|
|
| (10 intermediate revisions not shown) |
| Line 113: |
Line 113: |
| | <br> | | <br> |
| | <div class="heading"> | | <div class="heading"> |
| - | Descriptive Title of What You’re Doing
| + | Continuation of Plasmid Isolation |
| | </div> | | </div> |
| | <br> | | <br> |
| | <div class="desc"> | | <div class="desc"> |
| | </html> | | </html> |
| - | WIKI CODING HERE
| + | * Continued with Maxi-prep protocol via DNA precipitation. Unfortunately, no products were found. The washing step needs to be done with caution and the DNA product might have been discarded as well. |
| - | <html>
| + | * Prepared another overnight culture so that protocol can be repeated. |
| - | </div>
| + | |
| - | </td>
| + | |
| - | </tr>
| + | |
| - | | + | |
| - | <tr>
| + | |
| - | <td height="10px" bgcolor="#222222">
| + | |
| - | </td>
| + | |
| - | </tr>
| + | |
| - | | + | |
| - | <tr>
| + | |
| - | <td>
| + | |
| - | <a name="Chinmoyee"></a>
| + | |
| - | <br>
| + | |
| - | <div class="heading">
| + | |
| - | CHINMOYEE
| + | |
| - | </div>
| + | |
| - | <br>
| + | |
| - | <div class="heading">
| + | |
| - | CLASS
| + | |
| - | </div>
| + | |
| - | <br>
| + | |
| - | <div class="desc">
| + | |
| - | </html>
| + | |
| - | | + | |
| | | | |
| | <html> | | <html> |
| Line 149: |
Line 125: |
| | </td> | | </td> |
| | </tr> | | </tr> |
| - |
| |
| | | | |
| | <tr> | | <tr> |
| Line 174: |
Line 149: |
| | *Followed protocol from June 6th 2009 | | *Followed protocol from June 6th 2009 |
| | *PCR Prodcuts visualized on a 1% agarose gel run at 120 V. See gel photo below. Lanes 1-12 contain mastermix + LuxOD47E in pCR2.1-TOPO vector template. Lane 13 is a negative control with mastermix + ddH2O. | | *PCR Prodcuts visualized on a 1% agarose gel run at 120 V. See gel photo below. Lanes 1-12 contain mastermix + LuxOD47E in pCR2.1-TOPO vector template. Lane 13 is a negative control with mastermix + ddH2O. |
| - | [[Image:2009.06.12.LuxOD47E_Gradient_PCR.tif]] | + | [[Image:2009.06.12.LuxOD47E_Gradient_PCR.jpg|350px]] |
| | *Analysis: There is considerable contamination in the negative control, however the gradient looks as expected. The source of contamination may possibly be from the primers that were shared between Viki and I. We will try again with new Primer stocks diluted by Thane. | | *Analysis: There is considerable contamination in the negative control, however the gradient looks as expected. The source of contamination may possibly be from the primers that were shared between Viki and I. We will try again with new Primer stocks diluted by Thane. |
| | | | |
| Line 196: |
Line 171: |
| | <br> | | <br> |
| | <div class="heading"> | | <div class="heading"> |
| - | Descriptive Title of What You're Doing
| + | Marketing for June 11th 2009 |
| | </div> | | </div> |
| | <br> | | <br> |
| | <div class="desc"> | | <div class="desc"> |
| | </html> | | </html> |
| - | WIKI CODING HERE
| + | Today, we gad a meeting with Derek Gratz who is a University of Calgary Alumni and the owner of WestLink Innovations. He gave us good ideas on how to approach companies and how to important it is to build a network with these companies. He also gave us names of a few potential companies that might be interested in our project |
| - | | + | |
| - | <html>
| + | |
| - | </div>
| + | |
| - | </td>
| + | |
| - | </tr>
| + | |
| - | | + | |
| - | <tr>
| + | |
| - | <td height="10px" bgcolor="#222222">
| + | |
| - | </td>
| + | |
| - | </tr>
| + | |
| - | | + | |
| - | <tr>
| + | |
| - | <td>
| + | |
| - | <a name="Iman"></a>
| + | |
| - | <br>
| + | |
| - | <div class="heading">
| + | |
| - | IMAN
| + | |
| - | </div>
| + | |
| - | <br>
| + | |
| - | <div class="heading">
| + | |
| - | Descriptive Title of What You're Doing
| + | |
| - | </div>
| + | |
| - | <br>
| + | |
| - | <div class="desc">
| + | |
| - | </html>
| + | |
| - | WIKI CODING HERE
| + | |
| | | | |
| | <html> | | <html> |
| Line 248: |
Line 197: |
| | <br> | | <br> |
| | <div class="heading"> | | <div class="heading"> |
| - | Descriptive Title of What You're Doing
| + | Recheck Construction Technique for luxOU |
| | </div> | | </div> |
| | <br> | | <br> |
| | <div class="desc"> | | <div class="desc"> |
| | </html> | | </html> |
| - | WIKI CODING HERE
| + | No colonies grew on LB+chloramphenicol35 plates. Possible check points: construction digest and ligation/transformation. |
| | + | |
| | + | 1% agarose gel of construction digested vector showed cutting. Redo transformation with positive control |
| | | | |
| | <html> | | <html> |
| Line 274: |
Line 225: |
| | <br> | | <br> |
| | <div class="heading"> | | <div class="heading"> |
| - | Descriptive Title of What You're Doing
| + | Amplification of luxPQ with BBK using Gradient PCR |
| | </div> | | </div> |
| | <br> | | <br> |
| | <div class="desc"> | | <div class="desc"> |
| | </html> | | </html> |
| - | WIKI CODING HERE
| + | The purpose of today’s work was to begin moving the luxPQ operon from the TOPO vector to a BBK vector. luxPQ C24-1 TOPO was used as the DNA template, and in order to flank luxPQ with the appropriate BBK Restriction sites, LuxPQ-RS-F/R primers were used. A platinum-PFX gradient PCR was set up with the following conditions: 94ºC for 5 minutes; 36 X (94ºC for 30 seconds; 58-66ºC for 30 seconds; 72ºC for 4 minutes); 72ºC for 10 minutes; held at 4ºC. The extension, however, should have been set at 68ºC and not 72ºC as per the recommendation of the manufacturer. This PCR ran overnight. |
| | | | |
| | <html> | | <html> |
| Line 356: |
Line 307: |
| | <br> | | <br> |
| | <div class="heading"> | | <div class="heading"> |
| - | Descriptive Title of What You're Doing
| + | Gel Making in Second Life |
| | </div> | | </div> |
| | <br> | | <br> |
| | <div class="desc"> | | <div class="desc"> |
| | </html> | | </html> |
| - | WIKI CODING HERE
| + | With some inspiration from Sigma Aldrich, I began building a fumehood for the lab. As well, I read up and began writing a notecard describing the preparation of gels for gel electrophoresis. This is just for additional information (just like plate preparation), we are not planning on making this an activity for the virtual lab. |
| - | | + | |
| | <html> | | <html> |
| | </div> | | </div> |
| | + | <br> |
| | </td> | | </td> |
| | </tr> | | </tr> |
| Line 382: |
Line 333: |
| | <br> | | <br> |
| | <div class="heading"> | | <div class="heading"> |
| - | Descriptive Title of What You're Doing
| + | Physical Objects and Nonphysical Objects Do Not Get Along |
| | </div> | | </div> |
| | <br> | | <br> |
| | <div class="desc"> | | <div class="desc"> |
| | </html> | | </html> |
| - | WIKI CODING HERE
| + | Retrospective Notebook: This entry was not written on this day, but derived later from working notes I made that day. |
| | + | |
| | + | Began setting up a workaround for the problems with using SL's 'collision' event for object messaging. |
| | + | |
| | + | First started changing the object movement script to toggle its 'physics' state on and off, instead of continuously setting its velocity to zero, which would be a much smoother and simpler solution to that problem. |
| | + | |
| | + | I had by this time built a working RNAP, insofar as it would bind at the promoter, follow the DNA and detach at a terminator. |
| | + | |
| | + | I also encountered a problem with the RNAP polymerase and the DNA in collision, where because the DNA object was physical (ie, subject to the physics engine in second life) but the RNAP was not physical (so that it could occupy the same space as the DNA without colliding with it), the RNAP would try to move along the strand of DNA, but wound up 'pushing' the movable DNA along. If left alone, this setup would continue crawling across the world indefinitely! Ultimately, when two objects have to live in close proximity with each other, they must both be set to nonphysical or they will exhibit collision glitches like this. |
| | | | |
| | <html> | | <html> |
| Line 408: |
Line 367: |
| | <br> | | <br> |
| | <div class="heading"> | | <div class="heading"> |
| - | Descriptive Title of What You're Doing
| + | WestLink Innovations Meeting |
| | </div> | | </div> |
| | <br> | | <br> |
| | <div class="desc"> | | <div class="desc"> |
| | </html> | | </html> |
| - | WIKI CODING HERE
| + | Fahd, Thane and I went to meet the President from WestLink Innovations.He offered advise on how to contact companies, what to say, what not to say, who we should contact. He offered to help us with networking. It was nice of him to take time out of work to meet and discuss marketing strategies with us. |
| | + | What I got out of this: |
| | + | 1) Contact information for other companies |
| | + | 2) names to new companies which I hadn't thought of |
| | + | 3) established a relationship between iGEM and WestLink Innovations |
| | + | 4) Advice on marketing emails + follow up phone calls |
| | | | |
| | <html> | | <html> |
| Line 420: |
Line 384: |
| | </tr> | | </tr> |
| | | | |
| - | <tr>
| |
| - | <td height="10px" bgcolor="#222222">
| |
| - | </td>
| |
| - | </tr>
| |
| | | | |
| - | <tr>
| |
| - | <td>
| |
| - | <a name="Stefan"></a>
| |
| - | <br>
| |
| - | <div class="heading">
| |
| - | STEFAN
| |
| - | </div>
| |
| - | <br>
| |
| - | <div class="heading">
| |
| - | Descriptive Title of What You're Doing
| |
| - | </div>
| |
| - | <br>
| |
| - | <div class="desc">
| |
| - | </html>
| |
| - | WIKI CODING HERE
| |
| - |
| |
| - | <html>
| |
| - | </div>
| |
| - | </td>
| |
| - | </tr>
| |
| | | | |
| | <tr> | | <tr> |
| Line 475: |
Line 415: |
| | <br><br> | | <br><br> |
| | <center> | | <center> |
| - | [[Image:June11.png|700px]] | + | [[Image:LuxOD47AgradPCRannotated.jpg|700px]] |
| | </center> | | </center> |
| | | | |
|
|
CAROL
Continuation of Plasmid Isolation
- Continued with Maxi-prep protocol via DNA precipitation. Unfortunately, no products were found. The washing step needs to be done with caution and the DNA product might have been discarded as well.
- Prepared another overnight culture so that protocol can be repeated.
|
|
|
EMILY
Gradient PCR Take Two
- Objective: To biobrick LuxOD47E in the 2.1pCR-TOPO vector.
- Diluted LuxOD47E template to 40 ng/uL, (2.1uL template + 12.9 uL ddH20)
- Followed protocol from June 6th 2009
- PCR Prodcuts visualized on a 1% agarose gel run at 120 V. See gel photo below. Lanes 1-12 contain mastermix + LuxOD47E in pCR2.1-TOPO vector template. Lane 13 is a negative control with mastermix + ddH2O.
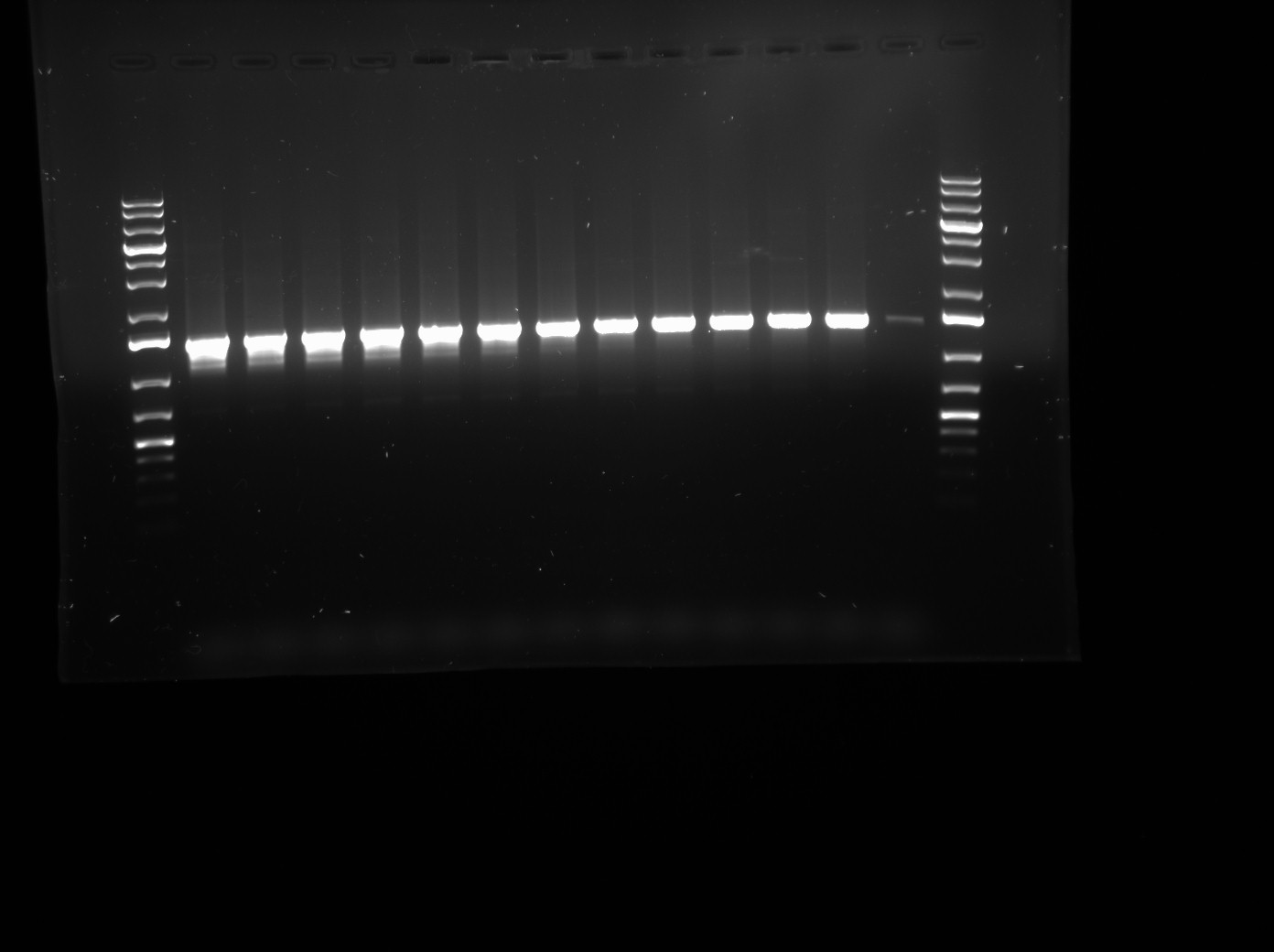
- Analysis: There is considerable contamination in the negative control, however the gradient looks as expected. The source of contamination may possibly be from the primers that were shared between Viki and I. We will try again with new Primer stocks diluted by Thane.
|
|
|
FAHD
Marketing for June 11th 2009
Today, we gad a meeting with Derek Gratz who is a University of Calgary Alumni and the owner of WestLink Innovations. He gave us good ideas on how to approach companies and how to important it is to build a network with these companies. He also gave us names of a few potential companies that might be interested in our project
|
|
|
JAMIE
Recheck Construction Technique for luxOU
No colonies grew on LB+chloramphenicol35 plates. Possible check points: construction digest and ligation/transformation.
1% agarose gel of construction digested vector showed cutting. Redo transformation with positive control
|
|
|
JEREMY
Amplification of luxPQ with BBK using Gradient PCR
The purpose of today’s work was to begin moving the luxPQ operon from the TOPO vector to a BBK vector. luxPQ C24-1 TOPO was used as the DNA template, and in order to flank luxPQ with the appropriate BBK Restriction sites, LuxPQ-RS-F/R primers were used. A platinum-PFX gradient PCR was set up with the following conditions: 94ºC for 5 minutes; 36 X (94ºC for 30 seconds; 58-66ºC for 30 seconds; 72ºC for 4 minutes); 72ºC for 10 minutes; held at 4ºC. The extension, however, should have been set at 68ºC and not 72ºC as per the recommendation of the manufacturer. This PCR ran overnight.
|
|
|
KATIE
Success
I was able to combine all three scripts into one by using the listen event and now it is possible for me to add more types of DNA to PCR and I can continue to easily add quite a few of them. I also was able to fix the error I was getting, which happened because as long as the first item stored in a list was found in the machine’s inventory, it believed everything was there, but when it went to do something with it, the item did not exist and an error occurred.
However, I would still like to change the way it is being used at this time since it is not really efficient to re-declare a variable every time you go through a loop. I will work on changing this at a later date.
|
|
|
KEVIN
Plasmid Isolation of LuxCDABE (luciferase)
The LuxCDABE is being isolated by Caron Chan, but she needed some hands on this, and I was willing to help. LuxCDABE was isolated with Quigen's EndoFree Plasmid Maxi (quigen) according to the specification of the provider. A different kit than the one used in previous experiments was used because the LuxCDABE is very big in size (6kb), and the Minprep would not be capable of isolating the plasmid at a high enough concentration.
Despite of the use of Maxiprep, however, the spectrophotometer seemed to have not picked up any plasmid in the isolated sample, as the purity and concentration were below acceptable. Another Maxiprep may be needed to be done.
|
|
|
MANDY
Gel Making in Second Life
With some inspiration from Sigma Aldrich, I began building a fumehood for the lab. As well, I read up and began writing a notecard describing the preparation of gels for gel electrophoresis. This is just for additional information (just like plate preparation), we are not planning on making this an activity for the virtual lab.
|
|
|
PATRICK
Physical Objects and Nonphysical Objects Do Not Get Along
Retrospective Notebook: This entry was not written on this day, but derived later from working notes I made that day.
Began setting up a workaround for the problems with using SL's 'collision' event for object messaging.
First started changing the object movement script to toggle its 'physics' state on and off, instead of continuously setting its velocity to zero, which would be a much smoother and simpler solution to that problem.
I had by this time built a working RNAP, insofar as it would bind at the promoter, follow the DNA and detach at a terminator.
I also encountered a problem with the RNAP polymerase and the DNA in collision, where because the DNA object was physical (ie, subject to the physics engine in second life) but the RNAP was not physical (so that it could occupy the same space as the DNA without colliding with it), the RNAP would try to move along the strand of DNA, but wound up 'pushing' the movable DNA along. If left alone, this setup would continue crawling across the world indefinitely! Ultimately, when two objects have to live in close proximity with each other, they must both be set to nonphysical or they will exhibit collision glitches like this.
|
|
|
PRIMA
WestLink Innovations Meeting
Fahd, Thane and I went to meet the President from WestLink Innovations.He offered advise on how to contact companies, what to say, what not to say, who we should contact. He offered to help us with networking. It was nice of him to take time out of work to meet and discuss marketing strategies with us.
What I got out of this:
1) Contact information for other companies
2) names to new companies which I hadn't thought of
3) established a relationship between iGEM and WestLink Innovations
4) Advice on marketing emails + follow up phone calls
|
|
|
VICKI
Re-do of gradient PCR because of contamination issues
Purpose: The last gradient PCR for BioBrick amplification failed because of contamination. This is the second attempt.
Materials and methods:
Please refer to the June 4th entry for the observed protocol.
Results: Shown below, with the lane key included. The blank negative control lane is reassuring. The ladder is a GeneElute 1kb Plus DNA ladder.
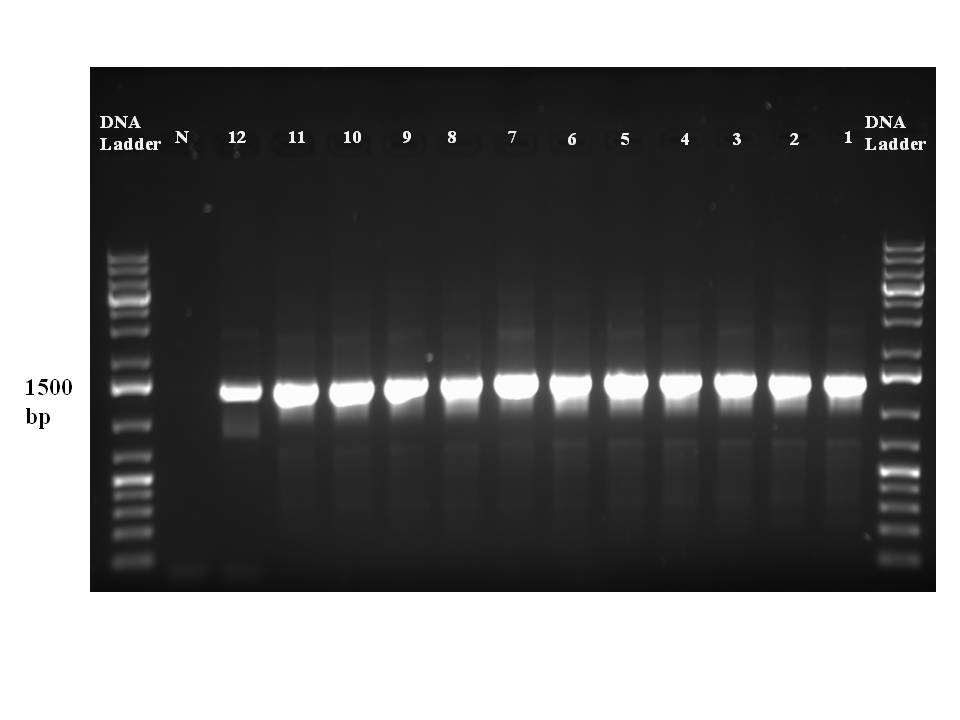
|

 "
"












