can see, the bands in most of the colony lanes match the positive control of Pqrr4+B0034, which means that the cells contain those pieces, not the one of interest. It seems that only the colony 9 is between 1000bp and 1500bp, which is what was expected. However, even this band is suspicious. The size of the circuit is about 1043bp, and with about 125bp addition due to Biobrick CP reverse primer annealing outside of the circuit; however, the band on colony 9 lane is at about 1250bp. Another problem can be noticed in the negative control lane, which is another contamination. It is very disappointing to see a whole day of work gone to waste :(! Yet, it is too early to give up hope. I can try to isolate the plasmid of the Colony 9 and perform a Restriction digest on it to see if I still do not see the expected band, which is at this point unlikely.
Distribution of our baking sale posters
Fahd left a stack of Baking sale posters for us to post throughout the building, and they were posted before 11am today.
|
|
|
MANDY
Descriptive Title of What You're Doing
|
|
|
PATRICK
Automagic Biobricks
I spent most of today being very productive with the Biobricker portion of the Biobrick Simulator. This is a little user interface upgrade that will permit you to build any biobrick device you please, from a small library of parts implemented in Second Life. In SL terms, the UI is an object owned by the user's avatar, that can be attached to your display. The simulator's UI will have quite a few features to make your digital bioengineering easier, of which the Biobricker is an important one.
I finished working on all of the controls for the biobricker, including some scroll buttons for the parts in the device under construction, and the ability to add parts from the UI's inventory, and delete them too. The Biobricker is all done except for one last control... the build button! This single button will be as much work as the rest of the Biobricker put together, though. The task is to create a copy of each individual Biobrick in world, inform each one where it belongs in the completed device, and link them all together. This will involve a lot of communication back and forth between the user interface object and the biobrick objects out in the world.
The Biobricker is in a very primitive state right now, with letters standing in for where all the pretty icons will be in the final version. So instead here's a screenshot of the Main Menu, with the link to the Biobricker visible.

The UI fits in at the top left corner of the screen. The red letters are buttons, that cause the UI to be redrawn with a new menu for whichever module you've chosen. I hope this gives you an idea of how the UI fits into the overall simulator.
|
|
|
PRIMA
Continuation of last day's work
Today, I edited the high school presentation proposition again and sent it out to Mrs. Ellechuk. I filled in the lab notebook and filled out the daily updates on the wiki.
I finished typing out the acknowledgement for each mentor/advisor of iGEM Calgary. I also helped to write references for Sonja. Then I finished off with mailing out more newsletters. We also had a very productive team meeting today.
On Monday I have to follow up with the companies which I have been contacting for the past few days. I will also call our hotel in Boston to change a few rooms around.
|
|
|
STEFAN
Continuing along the path
Construction of the path is almost complete and finals plans were mapped out but are subject to change to make room for future virtual demonstrations. In order to complete each station I've decided that a Hawaiian Bobtail Squid will be added to each that will tell visitors about what they see. It will preferably be very interactive (ie. move around and talk to you rather than give a notecard) so as to be fun and educational for the junior high and high school audience we are targeting. The squid that was already in the synthetic kingdom was slightly modified in order to match the physical characteristics of the Hawaiian Bobtail Squid.
|
|
|
VICKI
Discussion write-up, referencing and wiki posting
(This isn't terribly exciting. I continued with the discussion section of the technical report, I learned all about APA referencing, and I posted part of my notebook on the wiki).
Lab meeting
We discussed characterisation elements for our signalling circuit, as described in Carol's section. I focussed on dynamic performance and response time, or, in other words, what happens when you add a lot of AI-2 into a system where it wasn't there previously.
We also discussed how we could consolidate our modelling efforts and come up with something with a little depth to it in time for the competition. Some of the main points that came out of that are:
- The team members involved in modelling need to meet more and learn more about what the others are doing. We will address this by meeting on Tuesdays and Thursdays at 10h00 am.
- On the differential side, we have a lot of breadth to our approach but it's still fairly superficial. We have established a differential reaction-based model on SimBiology, but finding values for phosphorylation constants has been a much greater challenge than we had anticipated. We will need to set up a model anyway and collect some input/output data, and from that we can approximate a set of parameters. It's highly unlikely that they'll be unique, but they will give us something to work with and help us evaluate which ones are limiting steps and would be worth tweaking in a real system
- The point of standardisation and characterisation is to have a set of parts to work with and choose from, based on characteristics that can be compared among different parts. Endy's 2006 paper is interesting, but it only presents characterisation for one part. It would be useful to have more parts characterised, especially if they actually work. We are submitting a few parts to the Registry: a signalling circuit, a reporter, mutants - all with various combinations of promoter and terminator sequences. It would be useful to characterise as many of them as possible, even if it's just for static and dynamic respones, so that future users have a quantifiable way of comparing performance.
- We are testing a signalling-reporter combination, but we don't know how much the reporter will affect our model of the signalling circuit. For example, if we consider response time, we don't know if our circuit response is limited by the signalling circuit, the reporter circuit, or a combination of both. If the circuit response is slowed mostly by the reporter circuit, it would cast doubt on the validity of the model as soon as a different reporter or sexy circuit were implemented. If there is a way to characterise the reporter by itself along with the signalling + reporter (and if we could do the same with the mutants, that would be interesting too), it would help us assess how good our model is for the signalling circuit by itself, and how it will perform when used with a different reporter or sexy circuit at the end.
|

 "
"













