Team:Calgary/9 July 2009
From 2009.igem.org
(Difference between revisions)
Emily Hicks (Talk | contribs) |
|||
| (14 intermediate revisions not shown) | |||
| Line 157: | Line 157: | ||
<br> | <br> | ||
<div class="heading"> | <div class="heading"> | ||
| - | + | Colony PCR | |
</div> | </div> | ||
<br> | <br> | ||
<div class="desc"> | <div class="desc"> | ||
</html> | </html> | ||
| - | + | * Overnight plates resulted in 3 colonies | |
| + | * Performed colony PCR on the three colonies using BBK CP reverse and forward primers. | ||
| + | * The 0.6% agarose gel showed no viable results. Other smaller bands are found on the gel which suggests that we may be cloning primer dimers into the plasmid. | ||
| + | * Might have to attempt gel extraction. | ||
| + | |||
<html> | <html> | ||
</div> | </div> | ||
| Line 189: | Line 193: | ||
Today I went to the modelling meeting in ICT building . Then I decided to make a summary about math modelling so that on Friday both the modelling teams could compare the two system . This will ensure uniformity in parameter values , reaction equations used etc. Also a good written record will be present. | Today I went to the modelling meeting in ICT building . Then I decided to make a summary about math modelling so that on Friday both the modelling teams could compare the two system . This will ensure uniformity in parameter values , reaction equations used etc. Also a good written record will be present. | ||
| - | |||
<html> | <html> | ||
</div> | </div> | ||
| Line 210: | Line 213: | ||
<br> | <br> | ||
<div class="heading"> | <div class="heading"> | ||
| - | + | J13002-LuxOD47E-B0015 Construction Take Two | |
</div> | </div> | ||
<br> | <br> | ||
<div class="desc"> | <div class="desc"> | ||
</html> | </html> | ||
| - | *Today I ran a colony PCR of my J13002-LuxOD47E-B0015 colonies to verify the presence of the B0015 terminator in my construct. BBK-CP forward and reverse primers were used with p Taq and cycling conditions were as follows: 94°C for 6 min, 36x(94°C for 30 s, 55°C for 45s, 72°C for 90s), | + | *Today sequencing results came back and I was in fact successful in cloning in the J13002 promoter! So now I can proceed to verification of my J13002-LuxOD47E-B0015 circuit. |
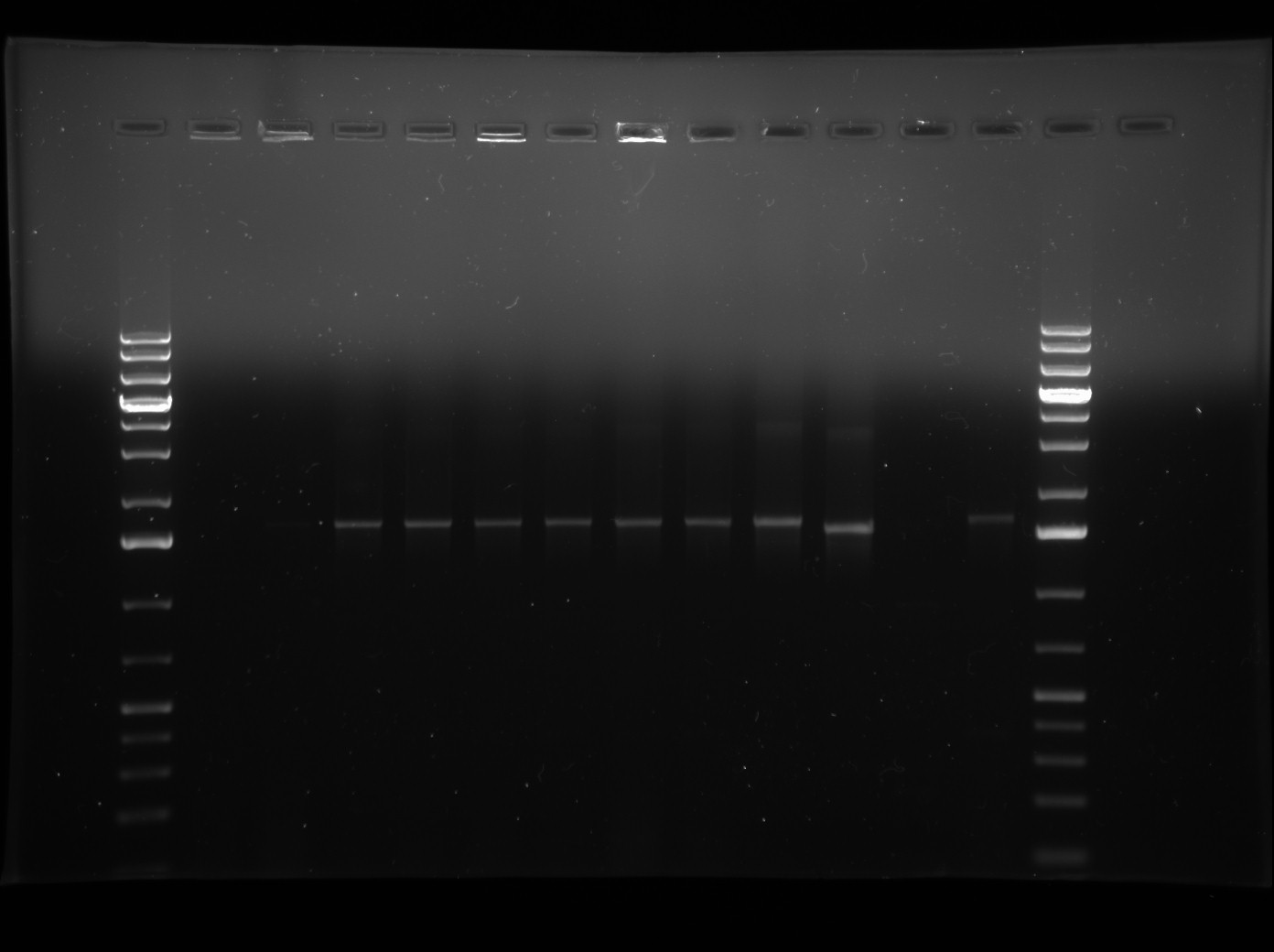
| + | *So I ran a colony PCR of my J13002-LuxOD47E-B0015 colonies to verify the presence of the B0015 terminator in my construct. BBK-CP forward and reverse primers were used with p Taq and cycling conditions were as follows: 94°C for 6 min, 36x(94°C for 30 s, 55°C for 45s, 72°C for 90s), 72°C for 10 min, held at 4°C. PCR products were visualized on a 1% agarose gel run at 120 V with J13002-LuxOD47E and LuxOD47E as size controls. Gel is pictured below. Lanes 1-4 are trial one where the B0015 terminator was treated as the insert. Lanes 5-8 are trial 2 where the J13002-LuxOD47E construct was treated as the insert. Lane 9 is J13002-LuxOD47E and Lane 10 is LuxOD47E. Lane 11 is blank and Lane 12 is the negative control. | ||
| + | [[Image:2009.07.09J13002-LuxOD47E-B0015.jpg|350px]] | ||
*Because all bands in the first 9 lanes appear to be the same size as the band in lane 9 with J13002-LuxOD47E, we can conclude that the B0015 terminator was not successfully cloned in and we will have to go back to contruction again. | *Because all bands in the first 9 lanes appear to be the same size as the band in lane 9 with J13002-LuxOD47E, we can conclude that the B0015 terminator was not successfully cloned in and we will have to go back to contruction again. | ||
| - | *Performed a restriction digest with EcoRI, XbaI and SpeI of B0015 and the J13002-LuxOD47E construct, digesting the insert with EcoRI and SpeI and digesting the recipient with EcoRI and XbaI. Left the | + | *Performed a restriction digest with EcoRI, XbaI and SpeI of B0015 and the J13002-LuxOD47E construct, digesting the insert with EcoRI and SpeI and digesting the recipient with EcoRI and XbaI. Left the digestion in the 37C waterbath for 2 hours, transformed into TOP10 cells on both AC and C plates and left for overnight growth. |
| + | *Today I also checked my KT1144 cells for glow. There wasn't any, but we wil let them grow at room temperature for a couple more days. | ||
<html> | <html> | ||
| Line 241: | Line 247: | ||
<br> | <br> | ||
<div class="heading"> | <div class="heading"> | ||
| - | + | Marketing for July 9th 2009 | |
</div> | </div> | ||
<br> | <br> | ||
<div class="desc"> | <div class="desc"> | ||
</html> | </html> | ||
| - | + | 1) I contacted a number of travel agencies(including Travel Cuts) and the cheapest i got was through travel cuts and that was 595 CDN dollars(includes taxes and this price is with student discounts). However, Prima got a really good deal a few weeks ago( 412 CDN dollars) and i think(suggestion) that we should book these flights before somebody else grabs the booking. I am still waiting for our team members to respond me back with regards to the HOTELS e-mail i had sent out on WEDNESDAY about their preference. SO far only Patrick, Mandy, Stefan and Prima have told me about their preference. | |
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
<br> | <br> | ||
| - | |||
| - | |||
| - | |||
<br> | <br> | ||
| - | + | 2) I did a research/invesyigation on the following research pharmaceutical companies: | |
| - | + | ||
| - | + | ||
<br> | <br> | ||
| - | < | + | <br> |
| - | < | + | a-Ambion Inc. |
| - | + | <br> | |
| + | <br> | ||
| + | b-Charles River Labs | ||
| + | <br> | ||
| + | <br> | ||
| + | c-Critical Outcome Technologies | ||
| + | <br> | ||
| + | <br> | ||
| + | d- E-Z-EM Canada Inc. | ||
| + | <br> | ||
| + | <br> | ||
| + | e- Genome Canada | ||
| + | <br> | ||
| + | <br> | ||
| + | f- i3 Canada Inc. | ||
| + | <br> | ||
| + | <br> | ||
| + | g- Ikaria Canada Inc. | ||
| + | <br> | ||
| + | <br> | ||
| + | h- Inimex Pharmaceuticals | ||
| + | <br> | ||
| + | <br> | ||
| + | i- Arc Pharma Canada Inc. | ||
| + | <br> | ||
| + | <br> | ||
| + | j- Dalto Pharma Services | ||
| + | <br> | ||
| + | <br> | ||
| + | k- Medicago Inc. | ||
| + | <br> | ||
| + | <br> | ||
| + | l- Neuroimage Inc. | ||
| + | <br> | ||
| + | <br> | ||
| + | m- Synbiosis | ||
| + | <br> | ||
| + | <br> | ||
| + | I e-mailed only 4 of the companies above because i want to spend as much time as i can on a company. I also have to confirm them with my team before sending them out so it takes time but i hope they respond back to us in a positive manner. | ||
<html> | <html> | ||
| Line 293: | Line 319: | ||
<br> | <br> | ||
<div class="heading"> | <div class="heading"> | ||
| - | + | Signalling circuit construction and Response circuit preparation | |
</div> | </div> | ||
<br> | <br> | ||
<div class="desc"> | <div class="desc"> | ||
</html> | </html> | ||
| - | + | See Jeremy's updates regarding luxPQ-luxOU construction from hereonin. | |
| + | |||
| + | cIIambda and aiiA were transformed from registry plates into TOP10 cells for plasmid isolation and verification. | ||
<html> | <html> | ||
| Line 319: | Line 347: | ||
<br> | <br> | ||
<div class="heading"> | <div class="heading"> | ||
| - | + | Constructing LuxPQ-LuxOU signaling circuit | |
</div> | </div> | ||
<br> | <br> | ||
<div class="desc"> | <div class="desc"> | ||
</html> | </html> | ||
| - | + | Purpose: to construct LuxPQ-B0015-R0040-LuxOU-B0015 (hereafter PQ-B-R-OU-B) by inserting LuxPQ (AC) into B-R-OU-B (AK). Construction was performed by setting up a restriction digest by cutting the insert (LuxPQ) with EcoRI and SpeI and cutting the vector (B-R-OU-B in AK) with EcoRI and XbaI. After this restriction digest, phosphatase and ligation treatment was performed and the plasmid was subsequently transformed into TOP10 competent cells and plated overnight on LB+Kanamycin. | |
<html> | <html> | ||
| Line 364: | Line 392: | ||
<br> | <br> | ||
<br> | <br> | ||
| - | I was also able to complete the script for the sequencing station as per Mandy’s request and it will now give any avatar who drops ‘Amplified | + | I was also able to complete the script for the sequencing station as per Mandy’s request and it will now give any avatar who drops ‘Amplified Colicin E2’ into the sequencing object a notecard, which as of now does not contain any useful information. I will be adding various other amplified DNA products as objects to give back sequencing for on Friday, which may continue into next week. |
| - | + | ||
<html> | <html> | ||
| Line 398: | Line 425: | ||
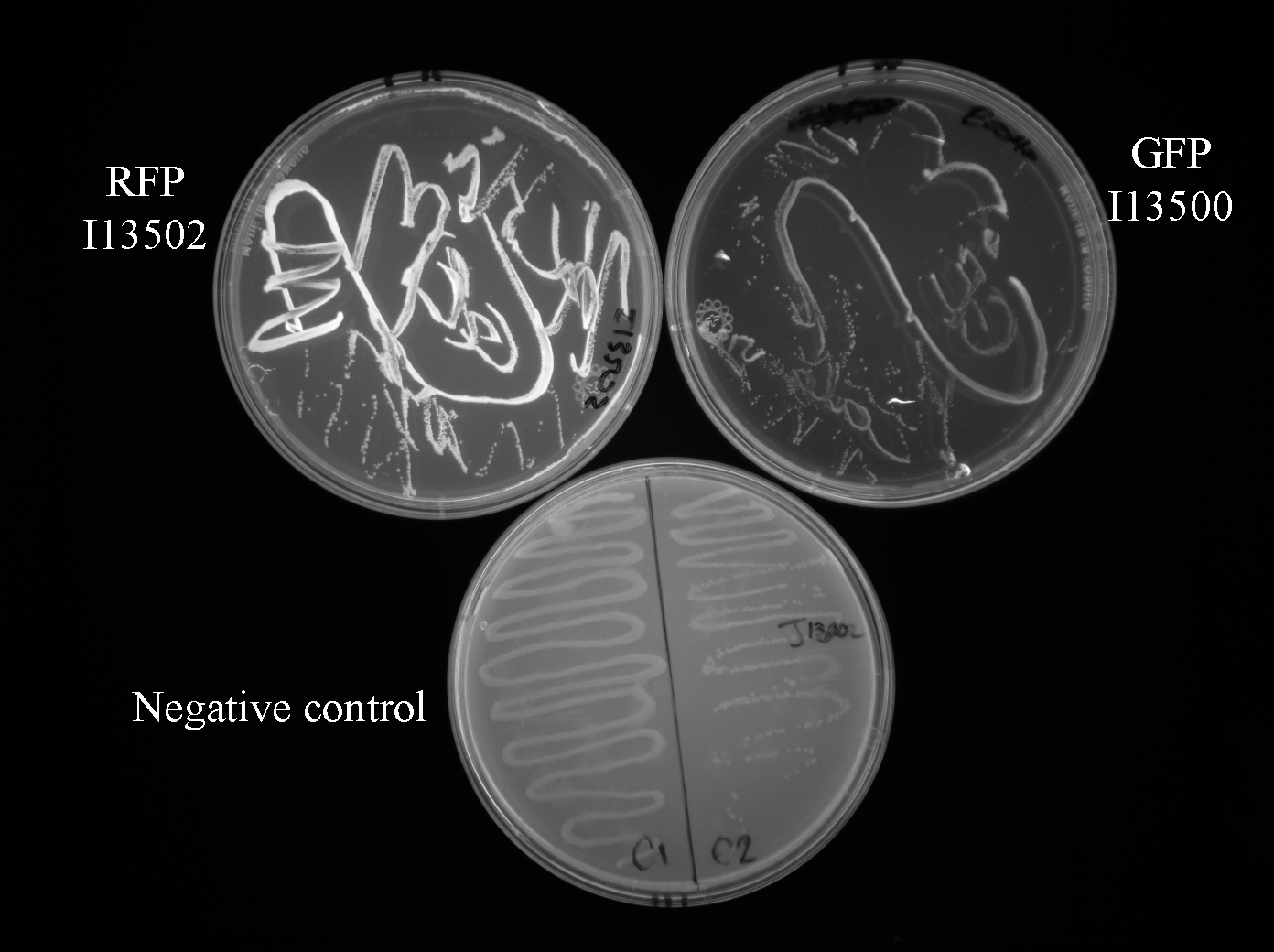
<br><br> | <br><br> | ||
From it, one can clearly see that RFP is glowing much brighter than the negative control, and that GFP is, although less obvious, also glowing. | From it, one can clearly see that RFP is glowing much brighter than the negative control, and that GFP is, although less obvious, also glowing. | ||
| + | |||
No other lab work was performed on this day. | No other lab work was performed on this day. | ||
| + | |||
<html> | <html> | ||
</div> | </div> | ||
| Line 418: | Line 447: | ||
<br> | <br> | ||
<div class="heading"> | <div class="heading"> | ||
| - | + | Second Life Biobert | |
</div> | </div> | ||
<br> | <br> | ||
<div class="desc"> | <div class="desc"> | ||
</html> | </html> | ||
| - | + | Today, we finished the prototype for ‘Biobert’, the lab helper robot. When clicked he can move and chat with people (using their names :D). He can also give instructions, note cards, and inventory like vectors. He also is the point where users can choose ‘lab missions’ to complete. | |
| - | + | <br><br> | |
| + | I finished designing/planning the intermediate lab mission level, which is PCR/Gel electrophoresis verification and then bacterial transformation of a circuit. I started designing the more difficult lab mission that involves the compilation of a biobricked circuit. | ||
| + | <br><br> | ||
| + | Biobert currently only responds to a couple phrases said to him by users. Tomorrow I plan to expand on both what he does and says (integrate him more in the lab, maybe have him give clues, etc.). Right now, he also likes to repeat himself a lot, which can get kind of annoying so I’m going to delete some of the scripting too. | ||
| + | <br><br> | ||
| + | As soon as I learn exactly how Katie’s scripted restriction digest works, I will be finishing the designing/planning of the more difficult lab mission, and then figure out what can be an easy one. | ||
<html> | <html> | ||
</div> | </div> | ||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
<br> | <br> | ||
<div class="heading"> | <div class="heading"> | ||
| - | + | Wiki Reformatting | |
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
</div> | </div> | ||
<br> | <br> | ||
<div class="desc"> | <div class="desc"> | ||
</html> | </html> | ||
| - | WIKI | + | Today, I tried laying out the wiki more, and getting it ready for sponsors to see. NO SUCH LUCK, HOWEVER: here is what I was playing with, NOT on a wiki site: http://igemtest.blogspot.com |
| + | (The purple is still uh…being debated). My css does not seem to combine with the wiki mark up, and Emily found that some teams used ‘templates’ with wiki markup to get around this problem, which I haven’t quite figured out how to do yet, but I’ve bookmarked some pages to read. THE WIKI pages have all been saved on Emily’s laptop and the pages online have been wiped in preparation for some trouble shooting (slowly adding elements and figuring out what isn’t working – why the formatting is off). | ||
| + | <br><br> | ||
| + | THE BLOG is filling up rather nicely, helped Prima upload her video and retagged/formatted the current posts (so they will show up if people search iGEM on blogspot). | ||
<html> | <html> | ||
</div> | </div> | ||
| + | <Br> | ||
</td> | </td> | ||
</tr> | </tr> | ||
| Line 470: | Line 493: | ||
<br> | <br> | ||
<div class="heading"> | <div class="heading"> | ||
| - | + | Company contacts | |
</div> | </div> | ||
<br> | <br> | ||
<div class="desc"> | <div class="desc"> | ||
</html> | </html> | ||
| - | + | I updated the gmail contact list. Sent a Thank you letter to Corning Life Sciences. | |
| + | Called Worley Parsons – the contact they gave me was apparently the wrong person. It was the HR person and she said she’ll find out who deals with sponsorship and get back to me. But I might just inquire and find out myself just in case she forgets. I’ve been calling company 2 for some time now… I’ll call him again tomorrow. | ||
| + | |||
| + | Sigma-Aldrich- called today but couldn’t get a hold of her so I left a message follow up tomorrow | ||
| + | |||
| + | Cayman Chemicals ..called…needs a final confirmation from their President follow up July 31 | ||
| + | |||
| + | Followed up with numerous companies (including oil companies such as Shell and Husky) but they declined our proposal. | ||
| + | I will continue to follow up and contact new companies. | ||
<html> | <html> | ||
| Line 496: | Line 527: | ||
<br> | <br> | ||
<div class="heading"> | <div class="heading"> | ||
| - | + | Freebies | |
</div> | </div> | ||
<br> | <br> | ||
<div class="desc"> | <div class="desc"> | ||
</html> | </html> | ||
| - | + | Most of today was spent traveling islands and the online Second Life marketplace XStreet in search of things to get for our sim. These included: | |
| + | <br> | ||
| + | <br> | ||
| + | *New textures | ||
| + | *Rocks, coral and seaweed | ||
| + | *Walkways | ||
| + | *Particle effects | ||
<html> | <html> | ||
| Line 522: | Line 559: | ||
<br> | <br> | ||
<div class="heading"> | <div class="heading"> | ||
| - | + | Modelling meeting | |
</div> | </div> | ||
<br> | <br> | ||
<div class="desc"> | <div class="desc"> | ||
</html> | </html> | ||
| - | + | Team MatLab met with Team Mathematica in order to better coordinate our efforts and ensure that we have something in place that will have an impact at the jamboree. More comprehensive notes can be found in Chinee’s notebook. We watched a simulation of what Afshin and Iman have come up with for membrane computing and discussed how to go about streamlining our work so that our models will actually have the potential to support each other. This would involve a thorough literature search to find relevant phosphorylation and reaction constants (and an analysis of the inherent assumptions). As well, we agreed to write up a set of rules/reactions so that a team member from the other modelling school of thought (or otherwise) can better follow the work that each modelling crew has done on screen. | |
<html> | <html> | ||
Latest revision as of 21:53, 20 October 2009
UNIVERSITY OF CALGARY

 "
"